[English] 日本語
 Yorodumi
Yorodumi- EMDB-6328: 3D reconstruction from 188 out of 450 fibrils of Abeta(1-40), sel... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6328 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | 3D reconstruction from 188 out of 450 fibrils of Abeta(1-40), selected by mean crossover-to-crossover distance (130 to 150 nm) and curvature (straightest 188 filaments were selected) | |||||||||
 Map data Map data | 3D reconstruction of Abeta(1-40) fibril before any filtering, symmetrization, or masking. Figure 5D of the primary citation was generated from this map by application of helical symmetry and a B factor of -600A^2, with low-pass filtering to 7.5A. | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
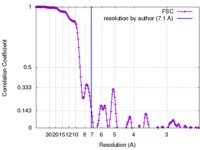
| Method | helical reconstruction / cryo EM / Resolution: 7.1 Å | |||||||||
 Authors Authors | Rohou A / Grigorieff N | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2008 Journal: Proc Natl Acad Sci U S A / Year: 2008Title: Paired beta-sheet structure of an Abeta(1-40) amyloid fibril revealed by electron microscopy. Authors: Carsten Sachse / Marcus Fändrich / Nikolaus Grigorieff /  Abstract: Alzheimer's disease is a neurodegenerative disorder that is characterized by the cerebral deposition of amyloid fibrils formed by Abeta peptide. Despite their prevalence in Alzheimer's and other ...Alzheimer's disease is a neurodegenerative disorder that is characterized by the cerebral deposition of amyloid fibrils formed by Abeta peptide. Despite their prevalence in Alzheimer's and other neurodegenerative diseases, important details of the structure of amyloid fibrils remain unknown. Here, we present a three-dimensional structure of a mature amyloid fibril formed by Abeta(1-40) peptide, determined by electron cryomicroscopy at approximately 8-A resolution. The fibril consists of two protofilaments, each containing approximately 5-nm-long regions of beta-sheet structure. A local twofold symmetry within each region suggests that pairs of beta-sheets are formed from equivalent parts of two Abeta(1-40) peptides contained in each protofilament. The pairing occurs via tightly packed interfaces, reminiscent of recently reported steric zipper structures. However, unlike these previous structures, the beta-sheet pairing is observed within an amyloid fibril and includes significantly longer amino acid sequences. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6328.map.gz emd_6328.map.gz | 365.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6328-v30.xml emd-6328-v30.xml emd-6328.xml emd-6328.xml | 10.1 KB 10.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_6328_fsc.xml emd_6328_fsc.xml | 17.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_6328.jpg emd_6328.jpg | 11.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6328 http://ftp.pdbj.org/pub/emdb/structures/EMD-6328 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6328 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6328 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6328.map.gz / Format: CCP4 / Size: 494.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6328.map.gz / Format: CCP4 / Size: 494.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D reconstruction of Abeta(1-40) fibril before any filtering, symmetrization, or masking. Figure 5D of the primary citation was generated from this map by application of helical symmetry and a B factor of -600A^2, with low-pass filtering to 7.5A. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.2 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : human Abeta(1-40)
| Entire | Name: human Abeta(1-40) |
|---|---|
| Components |
|
-Supramolecule #1000: human Abeta(1-40)
| Supramolecule | Name: human Abeta(1-40) / type: sample / ID: 1000 / Oligomeric state: Helical assembly / Number unique components: 1 |
|---|
-Macromolecule #1: Amyloid beta peptide (1-40)
| Macromolecule | Name: Amyloid beta peptide (1-40) / type: protein_or_peptide / ID: 1 / Name.synonym: Abeta(1-40) / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: human Homo sapiens (human) / synonym: human |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.8 / Details: 50 mM borate |
| Grid | Details: Quantifoil R1.2/1.3 Cu 400 mesh grids |
| Vitrification | Cryogen name: ETHANE / Instrument: HOMEMADE PLUNGER / Details: Vitrification carried out in cold room at 277 K. / Method: Blot for 7 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F30 |
|---|---|
| Date | Feb 9, 2006 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 7.0 µm / Number real images: 63 / Average electron dose: 35 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 3.629 µm / Nominal defocus min: 2.027 µm / Nominal magnification: 59000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)