[English] 日本語
 Yorodumi
Yorodumi- EMDB-4543: Structure of eIF2B-eIF2 (phosphorylated at Ser51) complex (map 1) -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4543 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of eIF2B-eIF2 (phosphorylated at Ser51) complex (map 1) | ||||||||||||
 Map data Map data | For optimal visualization of all eIF2 alpha and gamma, gauss-filter the map by 1.34 and display it at 0.022 contour level | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | integrated stress response / ISR / translation / initiation factors / phosphorylation / eIF2 / eIF2B / tRNAi / GEF / heat domain / eIF2 alpha | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of cellular response to amino acid starvation / positive regulation of cellular response to heat / Recycling of eIF2:GDP / Cellular response to mitochondrial stress / eukaryotic translation initiation factor 2B complex / ABC-family proteins mediated transport / methionyl-initiator methionine tRNA binding / eukaryotic translation initiation factor 2 complex / multi-eIF complex / cytoplasmic translational initiation ...negative regulation of cellular response to amino acid starvation / positive regulation of cellular response to heat / Recycling of eIF2:GDP / Cellular response to mitochondrial stress / eukaryotic translation initiation factor 2B complex / ABC-family proteins mediated transport / methionyl-initiator methionine tRNA binding / eukaryotic translation initiation factor 2 complex / multi-eIF complex / cytoplasmic translational initiation / protein-synthesizing GTPase / eukaryotic 43S preinitiation complex / guanyl-nucleotide exchange factor complex / formation of translation preinitiation complex / positive regulation of cellular response to amino acid starvation / eukaryotic 48S preinitiation complex / positive regulation of translational fidelity / regulation of translational initiation / Formation of the ternary complex, and subsequently, the 43S complex / Translation initiation complex formation / Ribosomal scanning and start codon recognition / L13a-mediated translational silencing of Ceruloplasmin expression / positive regulation of translational initiation / enzyme regulator activity / translation initiation factor binding / translation initiation factor activity / guanyl-nucleotide exchange factor activity / translational initiation / cytoplasmic stress granule / ribosome binding / ribosome / GTPase activity / mRNA binding / GTP binding / mitochondrion / RNA binding / metal ion binding / cytosol / cytoplasm Similarity search - Function | ||||||||||||
| Biological species |   | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.15 Å | ||||||||||||
 Authors Authors | Llacer JL / Gordiyenko Y / Ramakrishnan V | ||||||||||||
| Funding support |  United Kingdom, United Kingdom,  Spain, 3 items Spain, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2019 Journal: Nat Commun / Year: 2019Title: Structural basis for the inhibition of translation through eIF2α phosphorylation. Authors: Yuliya Gordiyenko / José Luis Llácer / V Ramakrishnan /   Abstract: One of the responses to stress by eukaryotic cells is the down-regulation of protein synthesis by phosphorylation of translation initiation factor eIF2. Phosphorylation results in low availability