[English] 日本語
 Yorodumi
Yorodumi- EMDB-6456: Cryo-electron microscopy reconstruction of the P. falciparum 80S ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6456 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-electron microscopy reconstruction of the P. falciparum 80S ribosome bound to P-tRNA | |||||||||
 Map data Map data | Cryo-EM reconstruction of the P. falciparum schizont-stage ribosome in a non-rotated state bound to P-tRNA | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cryo-EM / malaria / translational pausing / RACK1 | |||||||||
| Function / homology |  Function and homology information Function and homology informationRMTs methylate histone arginines / : / Protein methylation / Translesion synthesis by REV1 / : / Translesion Synthesis by POLH / Translesion synthesis by POLK / Translesion synthesis by POLI / Josephin domain DUBs / Metalloprotease DUBs ...RMTs methylate histone arginines / : / Protein methylation / Translesion synthesis by REV1 / : / Translesion Synthesis by POLH / Translesion synthesis by POLK / Translesion synthesis by POLI / Josephin domain DUBs / Metalloprotease DUBs / DNA Damage Recognition in GG-NER / Formation of Incision Complex in GG-NER / : / Formation of TC-NER Pre-Incision Complex / Dual incision in TC-NER / Gap-filling DNA repair synthesis and ligation in TC-NER / PTK6 Regulates RTKs and Their Effectors AKT1 and DOK1 / ER Quality Control Compartment (ERQC) / Iron uptake and transport / L13a-mediated translational silencing of Ceruloplasmin expression / SRP-dependent cotranslational protein targeting to membrane / : / Formation of a pool of free 40S subunits / Formation of the ternary complex, and subsequently, the 43S complex / Ribosomal scanning and start codon recognition / GTP hydrolysis and joining of the 60S ribosomal subunit / : / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / : / Synthesis of active ubiquitin: roles of E1 and E2 enzymes / Aggrephagy / Orc1 removal from chromatin / CDK-mediated phosphorylation and removal of Cdc6 / FBXL7 down-regulates AURKA during mitotic entry and in early mitosis / KEAP1-NFE2L2 pathway / UCH proteinases / Ub-specific processing proteases / Neddylation / Antigen processing: Ubiquitination & Proteasome degradation / MAPK6/MAPK4 signaling / ABC-family protein mediated transport / AUF1 (hnRNP D0) binds and destabilizes mRNA / preribosome / endonucleolytic cleavage to generate mature 3'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / protein-RNA complex assembly / maturation of LSU-rRNA / endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / cytosolic ribosome / ribosomal large subunit biogenesis / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of SSU-rRNA / small-subunit processome / maintenance of translational fidelity / modification-dependent protein catabolic process / mRNA 5'-UTR binding / protein tag activity / rRNA processing / large ribosomal subunit / ribosomal small subunit assembly / ribosome biogenesis / ribosomal small subunit biogenesis / 5S rRNA binding / small ribosomal subunit / ribosomal large subunit assembly / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / ubiquitin-dependent protein catabolic process / cytoplasmic translation / negative regulation of translation / rRNA binding / structural constituent of ribosome / protein ubiquitination / ribosome / translation / ribonucleoprotein complex / mRNA binding / ubiquitin protein ligase binding / nucleolus / mitochondrion / RNA binding / zinc ion binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 6.7 Å | |||||||||
 Authors Authors | Sun M / Li W / Blomqvist K / Das S / Hashem Y / Dvorin JD / Frank J | |||||||||
