+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM Structure of Mouse TLR4/MD-2/DLAM3 Complex | |||||||||
 Map data Map data | main map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Innate immune system / Toll-like receptors / TLR4 agonist / Vaccine adjuvants / Disaccharide-based Lipid A Mimetics / IMMUNE SYSTEM | |||||||||
| Function / homology |  Function and homology information Function and homology informationMyD88-independent TLR4 cascade / Caspase activation via Death Receptors in the presence of ligand / Toll Like Receptor 4 (TLR4) Cascade / Heme signaling / nitric oxide production involved in inflammatory response / Regulation of TLR by endogenous ligand / growth plate cartilage morphogenesis / TRIF-mediated programmed cell death / TRAF6-mediated induction of TAK1 complex within TLR4 complex / IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation ...MyD88-independent TLR4 cascade / Caspase activation via Death Receptors in the presence of ligand / Toll Like Receptor 4 (TLR4) Cascade / Heme signaling / nitric oxide production involved in inflammatory response / Regulation of TLR by endogenous ligand / growth plate cartilage morphogenesis / TRIF-mediated programmed cell death / TRAF6-mediated induction of TAK1 complex within TLR4 complex / IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation / positive regulation of cellular response to macrophage colony-stimulating factor stimulus / Regulation of TBK1, IKKε (IKBKE)-mediated activation of IRF3, IRF7 / NK T cell activation / Activation of IRF3, IRF7 mediated by TBK1, IKKε (IKBKE) / lipopolysaccharide immune receptor activity / positive regulation of nucleotide-binding oligomerization domain containing 1 signaling pathway / Toll-like receptor 4 binding / positive regulation of matrix metallopeptidase secretion / IKK complex recruitment mediated by RIP1 / mast cell activation / detection of lipopolysaccharide / regulation of dendritic cell cytokine production / innate immune response-activating signaling pathway / lipopolysaccharide receptor complex / positive regulation of lymphocyte proliferation / negative regulation of interleukin-23 production / cellular response to oxidised low-density lipoprotein particle stimulus / wound healing involved in inflammatory response / regulation of toll-like receptor 4 signaling pathway / B cell proliferation involved in immune response / positive regulation of nucleotide-binding oligomerization domain containing 2 signaling pathway / positive regulation of stress-activated MAPK cascade / leukotriene metabolic process / activation of NF-kappaB-inducing kinase activity / positive regulation of interleukin-1 production / TRIF-dependent toll-like receptor signaling pathway / positive regulation of interleukin-13 production / response to fatty acid / intestinal epithelial structure maintenance / macrophage activation / astrocyte development / microglia differentiation / regulation of tumor necrosis factor production / positive regulation of MHC class II biosynthetic process / positive regulation of platelet activation / signal transduction involved in regulation of gene expression / negative regulation of interleukin-17 production / positive regulation of macrophage activation / positive regulation of lipopolysaccharide-mediated signaling pathway / bone morphogenesis / positive regulation of chemokine (C-X-C motif) ligand 2 production / positive regulation of cytokine production involved in inflammatory response / negative regulation of cold-induced thermogenesis / positive regulation of extrinsic apoptotic signaling pathway / positive regulation of macrophage cytokine production / MyD88-dependent toll-like receptor signaling pathway / toll-like receptor 4 signaling pathway / positive regulation of smooth muscle cell migration / toll-like receptor signaling pathway / cellular response to lipoteichoic acid / positive regulation of NLRP3 inflammasome complex assembly / detection of temperature stimulus involved in sensory perception of pain / positive regulation of reactive oxygen species biosynthetic process / negative regulation of type II interferon production / negative regulation of interleukin-6 production / positive regulation of interferon-alpha production / positive regulation of interleukin-10 production / negative regulation of tumor necrosis factor production / detection of mechanical stimulus involved in sensory perception of pain / phagocytosis / phosphatidylinositol 3-kinase binding / phagocytic cup / positive regulation of B cell proliferation / ERK1 and ERK2 cascade / positive regulation of chemokine production / cellular response to platelet-derived growth factor stimulus / ruffle / positive regulation of interleukin-12 production / canonical NF-kappaB signal transduction / negative regulation of cytokine production involved in inflammatory response / activation of innate immune response / neurogenesis / positive regulation of interferon-beta production / positive regulation of smooth muscle cell proliferation / lipopolysaccharide-mediated signaling pathway / : / positive regulation of interleukin-1 beta production / positive regulation of interleukin-8 production / response to bacterium / lipopolysaccharide binding / cellular response to mechanical stimulus / microglial cell activation / negative regulation of ERK1 and ERK2 cascade / positive regulation of JNK cascade / positive regulation of interleukin-6 production / cellular response to type II interferon / positive regulation of type II interferon production / cellular response to amyloid-beta / positive regulation of nitric oxide biosynthetic process / positive regulation of tumor necrosis factor production Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.6 Å | |||||||||
 Authors Authors | Fu Y / Kim H / Kim HM | |||||||||
| Funding support |  Korea, Republic Of, 1 items Korea, Republic Of, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2025 Journal: Nat Commun / Year: 2025Title: Structural insight into TLR4/MD-2 activation by synthetic LPS mimetics with distinct binding modes. Authors: Yaoyao Fu / Hyojin Kim / Dong Sun Lee / Ah-Reum Han / Holger Heine / Alla Zamyatina / Ho Min Kim /    Abstract: The mammalian pattern-recognition receptor TLR4/MD-2 (Toll-like receptor 4/myeloid differentiation factor-2) can be activated by a wide variety of pathogen-associated and endogenous molecules, with ...The mammalian pattern-recognition receptor TLR4/MD-2 (Toll-like receptor 4/myeloid differentiation factor-2) can be activated by a wide variety of pathogen-associated and endogenous molecules, with Gram-negative bacterial lipopolysaccharide (LPS) being the primary natural TLR4 agonist. Activation of TLR4 triggers cellular signaling that enables the beneficial innate immune responses and enhances adaptive immunity, thereby emphasizing the potential of TLR4 agonists for the management of diseases with an immunopathological background and for use as vaccine adjuvants. Given the challenges associated with LPS-derived products, including structural complexity, heterogeneity, toxicity, and species specificity, synthetic molecules targeting TLR4/MD-2 offer a promising alternative. Here, we elucidate the structural basis for the recognition of synthetic LPS-mimicking glycolipids, Disaccharide Lipid A Mimetics (DLAMs), by human and mouse TLR4/MD-2 through cryo-EM structures of six dimeric [TLR4/MD-2/ligand] complexes resolved at 2.2-3.1 Å. We reveal that the specific binding modes of DLAMs, distinct from those of LPS, are essential for the species-independent TLR4 agonistic activity. DLAMs function as a molecular bridge, effectively induce the dimerization of TLR4/MD-2 complexes through specific carbohydrate structure-relevant ligand-protein interactions. Our findings reveal the distinct molecular modes of TLR4 activation, and provide a structural basis for the rationale design and development of innovative, highly potent TLR4-targeting immunotherapeutics and adjuvants. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_37794.map.gz emd_37794.map.gz | 121.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-37794-v30.xml emd-37794-v30.xml emd-37794.xml emd-37794.xml | 28.3 KB 28.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_37794_fsc.xml emd_37794_fsc.xml | 14.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_37794.png emd_37794.png | 73.2 KB | ||
| Filedesc metadata |  emd-37794.cif.gz emd-37794.cif.gz | 8.3 KB | ||
| Others |  emd_37794_half_map_1.map.gz emd_37794_half_map_1.map.gz emd_37794_half_map_2.map.gz emd_37794_half_map_2.map.gz | 226.4 MB 226.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-37794 http://ftp.pdbj.org/pub/emdb/structures/EMD-37794 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37794 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37794 | HTTPS FTP |