of ...One of the responses to stress by eukaryotic cells is the down-regulation of protein synthesis by phosphorylation of translation initiation factor eIF2. Phosphorylation results in low availability of the eIF2 ternary complex (eIF2-GTP-tRNAi) by affecting the interaction of eIF2 with its GTP-GDP exchange factor eIF2B. We have determined the cryo-EM structure of yeast eIF2B in complex with phosphorylated eIF2 at an overall resolution of 4.2 Å. Two eIF2 molecules bind opposite sides of an eIF2B hetero-decamer through eIF2α-D1, which contains the phosphorylated Ser51. eIF2α-D1 is mainly inserted between the N-terminal helix bundle domains of δ and α subunits of eIF2B. Phosphorylation of Ser51 enhances binding to eIF2B through direct interactions of phosphate groups with residues in eIF2Bα and indirectly by inducing contacts of eIF2α helix 58-63 with eIF2Bδ leading to a competition with Met-tRNA. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4543.map.gz emd_4543.map.gz | 77.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4543-v30.xml emd-4543-v30.xml emd-4543.xml emd-4543.xml | 29.9 KB 29.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_4543.png emd_4543.png | 149.4 KB | ||
| Filedesc metadata |  emd-4543.cif.gz emd-4543.cif.gz | 9 KB | ||
| Others |  emd_4543_half_map_1.map.gz emd_4543_half_map_1.map.gz emd_4543_half_map_2.map.gz emd_4543_half_map_2.map.gz | 64 MB 64 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4543 http://ftp.pdbj.org/pub/emdb/structures/EMD-4543 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4543 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4543 | HTTPS FTP |
-Validation report
| Summary document |  emd_4543_validation.pdf.gz emd_4543_validation.pdf.gz | 844.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_4543_full_validation.pdf.gz emd_4543_full_validation.pdf.gz | 844.1 KB | Display | |
| Data in XML |  emd_4543_validation.xml.gz emd_4543_validation.xml.gz | 12.5 KB | Display | |
| Data in CIF |  emd_4543_validation.cif.gz emd_4543_validation.cif.gz | 14.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4543 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4543 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4543 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4543 | HTTPS FTP |
-Related structure data
| Related structure data |  6qg0MC  4544C  4545C  4546C  4547C  4548C  6qg1C  6qg2C  6qg3C  6qg5C  6qg6C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4543.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4543.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | For optimal visualization of all eIF2 alpha and gamma, gauss-filter the map by 1.34 and display it at 0.022 contour level | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.34 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: #2
| File | emd_4543_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
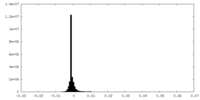
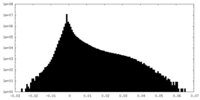
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_4543_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
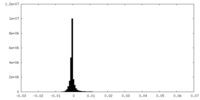
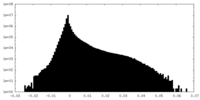
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Structure of eIF2B-eIF2 (phosphorylated at Ser51) complex (model 1)
