+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5934 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Dengue serotype 3 virus at 37 degrees C | |||||||||
 Map data Map data | Cryo-EM reconstruction of Dengue virus 3 at 37 degrees C | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | dengue virus | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated suppression of host JAK-STAT cascade via inhibition of host TYK2 activity / flavivirin / host cell mitochondrion / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT2 activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MAVS activity / protein complex oligomerization / ribonucleoside triphosphate phosphatase activity / monoatomic ion channel activity / channel activity / double-stranded RNA binding ...symbiont-mediated suppression of host JAK-STAT cascade via inhibition of host TYK2 activity / flavivirin / host cell mitochondrion / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT2 activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MAVS activity / protein complex oligomerization / ribonucleoside triphosphate phosphatase activity / monoatomic ion channel activity / channel activity / double-stranded RNA binding / nucleoside-triphosphate phosphatase / viral capsid / monoatomic ion transmembrane transport / mRNA (guanine-N7)-methyltransferase / methyltransferase cap1 / clathrin-dependent endocytosis of virus by host cell / mRNA (nucleoside-2'-O-)-methyltransferase activity / mRNA 5'-cap (guanine-N7-)-methyltransferase activity / RNA helicase activity / host cell perinuclear region of cytoplasm / protein dimerization activity / host cell endoplasmic reticulum membrane / RNA helicase / symbiont-mediated suppression of host type I interferon-mediated signaling pathway / induction by virus of host autophagy / RNA-directed RNA polymerase / viral RNA genome replication / serine-type endopeptidase activity / RNA-dependent RNA polymerase activity / virus-mediated perturbation of host defense response / fusion of virus membrane with host endosome membrane / viral envelope / host cell nucleus / virion attachment to host cell / structural molecule activity / virion membrane / ATP hydrolysis activity / proteolysis / extracellular region / ATP binding / membrane / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Dengue virus 3 Dengue virus 3 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 7.0 Å | |||||||||
 Authors Authors | Fibriansah G / Tan JL / Smith SA / de Alwis R / Ng T-S / Kostyuchenko VA / Kukkaro P / de Silva AM / Crowe Jr JE / Lok S-M | |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2015 Journal: Nat Commun / Year: 2015Title: A highly potent human antibody neutralizes dengue virus serotype 3 by binding across three surface proteins. Authors: Guntur Fibriansah / Joanne L Tan / Scott A Smith / Ruklanthi de Alwis / Thiam-Seng Ng / Victor A Kostyuchenko / Ramesh S Jadi / Petra Kukkaro / Aravinda M de Silva / James E Crowe / Shee-Mei Lok /   Abstract: Dengue virus (DENV) infects ~400 million people annually. There is no licensed vaccine or therapeutic drug. Only a small fraction of the total DENV-specific antibodies in a naturally occurring dengue ...Dengue virus (DENV) infects ~400 million people annually. There is no licensed vaccine or therapeutic drug. Only a small fraction of the total DENV-specific antibodies in a naturally occurring dengue infection consists of highly neutralizing antibodies. Here we show that the DENV-specific human monoclonal antibody 5J7 is exceptionally potent, neutralizing 50% of virus at nanogram-range antibody concentration. The 9 Å resolution cryo-electron microscopy structure of the Fab 5J7-DENV complex shows that a single Fab molecule binds across three envelope proteins and engages three functionally important domains, each from a different envelope protein. These domains are critical for receptor binding and fusion to the endosomal membrane. The ability to bind to multiple domains allows the antibody to fully coat the virus surface with only 60 copies of Fab, that is, half the amount compared with other potent antibodies. Our study reveals a highly efficient and unusual mechanism of molecular recognition by an antibody. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5934.map.gz emd_5934.map.gz | 99.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5934-v30.xml emd-5934-v30.xml emd-5934.xml emd-5934.xml | 11.3 KB 11.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5934.png emd_5934.png | 379.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5934 http://ftp.pdbj.org/pub/emdb/structures/EMD-5934 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5934 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5934 | HTTPS FTP |
-Validation report
| Summary document |  emd_5934_validation.pdf.gz emd_5934_validation.pdf.gz | 368.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_5934_full_validation.pdf.gz emd_5934_full_validation.pdf.gz | 367.8 KB | Display | |
| Data in XML |  emd_5934_validation.xml.gz emd_5934_validation.xml.gz | 8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5934 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5934 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5934 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5934 | HTTPS FTP |
-Related structure data
| Related structure data |  3j6tMC  5933C  5935C  3j6sC  3j6uC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_5934.map.gz / Format: CCP4 / Size: 500 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5934.map.gz / Format: CCP4 / Size: 500 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM reconstruction of Dengue virus 3 at 37 degrees C | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.7 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
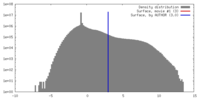
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Dengue virus serotype 3
| Entire | Name: Dengue virus serotype 3 |
|---|---|
| Components |
|
-Supramolecule #1000: Dengue virus serotype 3
| Supramolecule | Name: Dengue virus serotype 3 / type: sample / ID: 1000 Details: The sample was incubated at 37 degrees C for 30 min followed by another 2 h at 4 degrees C prior to freezing. Number unique components: 1 |
|---|
-Supramolecule #1: Dengue virus 3
| Supramolecule | Name: Dengue virus 3 / type: virus / ID: 1 / NCBI-ID: 11069 / Sci species name: Dengue virus 3 / Sci species strain: D3/SG/05K863DK1/2005 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) / synonym: VERTEBRATES Homo sapiens (human) / synonym: VERTEBRATES |
| Host system | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 / Details: 10 mM Tris-HCl, pH 8.0, 120 mM NaCl, 1 mM EDTA |
|---|---|
| Grid | Details: ultra-thin carbon-coated lacey carbon grid |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 100 K / Instrument: FEI VITROBOT MARK IV Method: Blotted with filter paper for 2 seconds prior to snap freezing |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Average: 100 K |
| Date | Mar 28, 2012 |
| Image recording | Category: CCD / Film or detector model: FEI FALCON I (4k x 4k) / Digitization - Sampling interval: 14 µm / Number real images: 138 / Average electron dose: 17.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.2 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 47000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | The particles were manually selected. |
|---|---|
| CTF correction | Details: each particle |
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 7.0 Å / Resolution method: OTHER / Software - Name: MPSA, EMAN, EMAN2 / Number images used: 3009 |
-Atomic model buiding 1
| Initial model | PDB ID: Chain - #0 - Chain ID: A / Chain - #1 - Chain ID: B / Chain - #2 - Chain ID: C / Chain - #3 - Chain ID: D / Chain - #4 - Chain ID: E / Chain - #5 - Chain ID: F |
|---|---|
| Software | Name: Chimera, NAMD/MDFF |
| Details | Initially fitted in Chimera and refined using NAMD/MDFF |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Target criteria: real space correlation |
| Output model |  PDB-3j6t: |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)