[English] 日本語
 Yorodumi
Yorodumi- EMDB-8977: Cryo-EM structure at 3.2 A resolution of HIV-1 fusion peptide-dir... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8977 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
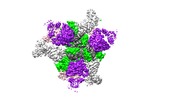
| Title | Cryo-EM structure at 3.2 A resolution of HIV-1 fusion peptide-directed antibody, A12V163-b.01, elicited by vaccination of Rhesus macaques, in complex with stabilized HIV-1 Env BG505 DS-SOSIP, which was also bound to antibodies VRC03 and PGT122 | |||||||||
 Map data Map data | primary map | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated perturbation of host defense response / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell ...symbiont-mediated perturbation of host defense response / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / host cell plasma membrane / virion membrane / structural molecule activity / membrane / identical protein binding Similarity search - Function | |||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /   Homo sapiens (human) Homo sapiens (human) | |||||||||
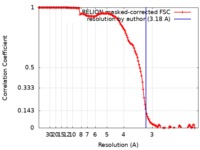
| Method | single particle reconstruction / cryo EM / Resolution: 3.18 Å | |||||||||
 Authors Authors | Acharya P / Eng ET / Kwong PD | |||||||||
 Citation Citation |  Journal: Cell / Year: 2019 Journal: Cell / Year: 2019Title: Antibody Lineages with Vaccine-Induced Antigen-Binding Hotspots Develop Broad HIV Neutralization. Authors: Rui Kong / Hongying Duan / Zizhang Sheng / Kai Xu / Priyamvada Acharya / Xuejun Chen / Cheng Cheng / Adam S Dingens / Jason Gorman / Mallika Sastry / Chen-Hsiang Shen / Baoshan Zhang / ...Authors: Rui Kong / Hongying Duan / Zizhang Sheng / Kai Xu / Priyamvada Acharya / Xuejun Chen / Cheng Cheng / Adam S Dingens / Jason Gorman / Mallika Sastry / Chen-Hsiang Shen / Baoshan Zhang / Tongqing Zhou / Gwo-Yu Chuang / Cara W Chao / Ying Gu / Alexander J Jafari / Mark K Louder / Sijy O'Dell / Ariana P Rowshan / Elise G Viox / Yiran Wang / Chang W Choi / Martin M Corcoran / Angela R Corrigan / Venkata P Dandey / Edward T Eng / Hui Geng / Kathryn E Foulds / Yicheng Guo / Young D Kwon / Bob Lin / Kevin Liu / Rosemarie D Mason / Martha C Nason / Tiffany Y Ohr / Li Ou / Reda Rawi / Edward K Sarfo / Arne Schön / John P Todd / Shuishu Wang / Hui Wei / Winston Wu / / James C Mullikin / Robert T Bailer / Nicole A Doria-Rose / Gunilla B Karlsson Hedestam / Diana G Scorpio / Julie Overbaugh / Jesse D Bloom / Bridget Carragher / Clinton S Potter / Lawrence Shapiro / Peter D Kwong / John R Mascola /   Abstract: The vaccine-mediated elicitation of antibodies (Abs) capable of neutralizing diverse HIV-1 strains has been a long-standing goal. To understand how broadly neutralizing antibodies (bNAbs) can be ...The vaccine-mediated elicitation of antibodies (Abs) capable of neutralizing diverse HIV-1 strains has been a long-standing goal. To understand how broadly neutralizing antibodies (bNAbs) can be elicited, we identified, characterized, and tracked five neutralizing Ab lineages targeting the HIV-1-fusion peptide (FP) in vaccinated macaques over time. Genetic and structural analyses revealed two of these lineages to belong to a reproducible class capable of neutralizing up to 59% of 208 diverse viral strains. B cell analysis indicated each of the five lineages to have been initiated and expanded by FP-carrier priming, with envelope (Env)-trimer boosts inducing cross-reactive neutralization. These Abs had binding-energy hotspots focused on FP, whereas several FP-directed Abs induced by immunization with Env trimer-only were less FP-focused and less broadly neutralizing. Priming with a conserved subregion, such as FP, can thus induce Abs with binding-energy hotspots coincident with the target subregion and capable of broad neutralization. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8977.map.gz emd_8977.map.gz | 14.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8977-v30.xml emd-8977-v30.xml emd-8977.xml emd-8977.xml | 27.3 KB 27.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_8977_fsc.xml emd_8977_fsc.xml | 13.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_8977.png emd_8977.png | 154 KB | ||
| Others |  emd_8977_additional.map.gz emd_8977_additional.map.gz | 14 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8977 http://ftp.pdbj.org/pub/emdb/structures/EMD-8977 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8977 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8977 | HTTPS FTP |
-Related structure data
| Related structure data |  6mpgMC  9189C  9319C  9320C  9359C  6mphC  6mqcC  6mqeC  6mqmC  6mqrC  6mqsC  6n16C  6n1vC  6n1wC  6nf2C  6osyC  6ot1C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_8977.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8977.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | primary map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
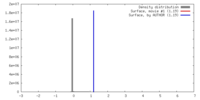
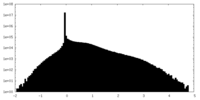
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Refined map from cryoSparc 0.6.5
| File | emd_8977_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Refined map from cryoSparc 0.6.5 | ||||||||||||
| Projections & Slices |
| ||||||||||||
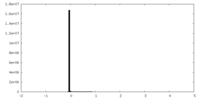
| Density Histograms |
- Sample components
Sample components
+Entire : Quaternary complex of HIV-1 Env BG505 DS-SOSIP with antibodies A1...
+Supramolecule #1: Quaternary complex of HIV-1 Env BG505 DS-SOSIP with antibodies A1...
+Supramolecule #2: HIV-1 Env BG505 DS-SOSIP gp120
+Supramolecule #3: HIV-1 Env BG505 DS-SOSIP gp41
+Supramolecule #4: A12V163-b.01
+Supramolecule #5: VRC03
+Supramolecule #6: PGT122
+Macromolecule #1: HIV-1 Env BG505 DS-SOSIP gp120
+Macromolecule #2: HIV-1 Env BG505 DS-SOSIP gp41
+Macromolecule #3: A12V163-b.01 Heavy chain
+Macromolecule #4: A12V163-b.01 Light chain
+Macromolecule #5: VRC03 heavy chain
+Macromolecule #6: VRC03 light chain
+Macromolecule #7: PGT122 heavy chain
+Macromolecule #8: PGT122 light chain
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Support film - Material: CARBON / Support film - topology: LACEY / Pretreatment - Type: GLOW DISCHARGE / Details: unspecified |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 298 K / Instrument: SPOTITON |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 69.98 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated defocus max: 3.0 µm / Calibrated defocus min: 0.8 µm / Calibrated magnification: 22500 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Target criteria: Correlation coefficient |
|---|---|
| Output model |  PDB-6mpg: |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)