| Entire | Name: Structure of eIF2B-eIF2 (phosphorylated at Ser51) complex (model 1) |
|---|---|
| Components |
|
-Supramolecule #1: Structure of eIF2B-eIF2 (phosphorylated at Ser51) complex (model 1)
| Supramolecule | Name: Structure of eIF2B-eIF2 (phosphorylated at Ser51) complex (model 1) type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 837 KDa |
-Macromolecule #1: Translation initiation factor eIF-2B subunit alpha
| Macromolecule | Name: Translation initiation factor eIF-2B subunit alpha / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 34.062027 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSEFNITETY LRFLEEDTEM TMPIAAIEAL VTLLRIKTPE TAAEMINTIK SSTEELIKSI PNSVSLRAGC DIFMRFVLRN LHLYGDWEN CKQHLIENGQ LFVSRAKKSR NKIAEIGVDF IADDDIILVH GYSRAVFSLL NHAANKFIRF RCVVTESRPS K QGNQLYTL ...String: MSEFNITETY LRFLEEDTEM TMPIAAIEAL VTLLRIKTPE TAAEMINTIK SSTEELIKSI PNSVSLRAGC DIFMRFVLRN LHLYGDWEN CKQHLIENGQ LFVSRAKKSR NKIAEIGVDF IADDDIILVH GYSRAVFSLL NHAANKFIRF RCVVTESRPS K QGNQLYTL LEQKGIPVTL IVDSAVGAVI DKVDKVFVGA EGVAESGGII NLVGTYSVGV LAHNARKPFY VVTESHKFVR MF PLSSDDL PMAGPPLDFT RRTDDLEDAL RGPTIDYTAQ EYITALITDL GVLTPSAVSE ELIKMWYD UniProtKB: Translation initiation factor eIF2B subunit alpha |
-Macromolecule #2: Translation initiation factor eIF-2B subunit beta
| Macromolecule | Name: Translation initiation factor eIF-2B subunit beta / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 42.621441 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSSQAFTSVH PNAATSDVNV TIDTFVAKLK RRQVQGSYAI ALETLQLLMR FISAARWNHV NDLIEQIRDL GNSLEKAHPT AFSCGNVIR RILAVLRDEV EEDTMSTTVT STSVAEPLIS SMFNLLQKPE QPHQNRKNSS GSSSMKTKTD YRQVAIQGIK D LIDEIKNI ...String: MSSQAFTSVH PNAATSDVNV TIDTFVAKLK RRQVQGSYAI ALETLQLLMR FISAARWNHV NDLIEQIRDL GNSLEKAHPT AFSCGNVIR RILAVLRDEV EEDTMSTTVT STSVAEPLIS SMFNLLQKPE QPHQNRKNSS GSSSMKTKTD YRQVAIQGIK D LIDEIKNI DEGIQQIAID LIHDHEILLT PTPDSKTVLK FLITARERSN RTFTVLVTEG FPNNTKNAHE FAKKLAQHNI ET LVVPDSA VFALMSRVGK VIIGTKAVFV NGGTISSNSG VSSVCECARE FRTPVFAVAG LYKLSPLYPF DVEKFVEFGG SQR ILPRMD PRKRLDTVNQ ITDYVPPENI DIYITNVGGF NPSFIYRIAW DNYKQIDVHL DKNKA UniProtKB: Translation initiation factor eIF2B subunit beta |
-Macromolecule #3: Translation initiation factor eIF-2B subunit gamma
| Macromolecule | Name: Translation initiation factor eIF-2B subunit gamma / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 65.76832 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSIQAFVFCG KGSNLAPFTQ PDFPFQTQNK DSTAATSGDK LNELVNSALD STVINEFMQH STRLPKALLP IGNRPMIEYV LDWCDQADF KEISVVAPVD EIELIESGLT SFLSLRKQQF ELIYKALSNS NHSHHLQDPK KINFIPSKAN STGESLQKEL L PRINGDFV ...String: MSIQAFVFCG KGSNLAPFTQ PDFPFQTQNK DSTAATSGDK LNELVNSALD STVINEFMQH STRLPKALLP IGNRPMIEYV LDWCDQADF KEISVVAPVD EIELIESGLT SFLSLRKQQF ELIYKALSNS NHSHHLQDPK KINFIPSKAN STGESLQKEL L PRINGDFV ILPCDFVTDI PPQVLVDQFR NRDDNNLAMT IYYKNSLDSS IDKKQQQKQK QQQFFTVYSE NEDSERQPIL LD VYSQRDV TKTKYLQIRS HLLWNYPNLT VSTKLLNSFI YFCSFELCQL LKLGPQSMSR QASFKDPFTG NQQQQNPPTT DDD EDRNHD DDDDYKPSAT SIQPTYFKKK NDLILDPINC NKSLSKVFRD LSRRSWQHSK PREPIGIFIL PNETLFIRAN NLNA YMDAN RFVLKIKSQT MFTKNIQIQS AAIGADAIVD PKCQISAHSN VKMSVLGTQA NIGSRCRVAG SLLFPGVHLG DEVIL ENCI IGPMAKIGSK CKLSNCYIEG HYVVEPKNNF KGETLANVYL DEDEEDELIY DDSVIAGESE IAEETDSDDR SDEDSD DSE YTDEYEYEDD GLFER UniProtKB: Translation initiation factor eIF2B subunit gamma |
