[English] 日本語
 Yorodumi
Yorodumi- EMDB-9250: CENP-A nucleosome bound by two copies of CENP-C(CD) and one copy ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9250 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CENP-A nucleosome bound by two copies of CENP-C(CD) and one copy CENP-N(NT) | |||||||||
 Map data Map data | CENP-A nucleosome bound by two copies of CENP-C(CD) and one copy CENP-N(NT) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | centromere / CENP-A / kinetochore / nucleosome / NUCLEAR PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationspindle attachment to meiosis I kinetochore / centromeric DNA binding / CENP-A containing chromatin assembly / protein localization to chromosome, centromeric region / kinetochore assembly / condensed chromosome, centromeric region / attachment of mitotic spindle microtubules to kinetochore / inner kinetochore / establishment of mitotic spindle orientation / mitotic cytokinesis ...spindle attachment to meiosis I kinetochore / centromeric DNA binding / CENP-A containing chromatin assembly / protein localization to chromosome, centromeric region / kinetochore assembly / condensed chromosome, centromeric region / attachment of mitotic spindle microtubules to kinetochore / inner kinetochore / establishment of mitotic spindle orientation / mitotic cytokinesis / chromosome, centromeric region / pericentric heterochromatin / negative regulation of megakaryocyte differentiation / protein localization to CENP-A containing chromatin / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / Packaging Of Telomere Ends / Amplification of signal from unattached kinetochores via a MAD2 inhibitory signal / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / Deposition of new CENPA-containing nucleosomes at the centromere / Mitotic Prometaphase / telomere organization / EML4 and NUDC in mitotic spindle formation / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / RNA Polymerase I Promoter Opening / Inhibition of DNA recombination at telomere / Assembly of the ORC complex at the origin of replication / Meiotic synapsis / SUMOylation of chromatin organization proteins / Regulation of endogenous retroelements by the Human Silencing Hub (HUSH) complex / Resolution of Sister Chromatid Cohesion / DNA methylation / Condensation of Prophase Chromosomes / Chromatin modifications during the maternal to zygotic transition (MZT) / SIRT1 negatively regulates rRNA expression / HCMV Late Events / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / PRC2 methylates histones and DNA / Regulation of endogenous retroelements by KRAB-ZFP proteins / Defective pyroptosis / HDACs deacetylate histones / Regulation of endogenous retroelements by Piwi-interacting RNAs (piRNAs) / Nonhomologous End-Joining (NHEJ) / RNA Polymerase I Promoter Escape / chromosome segregation / Transcriptional regulation by small RNAs / RHO GTPases Activate Formins / Formation of the beta-catenin:TCF transactivating complex / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / HDMs demethylate histones / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / G2/M DNA damage checkpoint / Negative Regulation of CDH1 Gene Transcription / NoRC negatively regulates rRNA expression / PKMTs methylate histone lysines / kinetochore / B-WICH complex positively regulates rRNA expression / DNA Damage/Telomere Stress Induced Senescence / Meiotic recombination / Pre-NOTCH Transcription and Translation / Activation of anterior HOX genes in hindbrain development during early embryogenesis / Transcriptional regulation of granulopoiesis / Metalloprotease DUBs / RMTs methylate histone arginines / HCMV Early Events / structural constituent of chromatin / Separation of Sister Chromatids / UCH proteinases / nucleosome / mitotic cell cycle / heterochromatin formation / nucleosome assembly / HATs acetylate histones / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / MLL4 and MLL3 complexes regulate expression of PPARG target genes in adipogenesis and hepatic steatosis / chromatin organization / RUNX1 regulates transcription of genes involved in differentiation of HSCs / Processing of DNA double-strand break ends / Senescence-Associated Secretory Phenotype (SASP) / midbody / Oxidative Stress Induced Senescence / Estrogen-dependent gene expression / chromosome, telomeric region / Ub-specific processing