+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the PP2A:B55-FAM122A complex | ||||||||||||
 Map data Map data | Relion 3D auto-refine merged map | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Protein Phosphatase 2A:B55 holoenzyme FAM122A inhibitor substrate binding cell cycle regulation / SIGNALING PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationprotein serine/threonine phosphatase inhibitor activity / meiotic spindle elongation / Integration of energy metabolism / PP2A-mediated dephosphorylation of key metabolic factors / regulation of microtubule binding / protein serine/threonine phosphatase complex / mitotic sister chromatid separation / MASTL Facilitates Mitotic Progression / regulation of meiotic cell cycle process involved in oocyte maturation / protein phosphatase type 2A complex ...protein serine/threonine phosphatase inhibitor activity / meiotic spindle elongation / Integration of energy metabolism / PP2A-mediated dephosphorylation of key metabolic factors / regulation of microtubule binding / protein serine/threonine phosphatase complex / mitotic sister chromatid separation / MASTL Facilitates Mitotic Progression / regulation of meiotic cell cycle process involved in oocyte maturation / protein phosphatase type 2A complex / meiotic sister chromatid cohesion, centromeric / peptidyl-serine dephosphorylation / peptidyl-threonine dephosphorylation / FAR/SIN/STRIPAK complex / positive regulation of microtubule binding / Regulation of glycolysis by fructose 2,6-bisphosphate metabolism / Inhibition of replication initiation of damaged DNA by RB1/E2F1 / female meiotic nuclear division / protein phosphatase regulator activity / protein antigen binding / GABA receptor binding / APC truncation mutants have impaired AXIN binding / AXIN missense mutants destabilize the destruction complex / Truncations of AMER1 destabilize the destruction complex / Initiation of Nuclear Envelope (NE) Reformation / ERKs are inactivated / positive regulation of extrinsic apoptotic signaling pathway in absence of ligand / Beta-catenin phosphorylation cascade / Signaling by GSK3beta mutants / CTNNB1 S33 mutants aren't phosphorylated / CTNNB1 S37 mutants aren't phosphorylated / CTNNB1 S45 mutants aren't phosphorylated / CTNNB1 T41 mutants aren't phosphorylated / mitotic G2/M transition checkpoint / negative regulation of epithelial to mesenchymal transition / response to morphine / regulation of growth / Disassembly of the destruction complex and recruitment of AXIN to the membrane / negative regulation of glycolytic process through fructose-6-phosphate / positive regulation of NLRP3 inflammasome complex assembly / myosin phosphatase activity / CTLA4 inhibitory signaling / protein serine/threonine phosphatase activity / Platelet sensitization by LDL / protein-serine/threonine phosphatase / Cyclin A/B1/B2 associated events during G2/M transition / regulation of cell differentiation / negative regulation of hippo signaling / ERK/MAPK targets / T cell homeostasis / regulation of G1/S transition of mitotic cell cycle / mesoderm development / phosphoprotein phosphatase activity / chromosome, centromeric region / negative regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction / DARPP-32 events / lateral plasma membrane / meiotic cell cycle / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / Amplification of signal from unattached kinetochores via a MAD2 inhibitory signal / Mitotic Prometaphase / EML4 and NUDC in mitotic spindle formation / Loss of Nlp from mitotic centrosomes / Loss of proteins required for interphase microtubule organization from the centrosome / Recruitment of mitotic centrosome proteins and complexes / Recruitment of NuMA to mitotic centrosomes / Anchoring of the basal body to the plasma membrane / Resolution of Sister Chromatid Cohesion / protein dephosphorylation / AURKA Activation by TPX2 / protein tyrosine phosphatase activity / chromosome segregation / RHO GTPases Activate Formins / response to lead ion / RAF activation / regulation of protein phosphorylation / Spry regulation of FGF signaling / Degradation of beta-catenin by the destruction complex / tau protein binding / positive regulation of protein serine/threonine kinase activity / PKR-mediated signaling / Cyclin D associated events in G1 / spindle pole / Regulation of TP53 Degradation / Negative regulation of MAPK pathway / Separation of Sister Chromatids / microtubule cytoskeleton / Regulation of PLK1 Activity at G2/M Transition / positive regulation of proteasomal ubiquitin-dependent protein catabolic process / mitotic cell cycle / PI5P, PP2A and IER3 Regulate PI3K/AKT Signaling / positive regulation of cell growth / protein-containing complex assembly / nuclear body / intracellular signal transduction / neuron projection / membrane raft / protein heterodimerization activity / neuronal cell body / glutamatergic synapse Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.79961 Å | ||||||||||||
