[English] 日本語
 Yorodumi
Yorodumi- EMDB-4868: Structure of the complete Vaccinia DNA-dependent RNA polymerase c... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4868 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the complete Vaccinia DNA-dependent RNA polymerase complex | |||||||||
 Map data Map data | Cryo-EM reconstruction map of the complete Vaccinia DNA-dependent RNA polymerase complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Vaccinia / RNA polymerase / RNA Polymerase complex / RNAP / vRNAP / complete vRNAP / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationpolynucleotide 5'-phosphatase / virion component => GO:0044423 / polynucleotide 5'-phosphatase activity / viral transcription / nucleotidyltransferase activity / ribonucleoside triphosphate phosphatase activity / DNA-templated transcription termination / ribonucleoside binding / DNA-directed 5'-3' RNA polymerase activity / DNA-directed RNA polymerase ...polynucleotide 5'-phosphatase / virion component => GO:0044423 / polynucleotide 5'-phosphatase activity / viral transcription / nucleotidyltransferase activity / ribonucleoside triphosphate phosphatase activity / DNA-templated transcription termination / ribonucleoside binding / DNA-directed 5'-3' RNA polymerase activity / DNA-directed RNA polymerase / nucleoside-triphosphate phosphatase / mRNA guanylyltransferase activity / mRNA guanylyltransferase / mRNA (guanine-N7)-methyltransferase / mRNA 5'-cap (guanine-N7-)-methyltransferase activity / DNA-binding transcription factor activity / DNA-templated transcription / positive regulation of DNA-templated transcription / DNA binding / zinc ion binding / ATP binding Similarity search - Function | |||||||||
| Biological species |  Vaccinia virus GLV-1H68 / Vaccinia virus GLV-1H68 /  Homo sapiens (human) / Homo sapiens (human) /  Vaccinia virus GLV-1h68 Vaccinia virus GLV-1h68 | |||||||||
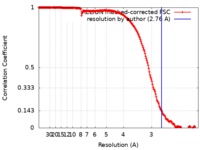
| Method | single particle reconstruction / cryo EM / Resolution: 2.76 Å | |||||||||
 Authors Authors | Grimm C / Hillen SH | |||||||||
| Funding support |  Germany, 2 items Germany, 2 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2019 Journal: Cell / Year: 2019Title: Structural Basis of Poxvirus Transcription: Vaccinia RNA Polymerase Complexes. Authors: Clemens Grimm / Hauke S Hillen / Kristina Bedenk / Julia Bartuli / Simon Neyer / Qian Zhang / Alexander Hüttenhofer / Matthias Erlacher / Christian Dienemann / Andreas Schlosser / Henning ...Authors: Clemens Grimm / Hauke S Hillen / Kristina Bedenk / Julia Bartuli / Simon Neyer / Qian Zhang / Alexander Hüttenhofer / Matthias Erlacher / Christian Dienemann / Andreas Schlosser / Henning Urlaub / Bettina Böttcher / Aladar A Szalay / Patrick Cramer / Utz Fischer /    Abstract: Poxviruses encode a multisubunit DNA-dependent RNA polymerase (vRNAP) that carries out viral gene expression in the host cytoplasm. We report cryo-EM structures of core and complete vRNAP enzymes ...Poxviruses encode a multisubunit DNA-dependent RNA polymerase (vRNAP) that carries out viral gene expression in the host cytoplasm. We report cryo-EM structures of core and complete vRNAP enzymes from Vaccinia virus at 2.8 Å resolution. The vRNAP core enzyme resembles eukaryotic RNA polymerase II (Pol II) but also reveals many virus-specific features, including the transcription factor Rap94. The complete enzyme additionally contains the transcription factor VETF, the mRNA processing factors VTF/CE and NPH-I, the viral core protein E11, and host tRNA. This complex can carry out the entire early transcription cycle. The structures show that Rap94 partially resembles the Pol II initiation factor TFIIB, that the vRNAP subunit Rpo30 resembles the Pol II elongation factor TFIIS, and that NPH-I resembles chromatin remodeling enzymes. Together with the accompanying paper (Hillen et al., 2019), these results provide the basis for unraveling the mechanisms of poxvirus transcription and RNA processing. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4868.map.gz emd_4868.map.gz | 30.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4868-v30.xml emd-4868-v30.xml emd-4868.xml emd-4868.xml | 35.1 KB 35.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4868_fsc.xml emd_4868_fsc.xml | 17.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_4868.png emd_4868.png | 67.6 KB | ||
| Filedesc metadata |  emd-4868.cif.gz emd-4868.cif.gz | 11.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4868 http://ftp.pdbj.org/pub/emdb/structures/EMD-4868 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4868 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4868 | HTTPS FTP |
-Validation report
| Summary document |  emd_4868_validation.pdf.gz emd_4868_validation.pdf.gz | 384.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_4868_full_validation.pdf.gz emd_4868_full_validation.pdf.gz | 383.9 KB | Display | |
| Data in XML |  emd_4868_validation.xml.gz emd_4868_validation.xml.gz | 16.2 KB | Display | |
| Data in CIF |  emd_4868_validation.cif.gz emd_4868_validation.cif.gz | 22.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4868 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4868 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4868 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4868 | HTTPS FTP |
-Related structure data
| Related structure data |  6rflMC  6rfgC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4868.map.gz / Format: CCP4 / Size: 488.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4868.map.gz / Format: CCP4 / Size: 488.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM reconstruction map of the complete Vaccinia DNA-dependent RNA polymerase complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0635 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Complete Vaccinia DNA-dependent RNA polymerase complex
+Supramolecule #1: Complete Vaccinia DNA-dependent RNA polymerase complex
+Supramolecule #2: Vaccinia DNA-dependent RNA polymerase complex
+Supramolecule #3: RNA
+Macromolecule #1: DNA-dependent RNA polymerase subunit rpo132
+Macromolecule #2: DNA-directed RNA polymerase subunit
+Macromolecule #3: DNA-directed RNA polymerase 19 kDa subunit
+Macromolecule #4: DNA-dependent RNA polymerase subunit rpo18
+Macromolecule #5: Putative H4L RNA polymerase-associated transcription factor RAP94
+Macromolecule #6: DNA-dependent RNA polymerase subunit rpo7
+Macromolecule #7: Small subunit of mRNA capping enzyme
+Macromolecule #8: Large subunit of mRNA capping enzyme
+Macromolecule #9: Virion core protein
+Macromolecule #10: Transcription factor VETF 82kDa large subunit
+Macromolecule #12: DNA-dependent RNA polymerase subunit rpo147
+Macromolecule #13: Nucleoside triphosphate phosphohydrolase-I
+Macromolecule #14: DNA-directed RNA polymerase 35 kDa subunit
+Macromolecule #15: DNA-directed RNA polymerase 30 kDa polypeptide
+Macromolecule #11: chr17.trna16-GlnTTG
+Macromolecule #16: ZINC ION
+Macromolecule #17: MAGNESIUM ION
+Macromolecule #18: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)