+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6746 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of PCV2 VLPs | |||||||||
 Map data Map data | Cryo-EM structure of full length PCV2 VLPs | |||||||||
 Sample Sample | Capsid protein != Porcine circovirus 2 Capsid protein
| |||||||||
| Function / homology | Circovirus capsid protein / Circovirus capsid superfamily / Circovirus capsid protein / viral capsid assembly / T=1 icosahedral viral capsid / viral penetration into host nucleus / symbiont entry into host cell / virion attachment to host cell / Capsid protein Function and homology information Function and homology information | |||||||||
| Biological species |   Porcine circovirus 2 Porcine circovirus 2 | |||||||||
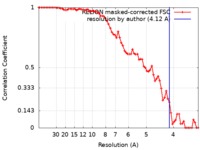
| Method | single particle reconstruction / cryo EM / Resolution: 4.12 Å | |||||||||
 Authors Authors | Mo X / Yuan AY | |||||||||
 Citation Citation |  Journal: PLoS Pathog / Year: 2019 Journal: PLoS Pathog / Year: 2019Title: Structural roles of PCV2 capsid protein N-terminus in PCV2 particle assembly and identification of PCV2 type-specific neutralizing epitope. Authors: Xiaobing Mo / Xiangdong Li / Bo Yin / Junhua Deng / Kegong Tian / Adam Yuan /   Abstract: Postweaning multisystemic wasting disease (PMWS) in piglets caused by porcine circovirus type 2 (PCV2) is one of the major threats to most pig farms worldwide. Among all the PCV types, PCV2 is the ...Postweaning multisystemic wasting disease (PMWS) in piglets caused by porcine circovirus type 2 (PCV2) is one of the major threats to most pig farms worldwide. Among all the PCV types, PCV2 is the dominant genotype causing PMWS and associated diseases. Considerable efforts were made to study the virus-like-particle (VLP) assembly and the specific PCV2-associated epitope(s) in order to establish the solid foundation for engineered PCV2 vaccine development. Although the N-terminal fragment including Nuclear Localization Signal (NLS) sequence seems important for recombinant PCV2 capsid protein expression and VLP assembly, the detailed structural and functional information regarding this important fragment are largely unknown. In this study, we report crystal structure of PCV2 VLP assembled from N-terminal NLS truncated PCV2 capsid protein at 2.8 Å resolution and cryo-EM structure of PCV2 VLP assembled from full-length PCV2 capsid protein at 4.1Å resolution. Our in vitro PCV2 VLP assembly results show that NLS-truncated PCV2 capsid protein only forms instable VLPs which were easily disassembled in solution, whereas full-length PCV2 capsid protein forms stable VLPs due to interaction between 15PRSHLGQILRRRP27 (α-helix) and 33RHRYRWRRKN42 (NLS-B) in a repeated manner. In addition, our results also showed that N-terminal truncation of PCV2 capsid protein up to 27 residues still forms PCV2 particles in solution with similar size and immunogenicity, while N-terminal truncation of PCV2 capsid protein with more than 30 residues is not able to form stable PCV2 particles in solution, demonstrating the importance of interaction between the α-helix at N-terminal and NLS-B in PCV2 VLP formation. Moreover, we also report the cryo-EM structure of PCV2 VLP in complex with 3H11-Fab, a PCV2 type-specific neutralizing antibody, at 15 Å resolution. MAb-3H11 specifically recognizes one exposed epitope located on the VLP surface EF-loop (residues 128-143), which is further confirmed by PCV1-PCV2 epitope swapping assay. Hence, our results have revealed the structural roles of N-terminal fragment of PCV2 capsid protein in PCV2 particle assembly and pinpointed one PCV2 type-specific neutralizing epitope for the first time, which could provide clear clue for next generation PCV2 vaccine and diagnostic kits development. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6746.map.gz emd_6746.map.gz | 27.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6746-v30.xml emd-6746-v30.xml emd-6746.xml emd-6746.xml | 12.6 KB 12.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_6746_fsc.xml emd_6746_fsc.xml | 7.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_6746.png emd_6746.png | 307.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6746 http://ftp.pdbj.org/pub/emdb/structures/EMD-6746 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6746 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6746 | HTTPS FTP |
-Validation report
| Summary document |  emd_6746_validation.pdf.gz emd_6746_validation.pdf.gz | 369.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_6746_full_validation.pdf.gz emd_6746_full_validation.pdf.gz | 369.3 KB | Display | |
| Data in XML |  emd_6746_validation.xml.gz emd_6746_validation.xml.gz | 9.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6746 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6746 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6746 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6746 | HTTPS FTP |
-Related structure data
| Related structure data |  5zboMC  6961C  5zjuC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6746.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6746.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of full length PCV2 VLPs | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.69 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
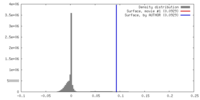
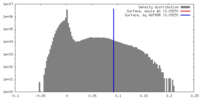
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Capsid protein
| Entire | Name: Capsid protein |
|---|---|
| Components |
|
-Supramolecule #1: Porcine circovirus 2
| Supramolecule | Name: Porcine circovirus 2 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: #1 / Details: Porcine circovirus 2 capsid protein / NCBI-ID: 85708 / Sci species name: Porcine circovirus 2 / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: Yes |
|---|---|
| Host system | Organism: |
| Molecular weight | Experimental: 1.7 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7 Component:
Details: 20mM Tris(pH 7.0), 500mM NaCl | |||||||||
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 400 / Support film - Material: FORMVAR / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 101.325 kPa | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 298 K / Instrument: FEI VITROBOT MARK IV / Details: blot for 5 seconds before pluning. | |||||||||
| Details | This sample was monodisperse |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN |
|---|---|
| Temperature | Min: 70.0 K / Max: 70.0 K |
| Details | Preliminary grid screening was performed manually. |
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Number grids imaged: 8 / Number real images: 151 / Average exposure time: 1.0 sec. / Average electron dose: 25.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated defocus max: 3.0 µm / Calibrated defocus min: 1.0 µm / Calibrated magnification: 47000 / Illumination mode: OTHER / Imaging mode: DARK FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 47000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: RECIPROCAL / Protocol: AB INITIO MODEL / Overall B value: 200 |
|---|---|
| Output model |  PDB-5zbo: |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)