 Authors Authors | Fuller JR / Padi SKR / Peti W / Page R | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation | Journal: Acta Crystallogr D Struct Biol / Year: 2018 Title: Real-space refinement in PHENIX for cryo-EM and crystallography. Authors: Pavel V Afonine / Billy K Poon / Randy J Read / Oleg V Sobolev / Thomas C Terwilliger / Alexandre Urzhumtsev / Paul D Adams /    Abstract: This article describes the implementation of real-space refinement in the phenix.real_space_refine program from the PHENIX suite. The use of a simplified refinement target function enables very fast ...This article describes the implementation of real-space refinement in the phenix.real_space_refine program from the PHENIX suite. The use of a simplified refinement target function enables very fast calculation, which in turn makes it possible to identify optimal data-restraint weights as part of routine refinements with little runtime cost. Refinement of atomic models against low-resolution data benefits from the inclusion of as much additional information as is available. In addition to standard restraints on covalent geometry, phenix.real_space_refine makes use of extra information such as secondary-structure and rotamer-specific restraints, as well as restraints or constraints on internal molecular symmetry. The re-refinement of 385 cryo-EM-derived models available in the Protein Data Bank at resolutions of 6 Å or better shows significant improvement of the models and of the fit of these models to the target maps. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40644.map.gz emd_40644.map.gz | 486.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40644-v30.xml emd-40644-v30.xml emd-40644.xml emd-40644.xml | 33 KB 33 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40644_fsc.xml emd_40644_fsc.xml | 19.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_40644.png emd_40644.png | 110.4 KB | ||
| Masks |  emd_40644_msk_1.map emd_40644_msk_1.map | 600.7 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-40644.cif.gz emd-40644.cif.gz | 8.3 KB | ||
| Others |  emd_40644_additional_1.map.gz emd_40644_additional_1.map.gz emd_40644_additional_2.map.gz emd_40644_additional_2.map.gz emd_40644_additional_3.map.gz emd_40644_additional_3.map.gz emd_40644_half_map_1.map.gz emd_40644_half_map_1.map.gz emd_40644_half_map_2.map.gz emd_40644_half_map_2.map.gz | 558.1 MB 362.9 MB 570 MB 487.5 MB 487.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40644 http://ftp.pdbj.org/pub/emdb/structures/EMD-40644 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40644 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40644 | HTTPS FTP |
-Related structure data
| Related structure data |  8so0MC  8ttbC  8tweC  8twiC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40644.map.gz / Format: CCP4 / Size: 600.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40644.map.gz / Format: CCP4 / Size: 600.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Relion 3D auto-refine merged map | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.534 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_40644_msk_1.map emd_40644_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
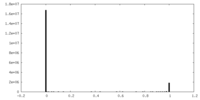
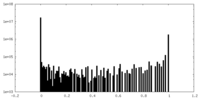
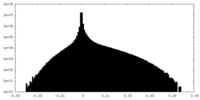
| Density Histograms |
-Additional map: Relion post-processed map, sharpened by automatically determined B-factor...
| File | emd_40644_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Relion post-processed map, sharpened by automatically determined B-factor (-32.3355) | ||||||||||||
| Projections & Slices |
| ||||||||||||
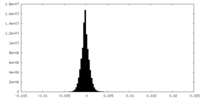
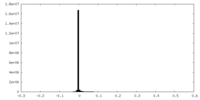
| Density Histograms |
-Additional map: Sharpened, local resolution-filtered map, as determined by the...