proteases / nuclear body / Amyloid fiber formation / protein heterodimerization activity / negative regulation of cell population proliferation / cell division / chromatin binding / protein-containing complex / DNA binding / RNA binding / extracellular exosome / extracellular region / nucleoplasm / membrane / identical protein binding Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Allu PK / Black BE | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Curr Biol / Year: 2019 Journal: Curr Biol / Year: 2019Title: Structure of the Human Core Centromeric Nucleosome Complex. Authors: Praveen Kumar Allu / Jennine M Dawicki-McKenna / Trevor Van Eeuwen / Moriya Slavin / Merav Braitbard / Chen Xu / Nir Kalisman / Kenji Murakami / Ben E Black /   Abstract: Centromeric nucleosomes are at the interface of the chromosome and the kinetochore that connects to spindle microtubules in mitosis. The core centromeric nucleosome complex (CCNC) harbors the ...Centromeric nucleosomes are at the interface of the chromosome and the kinetochore that connects to spindle microtubules in mitosis. The core centromeric nucleosome complex (CCNC) harbors the histone H3 variant, CENP-A, and its binding proteins, CENP-C (through its central domain; CD) and CENP-N (through its N-terminal domain; NT). CENP-C can engage nucleosomes through two domains: the CD and the CENP-C motif (CM). CENP-C is part of the CCNC by virtue of its high specificity for CENP-A nucleosomes and ability to stabilize CENP-A at the centromere. CENP-C is thought to engage a neighboring nucleosome, either one containing conventional H3 or CENP-A, and a crystal structure of a nucleosome complex containing two copies of CENP-C was reported. Recent structures containing a single copy of CENP-N bound to the CENP-A nucleosome in the absence of CENP-C were reported. Here, we find that one copy of CENP-N is lost for every two copies of CENP-C on centromeric chromatin just prior to kinetochore formation. We present the structures of symmetric and asymmetric forms of the CCNC that vary in CENP-N stoichiometry. Our structures explain how the central domain of CENP-C achieves its high specificity for CENP-A nucleosomes and how CENP-C and CENP-N sandwich the histone H4 tail. The natural centromeric DNA path in our structures corresponds to symmetric surfaces for CCNC assembly, deviating from what is observed in prior structures using artificial sequences. At mitosis, we propose that CCNC asymmetry accommodates its asymmetric connections at the chromosome/kinetochore interface. VIDEO ABSTRACT. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9250.map.gz emd_9250.map.gz | 28.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9250-v30.xml emd-9250-v30.xml emd-9250.xml emd-9250.xml | 23.1 KB 23.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_9250_fsc.xml emd_9250_fsc.xml | 7.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_9250.png emd_9250.png | 242.3 KB | ||
| Filedesc metadata |  emd-9250.cif.gz emd-9250.cif.gz | 6.8 KB | ||
| Others |  emd_9250_additional.map.gz emd_9250_additional.map.gz | 23.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9250 http://ftp.pdbj.org/pub/emdb/structures/EMD-9250 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9250 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9250 | HTTPS FTP |
-Related structure data
| Related structure data |  6muoMC  9251C  9252C  6mupC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9250.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9250.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CENP-A nucleosome bound by two copies of CENP-C(CD) and one copy CENP-N(NT) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: CENP-A nucleosome bound by two copies of CENP-C(CD)...
| File | emd_9250_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CENP-A nucleosome bound by two copies of CENP-C(CD) and one copy CENP-N(NT) | ||||||||||||
| Projections & Slices |
| ||||||||||||
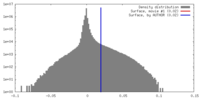
| Density Histograms |
- Sample components
Sample components
-Entire : CENP-A chromatin complex bound with CENP-C and CENP-N of CCAN kin...
| Entire | Name: CENP-A chromatin complex bound with CENP-C and CENP-N of CCAN kinetochore components |
|---|---|
| Components |
|
-Supramolecule #1: CENP-A chromatin complex bound with CENP-C and CENP-N of CCAN kin...
