+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7319 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Mtb RNAP Holo/RbpA/Fidaxomicin/upstream fork DNA | |||||||||
 Map data Map data | primary map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RNA Polymerase / Antibiotic / Inhibitor / TRANSCRIPTION / transcription-dna-antibiotic complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationbacterial-type RNA polymerase holo enzyme binding / response to water / Antimicrobial action and antimicrobial resistance in Mtb / bacterial-type RNA polymerase core enzyme binding / sigma factor activity / DNA-directed RNA polymerase complex / peptidoglycan-based cell wall / DNA-templated transcription initiation / ribonucleoside binding / DNA-directed 5'-3' RNA polymerase activity ...bacterial-type RNA polymerase holo enzyme binding / response to water / Antimicrobial action and antimicrobial resistance in Mtb / bacterial-type RNA polymerase core enzyme binding / sigma factor activity / DNA-directed RNA polymerase complex / peptidoglycan-based cell wall / DNA-templated transcription initiation / ribonucleoside binding / DNA-directed 5'-3' RNA polymerase activity / DNA-directed RNA polymerase / nucleic acid binding / protein dimerization activity / response to antibiotic / DNA-templated transcription / positive regulation of DNA-templated transcription / magnesium ion binding / DNA binding / zinc ion binding / plasma membrane / cytosol / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.38 Å | |||||||||
 Authors Authors | Darst SA / Campbell EA | |||||||||
 Citation Citation |  Journal: Elife / Year: 2018 Journal: Elife / Year: 2018Title: Fidaxomicin jams RNA polymerase motions needed for initiation via RbpA contacts. Authors: Hande Boyaci / James Chen / Mirjana Lilic / Margaret Palka / Rachel Anne Mooney / Robert Landick / Seth A Darst / Elizabeth A Campbell /  Abstract: Fidaxomicin (Fdx) is an antimicrobial RNA polymerase (RNAP) inhibitor highly effective against RNAP in vitro, but clinical use of Fdx is limited to treating intestinal infections due to poor ...Fidaxomicin (Fdx) is an antimicrobial RNA polymerase (RNAP) inhibitor highly effective against RNAP in vitro, but clinical use of Fdx is limited to treating intestinal infections due to poor absorption. To identify the structural determinants of Fdx binding to RNAP, we determined the 3.4 Å cryo-electron microscopy structure of a complete RNAP holoenzyme in complex with Fdx. We find that the actinobacteria general transcription factor RbpA contacts fidaxomycin, explaining its strong effect on . Additional structures define conformational states of RNAP between the free apo-holoenzyme and the promoter-engaged open complex ready for transcription. The results establish that Fdx acts like a doorstop to jam the enzyme in an open state, preventing the motions necessary to secure promoter DNA in the active site. Our results provide a structural platform to guide development of anti-tuberculosis antimicrobials based on the Fdx binding pocket. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7319.map.gz emd_7319.map.gz | 96.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7319-v30.xml emd-7319-v30.xml emd-7319.xml emd-7319.xml | 22.3 KB 22.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7319.png emd_7319.png | 130.1 KB | ||
| Filedesc metadata |  emd-7319.cif.gz emd-7319.cif.gz | 8.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7319 http://ftp.pdbj.org/pub/emdb/structures/EMD-7319 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7319 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7319 | HTTPS FTP |
-Validation report
| Summary document |  emd_7319_validation.pdf.gz emd_7319_validation.pdf.gz | 551.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_7319_full_validation.pdf.gz emd_7319_full_validation.pdf.gz | 550.7 KB | Display | |
| Data in XML |  emd_7319_validation.xml.gz emd_7319_validation.xml.gz | 6.3 KB | Display | |
| Data in CIF |  emd_7319_validation.cif.gz emd_7319_validation.cif.gz | 7.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7319 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7319 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7319 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7319 | HTTPS FTP |
-Related structure data
| Related structure data |  6bzoMC  7320C  7322C  7323C  6c04C  6c05C  6c06C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10887 (Title: CryoEM of Mycobacterium tuberculosis WT RNAP holoenzyme/RbpA/Fdx EMPIAR-10887 (Title: CryoEM of Mycobacterium tuberculosis WT RNAP holoenzyme/RbpA/FdxData size: 220.5 Data #1: Unaligned multi frame micrographs of M. tuberculosis RNAP/RbpA holoenzyme with Fdx [micrographs - multiframe])  EMPIAR-10896 (Title: CryoEM of Mycobacterium tuberculosis WT RNAP holoenzyme/RbpA EMPIAR-10896 (Title: CryoEM of Mycobacterium tuberculosis WT RNAP holoenzyme/RbpAData size: 838.9 Data #1: Unaligned multi-frame micrographs of M.tuberculosis RNAP holoenzyme with RbpA [micrographs - multiframe])  EMPIAR-10901 (Title: CryoEM of Mycobacterium tuberculosis WT RNAP holoenzyme/RbpA bound to the Fdx and upstream fork DNA EMPIAR-10901 (Title: CryoEM of Mycobacterium tuberculosis WT RNAP holoenzyme/RbpA bound to the Fdx and upstream fork DNAData size: 2.0 TB Data #1: Unaligned multiframe micrographs of M.tuberculosis RNAP holoenzyme with RbpA and inhibitor Fdx and upstream fork DNA [micrographs - multiframe] Data #2: Unaligned multiframe micrographs of M.tuberculosis RNAP holoenzyme with RbpA and inhibitor Fdx and upstream fork DNA [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_7319.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7319.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | primary map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
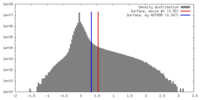
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Complex of Mtb RNAP with an antibiotic called fidaxomicin
+Supramolecule #1: Complex of Mtb RNAP with an antibiotic called fidaxomicin
+Macromolecule #1: DNA-directed RNA polymerase subunit alpha
+Macromolecule #2: DNA-directed RNA polymerase subunit beta
+Macromolecule #3: DNA-directed RNA polymerase subunit beta'
+Macromolecule #4: DNA-directed RNA polymerase subunit omega
+Macromolecule #5: RNA polymerase sigma factor SigA
+Macromolecule #6: RNA polymerase-binding protein RbpA
+Macromolecule #7: DNA (32-MER)
+Macromolecule #8: DNA (26-MER)
+Macromolecule #9: Fidaxomicin
+Macromolecule #10: ZINC ION
+Macromolecule #11: MAGNESIUM ION
+Macromolecule #12: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Grid | Model: C-flat / Material: GOLD / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 20 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 6.7 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.38 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 173509 |
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: PROJECTION MATCHING |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)