-Macromolecule #4: Translation initiation factor eIF-2B subunit delta
| Macromolecule | Name: Translation initiation factor eIF-2B subunit delta / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 70.945195 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSESEAKSRS ATPPSKAKQA TPTTTAAANG EKKLTNKELK ELKKQEKAAK RAAMKQANGI SIEQQQQQAQ MKKEKKQLQR EQQQKREQK QKNANKKKQN ERNVKKSTLF GHLETTEERR ATILALTSAV SSPKTSRITA AGLMVPVVAS ALSGSNVLTA S SLMPVGPN ...String: MSESEAKSRS ATPPSKAKQA TPTTTAAANG EKKLTNKELK ELKKQEKAAK RAAMKQANGI SIEQQQQQAQ MKKEKKQLQR EQQQKREQK QKNANKKKQN ERNVKKSTLF GHLETTEERR ATILALTSAV SSPKTSRITA AGLMVPVVAS ALSGSNVLTA S SLMPVGPN ASSTVSASAP ASTTTTLPAS SAALSAGTSS ASTNTPTAIQ QEIASSNASD VAKTLASISL EAGEFNVIPG IS SVIPTVL EQSFDNSSLI SSVKELLLNK DLIHPSILLL TSHLAHYKIV GSIPRCIAML EVFQIVIKDY QTPKGTTLSR NLT SYLSHQ IDLLKKARPL SVTMGNAIRW LKQEISLIDP STPDKAAKKD LCEKIGQFAK EKIELADQLI IDNASTQIEE STTI VTYGS SKVLTELLLH NAISLKKNIK VIVVDSRPLF EGRKMAETLR NAGVNVMYAL ITSLDTIFNM DVDYVFLGAH SILSN GFLY SRAGTAMLAM SAKRRNIPVL VCCESLKFSQ RVQLDSVTFN ELADPNDLVN IDYENPVERR GNKGALLNQF IKERKF EKK KLAMENKPKG NKIGGKKGSE GESKDASNEE DSNSKNILDG WQELPSLNIV NILYDLTPPE YIKKVITEFG ALPPSSV PV ILREYKGSA UniProtKB: Translation initiation factor eIF2B subunit delta |
-Macromolecule #5: Translation initiation factor eIF-2B subunit epsilon
| Macromolecule | Name: Translation initiation factor eIF-2B subunit epsilon / type: protein_or_peptide / ID: 5 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 81.249062 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAGKKGQKKS GLGNHGKNSD MDVEDRLQAV VLTDSYETRF MPLTAVKPRC LLPLANVPLI EYTLEFLAKA GVHEVFLICS SHANQINDY IENSKWNLPW SPFKITTIMS PEARCTGDVM RDLDNRGIIT GDFILVSGDV LTNIDFSKML EFHKKMHLQD K DHISTMCL ...String: MAGKKGQKKS GLGNHGKNSD MDVEDRLQAV VLTDSYETRF MPLTAVKPRC LLPLANVPLI EYTLEFLAKA GVHEVFLICS SHANQINDY IENSKWNLPW SPFKITTIMS PEARCTGDVM RDLDNRGIIT GDFILVSGDV LTNIDFSKML EFHKKMHLQD K DHISTMCL SKASTYPKTR TIEPAAFVLD KSTSRCIYYQ DLPLPSSREK TSIQIDPELL DNVDEFVIRN DLIDCRIDIC TS HVPLIFQ ENFDYQSLRT DFVKGVISSD ILGKHIYAYL TDEYAVRVES WQTYDTISQD FLGRWCYPLV LDSNIQDDQT YSY ESRHIY KEKDVVLAQS CKIGKCTAIG SGTKIGEGTK IENSVIGRNC QIGENIRIKN SFIWDDCIIG NNSIIDHSLI ASNA TLGSN VRLNDGCIIG FNVKIDDNMD LDRNTKISAS PLKNAGSRMY DNESNEQFDQ DLDDQTLAVS IVGDKGVGYI YESEV SDDE DSSTEACKEI NTLSNQLDEL YLSDDSISSA TKKTKKRRTM SVNSIYTDRE EIDSEFEDED FEKEGIATVE RAMENN HDL DTALLELNTL RMSMNVTYHE VRIATITALL RRVYHFIATQ TLGPKDAVVK VFNQWGLLFK RQAFDEEEYI DLMNIIM EK IVEQSFDKPD LILFSALVSL YDNDIIEEDV IYKWWDNVST DPRYDEVKKL TVKWVEWLQN ADEESSSEEE UniProtKB: Translation initiation factor eIF2B subunit epsilon |
-Macromolecule #6: Eukaryotic translation initiation factor 2 subunit alpha
| Macromolecule | Name: Eukaryotic translation initiation factor 2 subunit alpha type: protein_or_peptide / ID: 6 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 34.843633 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSTSHCRFYE NKYPEIDDIV MVNVQQIAEM GAYVKLLEYD NIEGMILLSE L(SEP)RRRIRSIQ KLIRVGKNDV AVVLRV DKE KGYIDLSKRR VSSEDIIKCE EKYQKSKTVH SILRYCAEKF QIPLEELYKT IAWPLSRKFG HAYEAFKLSI IDETVWE GI EPPSKDVLDE ...String: MSTSHCRFYE NKYPEIDDIV MVNVQQIAEM GAYVKLLEYD NIEGMILLSE L(SEP)RRRIRSIQ KLIRVGKNDV AVVLRV DKE KGYIDLSKRR VSSEDIIKCE EKYQKSKTVH SILRYCAEKF QIPLEELYKT IAWPLSRKFG HAYEAFKLSI IDETVWE GI EPPSKDVLDE LKNYISKRLT PQAVKIRADV EVSCFSYEGI DAIKDALKSA EDMSTEQMQV KVKLVAAPLY VLTTQALD K QKGIEQLESA IEKITEVITK YGGVCNITMP PKAVTATEDA ELQALLESKE LDNRSDSEDD EDESDDE UniProtKB: Eukaryotic translation initiation factor 2 subunit alpha |
