+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7n4y | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
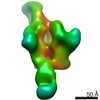
| Title | The structure of bovine thyroglobulin with iodinated tyrosines | |||||||||
 Components Components | Thyroglobulin | |||||||||
 Keywords Keywords | HORMONE / thyroid hormone synthesis | |||||||||
| Function / homology |  Function and homology information Function and homology informationhormone biosynthetic process / iodide transport / thyroid hormone generation / regulation of myelination / thyroid gland development / hormone activity / extracellular space / identical protein binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 2.61 Å | |||||||||
 Authors Authors | Kim, K. / Clarke, O.B. | |||||||||
| Funding support | 1items
| |||||||||
 Citation Citation |  Journal: Acta Crystallogr D Struct Biol / Year: 2021 Journal: Acta Crystallogr D Struct Biol / Year: 2021Title: The structure of natively iodinated bovine thyroglobulin. Authors: Kookjoo Kim / Mykhailo Kopylov / Daija Bobe / Kotaro Kelley / Edward T Eng / Peter Arvan / Oliver B Clarke /  Abstract: Thyroglobulin is a homodimeric glycoprotein that is essential for the generation of thyroid hormones in vertebrates. Upon secretion into the lumen of follicles in the thyroid gland, tyrosine residues ...Thyroglobulin is a homodimeric glycoprotein that is essential for the generation of thyroid hormones in vertebrates. Upon secretion into the lumen of follicles in the thyroid gland, tyrosine residues within the protein become iodinated to produce monoiodotyrosine (MIT) and diiodotyrosine (DIT). A subset of evolutionarily conserved pairs of DIT (and MIT) residues can then engage in oxidative coupling reactions that yield either thyroxine (T; produced from coupling of a DIT `acceptor' with a DIT `donor') or triiodothyronine (T; produced from coupling of a DIT acceptor with an MIT donor). Although multiple iodotyrosine residues have been identified as potential donors and acceptors, the specificity and structural context of the pairings (i.e. which donor is paired with which acceptor) have remained unclear. Here, single-particle cryogenic electron microscopy (cryoEM) was used to generate a high-resolution reconstruction of bovine thyroglobulin (2.3 Å resolution in the core region and 2.6 Å overall), allowing the structural characterization of two post-reaction acceptor-donor pairs as well as tyrosine residues modified as MIT and DIT. A substantial spatial separation between donor Tyr149 and acceptor Tyr24 was observed, suggesting that for thyroxine synthesis significant peptide motion is required for coupling at the evolutionarily conserved thyroglobulin amino-terminus. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7n4y.cif.gz 7n4y.cif.gz | 856.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7n4y.ent.gz pdb7n4y.ent.gz | 685.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7n4y.json.gz 7n4y.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  7n4y_validation.pdf.gz 7n4y_validation.pdf.gz | 1.6 MB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  7n4y_full_validation.pdf.gz 7n4y_full_validation.pdf.gz | 1.7 MB | Display | |
| Data in XML |  7n4y_validation.xml.gz 7n4y_validation.xml.gz | 131.7 KB | Display | |
| Data in CIF |  7n4y_validation.cif.gz 7n4y_validation.cif.gz | 207.5 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/n4/7n4y https://data.pdbj.org/pub/pdb/validation_reports/n4/7n4y ftp://data.pdbj.org/pub/pdb/validation_reports/n4/7n4y ftp://data.pdbj.org/pub/pdb/validation_reports/n4/7n4y | HTTPS FTP |
-Related structure data
| Related structure data |  24181MC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10833 (Title: The structure of natively iodinated bovine thyroglobulin EMPIAR-10833 (Title: The structure of natively iodinated bovine thyroglobulinData size: 1.2 TB Data #1: Unaligned multi-frame micrographs of bovine thyroglobulin [micrographs - multiframe]) |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
-Protein / Non-polymers , 2 types, 310 molecules AB

| #1: Protein | Mass: 303087.844 Da / Num. of mol.: 2 / Source method: isolated from a natural source / Source: (natural)  #6: Water | ChemComp-HOH / | |
|---|
-Sugars , 4 types, 18 molecules 
| #2: Polysaccharide | Source method: isolated from a genetically manipulated source #3: Polysaccharide | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose Source method: isolated from a genetically manipulated source #4: Polysaccharide | Source method: isolated from a genetically manipulated source #5: Sugar | ChemComp-NAG / |
|---|
-Details
| Has ligand of interest | Y |
|---|
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Bovine thyroglobulin purified from bovine thyroid / Type: COMPLEX / Entity ID: #1 / Source: NATURAL |
|---|---|
| Molecular weight | Value: 0.66 MDa / Experimental value: YES |
| Source (natural) | Organism:  |
| Buffer solution | pH: 7.4 |
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| Specimen support | Grid material: GOLD / Grid mesh size: 300 divisions/in. / Grid type: Quantifoil R0.6/1 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: TFS KRIOS |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD |
| Image recording | Electron dose: 71.32 e/Å2 / Detector mode: COUNTING / Film or detector model: GATAN K2 SUMMIT (4k x 4k) |
- Processing
Processing
| Software | Name: PHENIX / Version: 1.18.2_3874: / Classification: refinement | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| EM software |
| ||||||||||||||||||||||||
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||||||||||
| 3D reconstruction | Resolution: 2.61 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 554053 / Symmetry type: POINT | ||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller










 PDBj
PDBj