-Related structure data
| Related structure data |  8wryMC  8wo1C  8wqtC  8wsaC  8wtaC  9j03C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_37794.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_37794.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | main map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.848 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half B map
| File | emd_37794_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half B map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half A map
| File | emd_37794_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half A map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : mouse TLR4/MD2/DLAM3 complex
| Entire | Name: mouse TLR4/MD2/DLAM3 complex |
|---|---|
| Components |
|
-Supramolecule #1: mouse TLR4/MD2/DLAM3 complex
| Supramolecule | Name: mouse TLR4/MD2/DLAM3 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #2, #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Lymphocyte antigen 96
| Macromolecule | Name: Lymphocyte antigen 96 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 16.411711 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: EKQQWFCNSS DAIISYSYCD HLKFPISISS EPCIRLRGTN GFVHVEFIPR GNLKYLYFNL FISVNSIELP KRKEVLCHGH DDDYSFCRA LKGETVNTSI PFSFEGILFP KGHYRCVAEA IAGDTEEKLF CLNFTIIHRR DVN UniProtKB: Lymphocyte antigen 96 |
-Macromolecule #2: Toll-like receptor 4
| Macromolecule | Name: Toll-like receptor 4 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 68.651281 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: NPCIEVVPNI TYQCMDQKLS KVPDDIPSST KNIDLSFNPL KILKSYSFSN FSELQWLDLS RCEIETIEDK AWHGLHHLSN LILTGNPIQ SFSPGSFSGL TSLENLVAVE TKLASLESFP IGQLITLKKL NVAHNFIHSC KLPAYFSNLT NLVHVDLSYN Y IQTITVND ...String: NPCIEVVPNI TYQCMDQKLS KVPDDIPSST KNIDLSFNPL KILKSYSFSN FSELQWLDLS RCEIETIEDK AWHGLHHLSN LILTGNPIQ SFSPGSFSGL TSLENLVAVE TKLASLESFP IGQLITLKKL NVAHNFIHSC KLPAYFSNLT NLVHVDLSYN Y IQTITVND LQFLRENPQV NLSLDMSLNP IDFIQDQAFQ GIKLHELTLR GNFNSSNIMK TCLQNLAGLH VHRLILGEFK DE RNLEIFE PSIMEGLCDV TIDEFRLTYT NDFSDDIVKF HCLANVSAMS LAGVSIKYLE DVPKHFKWQS LSIIRCQLKQ FPT LDLPFL KSLTLTMNKG SISFKKVALP SLSYLDLSRN ALSFSGCCSY SDLGTNSLRH LDLSFNGAII MSANFMGLEE LQHL DFQHS TLKRVTEFSA FLSLEKLLYL DISYTNTKID FDGIFLGLTS LNTLKMAGNS FKDNTLSNVF ANTTNLTFLD LSKCQ LEQI SWGVFDTLHR LQLLNMSHNN LLFLDSSHYN QLYSLSTLDC SFNRIETSKG ILQHFPKSLA FFNLTNNSVA CICEHQ KFL QWVKEQKQFL VNVEQMTCAT PVEMNTSLVL DFNNSTCYMY K UniProtKB: Toll-like receptor 4 |
-Macromolecule #4: (3R)-3-(dodecanoyloxy)tetradecanoic acid
| Macromolecule | Name: (3R)-3-(dodecanoyloxy)tetradecanoic acid / type: ligand / ID: 4 / Number of copies: 4 / Formula: 2IL |
|---|---|
| Molecular weight | Theoretical: 426.673 Da |
| Chemical component information |  ChemComp-2IL: |
-Macromolecule #5: (3R)-3-(tetradecanoyloxy)tetradecanoic acid
| Macromolecule | Name: (3R)-3-(tetradecanoyloxy)tetradecanoic acid / type: ligand / ID: 5 / Number of copies: 2 / Formula: 0IL |
|---|---|
| Molecular weight | Theoretical: 454.726 Da |
-Macromolecule #6: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 6 / Number of copies: 16 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #7: 2-amino-2-deoxy-4-O-phosphono-alpha-D-glucopyranose
| Macromolecule | Name: 2-amino-2-deoxy-4-O-phosphono-alpha-D-glucopyranose / type: ligand / ID: 7 / Number of copies: 2 / Formula: GP4 |
|---|---|
| Molecular weight | Theoretical: 259.151 Da |
| Chemical component information | 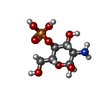 ChemComp-GP4: |
-Macromolecule #8: 2-(hydroxymethyl)-5-methoxy-3,6-bis(oxidanyl)pyran-4-one
| Macromolecule | Name: 2-(hydroxymethyl)-5-methoxy-3,6-bis(oxidanyl)pyran-4-one type: ligand / ID: 8 / Number of copies: 2 / Formula: XIQ |
|---|---|
| Molecular weight | Theoretical: 188.135 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 5 sec. / Details: 10mA | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 18733 / Average electron dose: 66.2 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 58963 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.9000000000000001 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 105000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)






