| File | emd_40644_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened, local resolution-filtered map, as determined by the algorithm implemented in Relion | ||||||||||||
| Projections & Slices |
| ||||||||||||
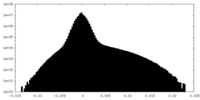
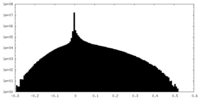
| Density Histograms |
-Additional map: LAFTER filtered map (using --sharp option)
| File | emd_40644_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | LAFTER filtered map (using --sharp option) | ||||||||||||
| Projections & Slices |
| ||||||||||||
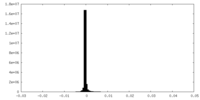
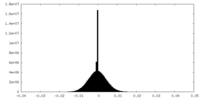
| Density Histograms |
-Half map: Relion 3D auto-refine half map 1
| File | emd_40644_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Relion 3D auto-refine half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Relion 3D auto-refine half map 2
| File | emd_40644_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Relion 3D auto-refine half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Quadruple complex of PP2A:B55 (PP2Aa:PP2Ac:B55) bound to FAM122A
| Entire | Name: Quadruple complex of PP2A:B55 (PP2Aa:PP2Ac:B55) bound to FAM122A |
|---|---|
| Components |
|
-Supramolecule #1: Quadruple complex of PP2A:B55 (PP2Aa:PP2Ac:B55) bound to FAM122A
| Supramolecule | Name: Quadruple complex of PP2A:B55 (PP2Aa:PP2Ac:B55) bound to FAM122A type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|
-Macromolecule #1: Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit...
| Macromolecule | Name: Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A alpha isoform type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 64.95798 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GHMSLYPIAV LIDELRNEDV QLRLNSIKKL STIALALGVE RTRSELLPFL TDTIYDEDEV LLALAEQLGT FTTLVGGPEY VHCLLPPLE SLATVEETVV RDKAVESLRA ISHEHSPSDL EAHFVPLVKR LAGGDWFTSR TSACGLFSVC YPRVSSAVKA E LRQYFRNL ...String: GHMSLYPIAV LIDELRNEDV QLRLNSIKKL STIALALGVE RTRSELLPFL TDTIYDEDEV LLALAEQLGT FTTLVGGPEY VHCLLPPLE SLATVEETVV RDKAVESLRA ISHEHSPSDL EAHFVPLVKR LAGGDWFTSR TSACGLFSVC YPRVSSAVKA E LRQYFRNL CSDDTPMVRR AAASKLGEFA KVLELDNVKS EIIPMFSNLA SDEQDSVRLL AVEACVNIAQ LLPQEDLEAL VM PTLRQAA EDKSWRVRYM VADKFTELQK AVGPEITKTD LVPAFQNLMK DCEAEVRAAA SHKVKEFCEN LSADCRENVI MSQ ILPCIK ELVSDANQHV KSALASVIMG LSPILGKDNT IEHLLPLFLA QLKDECPEVR LNIISNLDCV NEVIGIRQLS QSLL PAIVE LAEDAKWRVR LAIIEYMPLL AGQLGVEFFD EKLNSLCMAW LVDHVYAIRE AATSNLKKLV EKFGKEWAHA TIIPK VLAM SGDPNYLHRM TTLFCINVLS EVCGQDITTK HMLPTVLRMA GDPVANVRFN VAKSLQKIGP ILDNSTLQSE VKPILE KLT QDQDVDVKYF AQEALTVLSL A UniProtKB: Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A alpha isoform |
-Macromolecule #2: Serine/threonine-protein phosphatase 2A 55 kDa regulatory subunit...