| Supramolecule | Name: CENP-A chromatin complex bound with CENP-C and CENP-N of CCAN kinetochore components type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1-#8 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 276 KDa |
-Macromolecule #1: Histone H3-like centromeric protein A
| Macromolecule | Name: Histone H3-like centromeric protein A / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.993037 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: HQHSRRRQGW LKEIRKLQKS THLLIRKLPF SRLAREICVK FTRGVDFNWQ AQALLALQEA AEAFLVHLFE DAYLLTLHAG RVTLFPKDV QLARRIRGLE EGL UniProtKB: Histone H3-like centromeric protein A |
-Macromolecule #2: Histone H4
| Macromolecule | Name: Histone H4 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 10.604521 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: KGLGKGGAKR HRKVLRDNIQ GITKPAIRRL ARRGGVKRIS GLIYEETRGV LKVFLENVIR DAVTYTEHAK RKTVTAMDVV YALKRQGRT LYGFG UniProtKB: Histone H4 |
-Macromolecule #3: Histone H2A type 1-C
| Macromolecule | Name: Histone H2A type 1-C / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.494393 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: KAKSRSSRAG LQFPVGRVHR LLRKGNYAER VGAGAPVYLA AVLEYLTAEI LELAGNAARD NKKTRIIPRH LQLAIRNDEE LNKLLGRVT IAQGGVLPNI QSVLLP UniProtKB: Histone H2A type 1-C |
-Macromolecule #4: Histone H2B type 2-F
| Macromolecule | Name: Histone H2B type 2-F / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 10.249723 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: RKESYSVYVY KVLKQVHPDT GISSKAMGIM NSFVNDIFER IAGEASRLAH YNKRSTITSR EIQTAVRLLL PGELAKHAVS EGTKAVTKY TSS UniProtKB: Histone H2B type 2-F |
-Macromolecule #7: Centromere protein C
| Macromolecule | Name: Centromere protein C / type: protein_or_peptide / ID: 7 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 2.508834 KDa |
| Sequence | String: TKSRRISRRP SDWWVVKSEE UniProtKB: Centromere protein C |
-Macromolecule #8: Centromere protein N
| Macromolecule | Name: Centromere protein N / type: protein_or_peptide / ID: 8 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 25.052885 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDETVAEFIK RTILKIPMNE LTTILKAWDF LSENQLQTVN FRQRKESVVQ HLIHLCEEKR ASISDAALLD IIYMQFHQHQ KVWDVFQMS KGPGEDVDLF DMKQFKNSFK KILQRALKNV TVSFRETEEN AVWIRIAWGT QYTKPNQYKP TYVVYYSQTP Y AFTSSSML ...String: MDETVAEFIK RTILKIPMNE LTTILKAWDF LSENQLQTVN FRQRKESVVQ HLIHLCEEKR ASISDAALLD IIYMQFHQHQ KVWDVFQMS KGPGEDVDLF DMKQFKNSFK KILQRALKNV TVSFRETEEN AVWIRIAWGT QYTKPNQYKP TYVVYYSQTP Y AFTSSSML RRNTPLLGQA LTIASKHHQI VKMDLRSRYL DSLKAIVFKQ YNQT UniProtKB: Centromere protein N |
