[English] 日本語
 Yorodumi
Yorodumi- SASDFV3: Cytokine-like nuclear factor dehydrogenase domain, NPAC DH (NPAC ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | 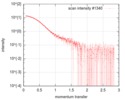 |
|---|---|
 Sample Sample | Cytokine-like nuclear factor dehydrogenase domain, NPAC DH (NPAC delta-261)
|
| Function / homology |  Function and homology information Function and homology informationtranscription elongation-coupled chromatin remodeling / chromatin-protein adaptor activity / nucleosome binding / transcription initiation-coupled chromatin remodeling / NAD binding / NADP binding / nucleosome / histone binding / chromatin binding / chromatin ...transcription elongation-coupled chromatin remodeling / chromatin-protein adaptor activity / nucleosome binding / transcription initiation-coupled chromatin remodeling / NAD binding / NADP binding / nucleosome / histone binding / chromatin binding / chromatin / DNA binding / nucleoplasm / cytosol Similarity search - Function |
| Biological species |  Homo sapiens (human) Homo sapiens (human) |
 Citation Citation |  Journal: Cell Rep / Year: 2019 Journal: Cell Rep / Year: 2019Title: A Tail-Based Mechanism Drives Nucleosome Demethylation by the LSD2/NPAC Multimeric Complex. Authors: Chiara Marabelli / Biagina Marrocco / Simona Pilotto / Sagar Chittori / Sarah Picaud / Sara Marchese / Giuseppe Ciossani / Federico Forneris / Panagis Filippakopoulos / Guy Schoehn / Daniela ...Authors: Chiara Marabelli / Biagina Marrocco / Simona Pilotto / Sagar Chittori / Sarah Picaud / Sara Marchese / Giuseppe Ciossani / Federico Forneris / Panagis Filippakopoulos / Guy Schoehn / Daniela Rhodes / Sriram Subramaniam / Andrea Mattevi /       Abstract: LSD1 and LSD2 are homologous histone demethylases with opposite biological outcomes related to chromatin silencing and transcription elongation, respectively. Unlike LSD1, LSD2 nucleosome-demethylase ...LSD1 and LSD2 are homologous histone demethylases with opposite biological outcomes related to chromatin silencing and transcription elongation, respectively. Unlike LSD1, LSD2 nucleosome-demethylase activity relies on a specific linker peptide from the multidomain protein NPAC. We used single-particle cryoelectron microscopy (cryo-EM), in combination with kinetic and mutational analysis, to analyze the mechanisms underlying the function of the human LSD2/NPAC-linker/nucleosome complex. Weak interactions between LSD2 and DNA enable multiple binding modes for the association of the demethylase to the nucleosome. The demethylase thereby captures mono- and dimethyl Lys4 of the H3 tail to afford histone demethylation. Our studies also establish that the dehydrogenase domain of NPAC serves as a catalytically inert oligomerization module. While LSD1/CoREST forms a nucleosome docking platform at silenced gene promoters, LSD2/NPAC is a multifunctional enzyme complex with flexible linkers, tailored for rapid chromatin modification, in conjunction with the advance of the RNA polymerase on actively transcribed genes. |
 Contact author Contact author |
|
- Structure visualization
Structure visualization
- Downloads & links
Downloads & links
-Data source
| SASBDB page |  SASDFV3 SASDFV3 |
|---|
-Related structure data
| Related structure data |  4704C  4705C  4710C  4711C  4712C  6r1tC  6r1uC  6r25C C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- External links
External links
| Related items in Molecule of the Month |
|---|
-Models
- Sample
Sample
 Sample Sample | Name: Cytokine-like nuclear factor dehydrogenase domain, NPAC DH (NPAC delta-261) Specimen concentration: 1.5 mg/ml |
|---|---|
| Buffer | Name: 15 mM HEPES, 200 mM NaCl / pH: 7.3 |
| Entity #1518 | Name: NPAC DH / Type: protein / Description: NPAC dehydrogenase domain / Formula weight: 31.369 / Num. of mol.: 4 / Source: Homo sapiens / References: UniProt: Q49A26 Sequence: GSITPTDKKI GFLGLGLMGS GIVSNLLKMG HTVTVWNRTA EKCDLFIQEG ARLGRTPAEV VSTCDITFAC VSDPKAAKDL VLGPSGVLQG IRPGKCYVDM STVDADTVTE LAQVIVSRGG RFLEAPVSGN QQLSNDGMLV ILAAGDRGLY EDCSSCFQAM GKTSFFLGEV ...Sequence: GSITPTDKKI GFLGLGLMGS GIVSNLLKMG HTVTVWNRTA EKCDLFIQEG ARLGRTPAEV VSTCDITFAC VSDPKAAKDL VLGPSGVLQG IRPGKCYVDM STVDADTVTE LAQVIVSRGG RFLEAPVSGN QQLSNDGMLV ILAAGDRGLY EDCSSCFQAM GKTSFFLGEV GNAAKMMLIV NMVQGSFMAT IAEGLTLAQV TGQSQQTLLD ILNQGQLASI FLDQKCQNIL QGNFKPDFYL KYIQKDLRLA IALGDAVNHP TPMAAAANEV YKRAKALDQS DNDMSAVYRA YIH |
-Experimental information
| Beam | Instrument name: ESRF BM29 / City: Grenoble / 国: France  / Type of source: X-ray synchrotron / Wavelength: 0.1 Å / Dist. spec. to detc.: 2.87 mm / Type of source: X-ray synchrotron / Wavelength: 0.1 Å / Dist. spec. to detc.: 2.87 mm | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Name: Pilatus 1M / Type: Dectris / Pixsize x: 172 mm | ||||||||||||||||||||||||||||||
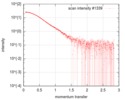
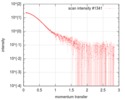
| Scan | Measurement date: Jan 26, 2018 / Storage temperature: 4 °C / Cell temperature: 20 °C / Exposure time: 1 sec. / Number of frames: 173 / Unit: 1/nm /
| ||||||||||||||||||||||||||||||
| Distance distribution function P(R) |
| ||||||||||||||||||||||||||||||
| Result |
|
 Movie
Movie Controller
Controller