| Macromolecule | Name: Serine/threonine-protein phosphatase 2A 55 kDa regulatory subunit B alpha isoform type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 52.10134 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GHMGSAGAGG GNDIQWCFSQ VKGAVDDDVA EADIISTVEF NHSGELLATG DKGGRVVIFQ QEQENKIQSH SRGEYNVYST FQSHEPEFD YLKSLEIEEK INKIRWLPQK NAAQFLLSTN DKTIKLWKIS ERDKRPEGYN LKEEDGRYRD PTTVTTLRVP V FRPMDLMV ...String: GHMGSAGAGG GNDIQWCFSQ VKGAVDDDVA EADIISTVEF NHSGELLATG DKGGRVVIFQ QEQENKIQSH SRGEYNVYST FQSHEPEFD YLKSLEIEEK INKIRWLPQK NAAQFLLSTN DKTIKLWKIS ERDKRPEGYN LKEEDGRYRD PTTVTTLRVP V FRPMDLMV EASPRRIFAN AHTYHINSIS INSDYETYLS ADDLRINLWH LEITDRSFNI VDIKPANMEE LTEVITAAEF HP NSCNTFV YSSSKGTIRL CDMRASALCD RHSKLFEEPE DPSNRSFFSE IISSISDVKF SHSGRYMMTR DYLSVKIWDL NME NRPVET YQVHEYLRSK LCSLYENDCI FDKFECCWNG SDSVVMTGSY NNFFRMFDRN TKRDITLEAS RENNKPRTVL KPRK VCASG KRKKDEISVD SLDFNKKILH TAWHPKENII AVATTNNLYI FQDKVN UniProtKB: Serine/threonine-protein phosphatase 2A 55 kDa regulatory subunit B alpha isoform |
-Macromolecule #3: Serine/threonine-protein phosphatase 2A catalytic subunit alpha i...
| Macromolecule | Name: Serine/threonine-protein phosphatase 2A catalytic subunit alpha isoform type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO / EC number: protein-serine/threonine phosphatase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 35.845375 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GHMDEKVFTK ELDQWIEQLN ECKQLSESQV KSLCEKAKEI LTKESNVQEV RCPVTVCGDV HGQFHDLMEL FRIGGKSPDT NYLFMGDYV DRGYYSVETV TLLVALKVRY RERITILRGN HESRQITQVY GFYDECLRKY GNANVWKYFT DLFDYLPLTA L VDGQIFCL ...String: GHMDEKVFTK ELDQWIEQLN ECKQLSESQV KSLCEKAKEI LTKESNVQEV RCPVTVCGDV HGQFHDLMEL FRIGGKSPDT NYLFMGDYV DRGYYSVETV TLLVALKVRY RERITILRGN HESRQITQVY GFYDECLRKY GNANVWKYFT DLFDYLPLTA L VDGQIFCL HGGLSPSIDT LDHIRALDRL QEVPHEGPMC DLLWSDPDDR GGWGISPRGA GYTFGQDISE TFNHANGLTL VS RAHQLVM EGYNWCHDRN VVTIFSAPNY CYRCGNQAAI MELDDTLKYS FLQFDPAPRR GEPHVTRRTP DYF(MLL) UniProtKB: Serine/threonine-protein phosphatase 2A catalytic subunit alpha isoform |
-Macromolecule #4: PPP2R1A-PPP2R2A-interacting phosphatase regulator 1
| Macromolecule | Name: PPP2R1A-PPP2R2A-interacting phosphatase regulator 1 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 10.680907 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GHMGGGLRRS NSAPLIHGLS DSSPVFQDEA PSARRNRTTF PSRHGLLLPA SPVRMHSSRL HQIKQEEGMD LINRETVHER EVQTAMQIS HSWEES UniProtKB: PPP2R1A-PPP2R2A-interacting phosphatase regulator 1 |
-Macromolecule #5: FE (III) ION
| Macromolecule | Name: FE (III) ION / type: ligand / ID: 5 / Number of copies: 1 / Formula: FE |
|---|---|
| Molecular weight | Theoretical: 55.845 Da |
-Macromolecule #6: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 6 / Number of copies: 1 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.2 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
Details: CHAPSO was added only immediately prior to vitrification | ||||||||||||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 35 sec. / Pretreatment - Atmosphere: AIR / Details: Gatan Solarus, 20W power for 35s using room air | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 291 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Average electron dose: 70.0 e/Å2 Details: Camera was operated in CDS mode, writing super-resolution movies with 59 frames |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated defocus max: 2.5300000000000002 µm / Calibrated defocus min: 0.55 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.9000000000000001 µm / Nominal defocus min: 0.7000000000000001 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||
|---|---|---|---|---|---|---|---|
| Details | Iterating between manual refinement in Coot and automated real-space refinement in Phenix | ||||||
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Target criteria: Cross-correlation | ||||||
| Output model |  PDB-8so0: |
 Movie
Movie Controller
Controller

























 Z
Z Y
Y X
X