[English] 日本語
 Yorodumi
Yorodumi- EMDB-9373: Single particle reconstruction of the symmetric core an engineere... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9373 | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Single particle reconstruction of the symmetric core an engineered protein scaffold | |||||||||||||||||||||
 Map data Map data | Single particle reconstruction of the symmetric core of an engineered protein scaffold (DARP14) | |||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
 Keywords Keywords | protein engineering / symmetric scaffold / small protein cryo-EM / display platform / BIOSYNTHETIC PROTEIN | |||||||||||||||||||||
| Function / homology | 5-carboxymethyl-2-hydroxymuconate isomerase / 5-carboxymethyl-2-hydroxymuconate isomerase / 5-carboxymethyl-2-hydroxymuconate delta-isomerase activity / : / Tautomerase/MIF superfamily / 5-carboxymethyl-2-hydroxymuconate isomerase Function and homology information Function and homology information | |||||||||||||||||||||
| Biological species |  Pseudomonas aeruginosa PAO1 (bacteria) / Pseudomonas aeruginosa PAO1 (bacteria) /   Pyrococcus horikoshii OT3 (archaea) / Pyrococcus horikoshii OT3 (archaea) /   Pyrococcus horikoshii (strain ATCC 700860 / DSM 12428 / JCM 9974 / NBRC 100139 / OT-3) (archaea) / Pyrococcus horikoshii (strain ATCC 700860 / DSM 12428 / JCM 9974 / NBRC 100139 / OT-3) (archaea) /  Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria) Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria) | |||||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||||||||||||||
 Authors Authors | Liu Y / Huynh D | |||||||||||||||||||||
| Funding support |  United States, 6 items United States, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2019 Journal: Nat Commun / Year: 2019Title: A 3.8 Å resolution cryo-EM structure of a small protein bound to an imaging scaffold. Authors: Yuxi Liu / Duc T Huynh / Todd O Yeates /  Abstract: Proteins smaller than about 50 kDa are currently too small to be imaged at high resolution by cryo-electron microscopy (cryo-EM), leaving most protein molecules in the cell beyond the reach of this ...Proteins smaller than about 50 kDa are currently too small to be imaged at high resolution by cryo-electron microscopy (cryo-EM), leaving most protein molecules in the cell beyond the reach of this powerful structural technique. Here we use a designed protein scaffold to bind and symmetrically display 12 copies of a small 26 kDa protein, green fluorescent protein (GFP). We show that the bound cargo protein is held rigidly enough to visualize it at a resolution of 3.8 Å by cryo-EM, where specific structural features of the protein are visible. The designed scaffold is modular and can be modified through modest changes in its amino acid sequence to bind and display diverse proteins for imaging, thus providing a general method to break through the lower size limitation in cryo-EM. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9373.map.gz emd_9373.map.gz | 85.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9373-v30.xml emd-9373-v30.xml emd-9373.xml emd-9373.xml | 21.5 KB 21.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9373.png emd_9373.png | 142.6 KB | ||
| Masks |  emd_9373_msk_1.map emd_9373_msk_1.map | 91.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-9373.cif.gz emd-9373.cif.gz | 6.7 KB | ||
| Others |  emd_9373_half_map_1.map.gz emd_9373_half_map_1.map.gz emd_9373_half_map_2.map.gz emd_9373_half_map_2.map.gz | 84.7 MB 84.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9373 http://ftp.pdbj.org/pub/emdb/structures/EMD-9373 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9373 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9373 | HTTPS FTP |
-Validation report
| Summary document |  emd_9373_validation.pdf.gz emd_9373_validation.pdf.gz | 945 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_9373_full_validation.pdf.gz emd_9373_full_validation.pdf.gz | 944.6 KB | Display | |
| Data in XML |  emd_9373_validation.xml.gz emd_9373_validation.xml.gz | 13.4 KB | Display | |