-Macromolecule #7: Eukaryotic translation initiation factor 2 subunit gamma
| Macromolecule | Name: Eukaryotic translation initiation factor 2 subunit gamma type: protein_or_peptide / ID: 7 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 57.942699 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSDLQDQEPS IIINGNLEPV GEPDIVEETE VVAQETQETQ DADKPKKKVA FTGLEEDGET EEEKRKREFE EGGGLPEQPL NPDFSKLNP LSAEIINRQA TINIGTIGHV AHGKSTVVRA ISGVQTVRFK DELERNITIK LGYANAKIYK CQEPTCPEPD C YRSFKSDK ...String: MSDLQDQEPS IIINGNLEPV GEPDIVEETE VVAQETQETQ DADKPKKKVA FTGLEEDGET EEEKRKREFE EGGGLPEQPL NPDFSKLNP LSAEIINRQA TINIGTIGHV AHGKSTVVRA ISGVQTVRFK DELERNITIK LGYANAKIYK CQEPTCPEPD C YRSFKSDK EISPKCQRPG CPGRYKLVRH VSFVDCPGHD ILMSTMLSGA AVMDAALLLI AGNESCPQPQ TSEHLAAIEI MK LKHVIIL QNKVDLMREE SALEHQKSIL KFIRGTIADG APIVPISAQL KYNIDAVNEF IVKTIPVPPR DFMISPRLIV IRS FDVNKP GAEIEDLKGG VAGGSILNGV FKLGDEIEIR PGIVTKDDKG KIQCKPIFSN IVSLFAEQND LKFAVPGGLI GVGT KVDPT LCRADRLVGQ VVGAKGHLPN IYTDIEINYF LLRRLLGVKT DGQKQAKVRK LEPNEVLMVN IGSTATGARV VAVKA DMAR LQLTSPACTE INEKIALSRR IEKHWRLIGW ATIKKGTTLE PIA UniProtKB: Eukaryotic translation initiation factor 2 subunit gamma |
-Macromolecule #8: Eukaryotic translation initiation factor 2 subunit beta
| Macromolecule | Name: Eukaryotic translation initiation factor 2 subunit beta type: protein_or_peptide / ID: 8 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 31.631309 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSSDLAAELG FDPALKKKKK TKKVIPDDFD AAVNGKENGS GDDLFAGLKK KKKKSKSVSA DAEAEKEPTD DIAEALGELS LKKKKKKTK DSSVDAFEKE LAKAGLDNVD AESKEGTPSA NSSIQQEVGL PYSELLSRFF NILRTNNPEL AGDRSGPKFR I PPPVCLRD ...String: MSSDLAAELG FDPALKKKKK TKKVIPDDFD AAVNGKENGS GDDLFAGLKK KKKKSKSVSA DAEAEKEPTD DIAEALGELS LKKKKKKTK DSSVDAFEKE LAKAGLDNVD AESKEGTPSA NSSIQQEVGL PYSELLSRFF NILRTNNPEL AGDRSGPKFR I PPPVCLRD GKKTIFSNIQ DIAEKLHRSP EHLIQYLFAE LGTSGSVDGQ KRLVIKGKFQ SKQMENVLRR YILEYVTCKT CK SINTELK REQSNRLFFM VCKSCGSTRS VSSIKTGFQA TVGKRRRM UniProtKB: Eukaryotic translation initiation factor 2 subunit beta |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.17 mg/mL | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||
| Grid | Model: Quantifoil, UltrAuFoil / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 15 sec. | ||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK I |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 90.0 K / Max: 100.0 K |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 2 / Number real images: 1241 / Average exposure time: 60.0 sec. / Average electron dose: 21.0 e/Å2 Details: Images were collected in movie-mode at 32 frames per second |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 104478 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 59000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: RECIPROCAL / Protocol: OTHER / Target criteria: FSC |
|---|---|
| Output model |  PDB-6qg0: |
 Movie
Movie Controller
Controller



























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)