 Citation Citation |  Journal: Elife / Year: 2014 Journal: Elife / Year: 2014Title: Cryo-EM structure of the Plasmodium falciparum 80S ribosome bound to the anti-protozoan drug emetine. Authors: Wilson Wong / Xiao-chen Bai / Alan Brown / Israel S Fernandez / Eric Hanssen / Melanie Condron / Yan Hong Tan / Jake Baum / Sjors H W Scheres /   Abstract: Malaria inflicts an enormous burden on global human health. The emergence of parasite resistance to front-line drugs has prompted a renewed focus on the repositioning of clinically approved drugs as ...Malaria inflicts an enormous burden on global human health. The emergence of parasite resistance to front-line drugs has prompted a renewed focus on the repositioning of clinically approved drugs as potential anti-malarial therapies. Antibiotics that inhibit protein translation are promising candidates for repositioning. We have solved the cryo-EM structure of the cytoplasmic ribosome from the human malaria parasite, Plasmodium falciparum, in complex with emetine at 3.2 Å resolution. Emetine is an anti-protozoan drug used in the treatment of ameobiasis that also displays potent anti-malarial activity. Emetine interacts with the E-site of the ribosomal small subunit and shares a similar binding site with the antibiotic pactamycin, thereby delivering its therapeutic effect by blocking mRNA/tRNA translocation. As the first cryo-EM structure that visualizes an antibiotic bound to any ribosome at atomic resolution, this establishes cryo-EM as a powerful tool for screening and guiding the design of drugs that target parasite translation machinery. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6456.map.gz emd_6456.map.gz | 48 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6456-v30.xml emd-6456-v30.xml emd-6456.xml emd-6456.xml | 10.9 KB 10.9 KB | Display Display |  EMDB header EMDB header |
| Images |  400_6456.gif 400_6456.gif 80_6456.gif 80_6456.gif | 59.1 KB 4.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6456 http://ftp.pdbj.org/pub/emdb/structures/EMD-6456 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6456 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6456 | HTTPS FTP |
-Related structure data
| Related structure data |  3jbnMC  6452C  6454C  3jboC  3jbpC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6456.map.gz / Format: CCP4 / Size: 96.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6456.map.gz / Format: CCP4 / Size: 96.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM reconstruction of the P. falciparum schizont-stage ribosome in a non-rotated state bound to P-tRNA | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.66 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
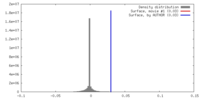
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : The schizont-stage Plasmodium falciparum 80S ribosome bound to P-tRNA
| Entire | Name: The schizont-stage Plasmodium falciparum 80S ribosome bound to P-tRNA |
|---|---|
| Components |
|
-Supramolecule #1000: The schizont-stage Plasmodium falciparum 80S ribosome bound to P-tRNA
| Supramolecule | Name: The schizont-stage Plasmodium falciparum 80S ribosome bound to P-tRNA type: sample / ID: 1000 / Number unique components: 1 |
|---|
-Supramolecule #1: 80S ribosome
| Supramolecule | Name: 80S ribosome / type: complex / ID: 1 / Recombinant expression: No / Database: NCBI / Ribosome-details: ribosome-eukaryote: ALL |
|---|---|
| Source (natural) | Organism:  Strain: 3D7 / synonym: Plasmodium |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Details: 10 mM HEPES potassium, 50 mM KOAc, 10 mM NH4Cl, 2 mM DTT, 5 mM Mg(OAc)2 |
|---|---|
| Grid | Details: 300 mesh Copper/Molbydenum holey carton-coated Quantifoil R2/4 grid, containing an additional continuous thin layer of carbon |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV / Method: Blot for 4 seconds before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Date | Oct 13, 2013 |
| Image recording | Category: CCD / Film or detector model: GATAN K2 (4k x 4k) / Number real images: 5734 / Average electron dose: 25 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 30120 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.26 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 23000 |
| Sample stage | Specimen holder model: GATAN HELIUM |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | See 'Materials and Methods' in Sun, M., et al 2015 |
|---|---|
| CTF correction | Details: Each micrograph |
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 6.7 Å / Resolution method: OTHER / Software - Name: Arachnid, CTFFIND3, RELION / Number images used: 14696 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name: MDFF |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
| Output model |  PDB-3jbn: |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)