| Data in CIF |  emd_9373_validation.cif.gz emd_9373_validation.cif.gz | 15.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9373 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9373 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9373 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9373 | HTTPS FTP |
-Related structure data
| Related structure data |  6nhtMC  9374C  6nhvC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9373.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9373.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Single particle reconstruction of the symmetric core of an engineered protein scaffold (DARP14) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_9373_msk_1.map emd_9373_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
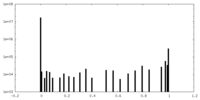
| Density Histograms |
-Half map: Half map A
| File | emd_9373_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
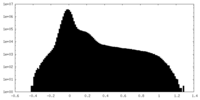
| Density Histograms |
-Half map: Half map B
| File | emd_9373_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Subunit A with DARPin Subunit B
| Entire | Name: Subunit A with DARPin Subunit B |
|---|---|
| Components |
|
-Supramolecule #1: Subunit A with DARPin Subunit B
| Supramolecule | Name: Subunit A with DARPin Subunit B / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|
-Supramolecule #2: Subunit B
| Supramolecule | Name: Subunit B / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Pseudomonas aeruginosa PAO1 (bacteria) Pseudomonas aeruginosa PAO1 (bacteria) |
-Supramolecule #3: Subunit A with DARPin
| Supramolecule | Name: Subunit A with DARPin / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Pyrococcus horikoshii OT3 (archaea) Pyrococcus horikoshii OT3 (archaea) |
-Macromolecule #1: DARP14 - Subunit A with DARPin
| Macromolecule | Name: DARP14 - Subunit A with DARPin / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Pyrococcus horikoshii (strain ATCC 700860 / DSM 12428 / JCM 9974 / NBRC 100139 / OT-3) (archaea) Pyrococcus horikoshii (strain ATCC 700860 / DSM 12428 / JCM 9974 / NBRC 100139 / OT-3) (archaea)Strain: ATCC 700860 / DSM 12428 / JCM 9974 / NBRC 100139 / OT-3 |
| Molecular weight | Theoretical: 34.71782 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MRITTKVGDK GSTRLFGGEE VWKDSPIIEA NGTLDELTSF IGEAKHYVDE EMKGILEEIQ NDIYKIMGEI GSKGKIEGIS EERIAWLLK LILRYMEMVN LKSFVLPGGT LESAKLDVCR TIARRALRKV LTVTREFGIG AEAAAYLLAL SDLLFLLARV I EIEQGKKL ...String: MRITTKVGDK GSTRLFGGEE VWKDSPIIEA NGTLDELTSF IGEAKHYVDE EMKGILEEIQ NDIYKIMGEI GSKGKIEGIS EERIAWLLK LILRYMEMVN LKSFVLPGGT LESAKLDVCR TIARRALRKV LTVTREFGIG AEAAAYLLAL SDLLFLLARV I EIEQGKKL LEAARAGQDD EVRILMANGA DVNAADDVGV TPLHLAAQRG HLEIVEVLLK CGADVNAADL WGQTPLHLAA TA GHLEIVE VLLKNGADVN ARDNIGHTPL HLAAWAGHLE IVEVLLKYGA DVNAQDKFGK TPFDLAIDNG NEDIAEVLQK AA |
-Macromolecule #2: DARP14 - Subunit B
| Macromolecule | Name: DARP14 - Subunit B / type: protein_or_peptide / ID: 2 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria) Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria)Strain: ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1 |
| Molecular weight | Theoretical: 14.346274 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPHLVIEATA NLRLETSPGE LLEQANKALF ASGQFGEADI KSRFVTLEAY RQGTAAVERA YLHACLSILD GRDIATRTLL GASLCAVLA EAVAGGGEEG VQVSVEVREM ERLSYAKRVV ARQRLEHHHH HH UniProtKB: 5-carboxymethyl-2-hydroxymuconate isomerase |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: 2.5 microliter of sample, 0 sec wait, 0 sec drain, 3 sec blot, -15 blot force, grids pre-treated with 0.1% poly-lysine for 6 hours. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Frames/image: 3-20 / Number grids imaged: 1 / Number real images: 1929 / Average exposure time: 8.0 sec. / Average electron dose: 56.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Details | Initial local fitting by Chimera and individual residues refined using phenix.real_space_refine |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
| Output model |  PDB-6nht: |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)