-Macromolecule #5: DNA/RNA (147-MER)
| Macromolecule | Name: DNA/RNA (147-MER) / type: dna / ID: 5 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 45.141918 KDa |
| Sequence | String: (DA)(DT)(DC)(DA)(DA)(DA)(DT)(DA)(DT)(DC) (DC)(DA)(DC)(DC)(DT)(DG)(DC)(DA)(DG)(DA) (DT)(DT)(DC)(DT)(DA)(DC)(DC)(DA)(DA) (DA)(DA)(DG)(DT)(DG)(DT)(DA)(DT)(DT)(DT) (DG) (DG)(DA)(DA)(DA)(DC)(DT) ...String: (DA)(DT)(DC)(DA)(DA)(DA)(DT)(DA)(DT)(DC) (DC)(DA)(DC)(DC)(DT)(DG)(DC)(DA)(DG)(DA) (DT)(DT)(DC)(DT)(DA)(DC)(DC)(DA)(DA) (DA)(DA)(DG)(DT)(DG)(DT)(DA)(DT)(DT)(DT) (DG) (DG)(DA)(DA)(DA)(DC)(DT)(DG)(DC) (DT)(DC)(DC)(DA)(DT)(DC)(DA)(DA)(DA)(DA) (DG)(DG) (DC)(DA)(DT)(DG)(DT)(DT)(DC) (DA)(DG)(DC)(DT)(DC)(DT)(DG)(DT)(DG)(DA) (DG)(DT)(DG) (DA)(DA)(DA)(DC)(DT)(DC) (DC)(DA)(DT)(DC)(DA)(DT)(DC)(DA)(DC)(DA) (DA)(DA)(DG)(DA) (DA)(DT)(DA)(DT)(DT) (DC)(DT)(DG)(DA)(DG)(DA)(DA)(DT)(DG)(DC) (DT)(DT)(DC)(DC)(DG) (DT)(DT)(DT)(DG) (DC)(DC)(DT)(DT)(DT)(DT)(DA)(DT)(DA)(DT) (DG)(DA)(DA)(DC)(DT)(DT) (DC)(DC)(DT) (DC)(DG)(DA)(DT) |
-Macromolecule #6: DNA/RNA (147-MER)
| Macromolecule | Name: DNA/RNA (147-MER) / type: dna / ID: 6 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 45.582188 KDa |
| Sequence | String: (DA)(DT)(DC)(DG)(DA)(DG)(DG)(DA)(DA)(DG) (DT)(DT)(DC)(DA)(DT)(DA)(DT)(DA)(DA)(DA) (DA)(DG)(DG)(DC)(DA)(DA)(DA)(DC)(DG) (DG)(DA)(DA)(DG)(DC)(DA)(DT)(DT)(DC)(DT) (DC) (DA)(DG)(DA)(DA)(DT)(DA) ...String: (DA)(DT)(DC)(DG)(DA)(DG)(DG)(DA)(DA)(DG) (DT)(DT)(DC)(DA)(DT)(DA)(DT)(DA)(DA)(DA) (DA)(DG)(DG)(DC)(DA)(DA)(DA)(DC)(DG) (DG)(DA)(DA)(DG)(DC)(DA)(DT)(DT)(DC)(DT) (DC) (DA)(DG)(DA)(DA)(DT)(DA)(DT)(DT) (DC)(DT)(DT)(DT)(DG)(DT)(DG)(DA)(DT)(DG) (DA)(DT) (DG)(DG)(DA)(DG)(DT)(DT)(DT) (DC)(DA)(DC)(DT)(DC)(DA)(DC)(DA)(DG)(DA) (DG)(DC)(DT) (DG)(DA)(DA)(DC)(DA)(DT) (DG)(DC)(DC)(DT)(DT)(DT)(DT)(DG)(DA)(DT) (DG)(DG)(DA)(DG) (DC)(DA)(DG)(DT)(DT) (DT)(DC)(DC)(DA)(DA)(DA)(DT)(DA)(DC)(DA) (DC)(DT)(DT)(DT)(DT) (DG)(DG)(DT)(DA) (DG)(DA)(DA)(DT)(DC)(DT)(DG)(DC)(DA)(DG) (DG)(DT)(DG)(DG)(DA)(DT) (DA)(DT)(DT) (DT)(DG)(DA)(DT) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Details: 10 mM HEPES, 50 mN NaCl, 1 mM EDTA, 1mM DTT |
| Grid | Model: C-flat-1.2/1.3 4C / Material: COPPER / Mesh: 400 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 40 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 30.0 kPa / Details: unspecified |
| Vitrification | Cryogen name: ETHANE / Instrument: LEICA EM CPC / Details: Blot for 8 seconds before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: SUPER-RESOLUTION / Number grids imaged: 2 / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)