+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | cryo-EM structure of the CED-4/CED-3 holoenzyme | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CED-4 / CED-3 / holoenzyme / APOPTOSIS | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of cellular response to manganese ion / positive regulation of cellular response to gamma radiation / : / Apoptosis induced DNA fragmentation / AKT phosphorylates targets in the cytosol / Signaling by Hippo / : / : / : / Regulation of TNFR1 signaling ...negative regulation of cellular response to manganese ion / positive regulation of cellular response to gamma radiation / : / Apoptosis induced DNA fragmentation / AKT phosphorylates targets in the cytosol / Signaling by Hippo / : / : / : / Regulation of TNFR1 signaling / BH1 domain binding / regulation of vulval development / regulation of development, heterochronic / caspase complex / positive regulation of apoptotic process involved in development / positive regulation of synapse pruning / peptidase activator activity involved in apoptotic process / caspase-7 / caspase binding / positive regulation of protein processing / embryonic morphogenesis / regulation of cell fate specification / apoptotic process involved in development / cysteine-type endopeptidase activator activity / actin filament depolymerization / programmed cell death / embryo development ending in birth or egg hatching / negative regulation of execution phase of apoptosis / execution phase of apoptosis / cysteine-type endopeptidase activator activity involved in apoptotic process / muscle cell cellular homeostasis / regulation of synapse organization / regulation of locomotion / regulation of cell size / protein autoprocessing / BH3 domain binding / endopeptidase activator activity / regulation of cell adhesion / protein catabolic process / enzyme activator activity / ADP binding / regulation of protein stability / positive regulation of neuron apoptotic process / presynapse / endopeptidase activity / nuclear membrane / defense response to Gram-negative bacterium / perikaryon / positive regulation of apoptotic process / cysteine-type endopeptidase activity / neuronal cell body / apoptotic process / negative regulation of apoptotic process / perinuclear region of cytoplasm / magnesium ion binding / protein-containing complex / mitochondrion / proteolysis / ATP binding / identical protein binding / nucleus / membrane / cytosol / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
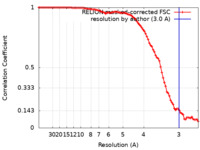
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Li Y / Tian L / Zhang Y / Shi Y | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Life Sci Alliance / Year: 2023 Journal: Life Sci Alliance / Year: 2023Title: Structural insights into CED-3 activation. Authors: Yini Li / Lu Tian / Ying Zhang / Yigong Shi /  Abstract: In , onset of programmed cell death is marked with the activation of CED-3, a process that requires assembly of the CED-4 apoptosome. Activated CED-3 forms a holoenzyme with the CED-4 apoptosome to ...In , onset of programmed cell death is marked with the activation of CED-3, a process that requires assembly of the CED-4 apoptosome. Activated CED-3 forms a holoenzyme with the CED-4 apoptosome to cleave a wide range of substrates, leading to irreversible cell death. Despite decades of investigations, the underlying mechanism of CED-4-facilitated CED-3 activation remains elusive. Here, we report cryo-EM structures of the CED-4 apoptosome and three distinct CED-4/CED-3 complexes that mimic different activation stages for CED-3. In addition to the previously reported octamer in crystal structures, CED-4, alone or in complex with CED-3, exists in multiple oligomeric states. Supported by biochemical analyses, we show that the conserved CARD-CARD interaction promotes CED-3 activation, and initiation of programmed cell death is regulated by the dynamic organization of the CED-4 apoptosome. #1:  Journal: Life Sci Alliance / Year: 2023 Journal: Life Sci Alliance / Year: 2023Title: Structural insights into CED-3 activation Authors: Li Y / Tian L / Zhang Y / Shi Y | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36459.map.gz emd_36459.map.gz | 45.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36459-v30.xml emd-36459-v30.xml emd-36459.xml emd-36459.xml | 18.2 KB 18.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_36459_fsc.xml emd_36459_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_36459.png emd_36459.png | 51.7 KB | ||
| Filedesc metadata |  emd-36459.cif.gz emd-36459.cif.gz | 6 KB | ||
| Others |  emd_36459_additional_1.map.gz emd_36459_additional_1.map.gz emd_36459_half_map_1.map.gz emd_36459_half_map_1.map.gz emd_36459_half_map_2.map.gz emd_36459_half_map_2.map.gz | 45.4 MB 45.9 MB 45.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36459 http://ftp.pdbj.org/pub/emdb/structures/EMD-36459 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36459 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36459 | HTTPS FTP |
-Related structure data
| Related structure data |  8jolMC  8jnsC  8jo0C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_36459.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36459.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.32 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_36459_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_36459_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_36459_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : ternary complex of CED-4 with CED-3 CARD and CED-3 catalytic domain
| Entire | Name: ternary complex of CED-4 with CED-3 CARD and CED-3 catalytic domain |
|---|---|
| Components |
|
-Supramolecule #1: ternary complex of CED-4 with CED-3 CARD and CED-3 catalytic domain
| Supramolecule | Name: ternary complex of CED-4 with CED-3 CARD and CED-3 catalytic domain type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Cell death protein 4
| Macromolecule | Name: Cell death protein 4 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 62.953266 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MLCEIECRAL STAHTRLIHD FEPRDALTYL EGKNIFTEDH SELISKMSTR LERIANFLRI YRRQASELGP LIDFFNYNNQ SHLADFLED YIDFAINEPD LLRPVVIAPQ FSRQMLDRKL LLGNVPKQMT CYIREYHVDR VIKKLDEMCD LDSFFLFLHG R AGSGKSVI ...String: MLCEIECRAL STAHTRLIHD FEPRDALTYL EGKNIFTEDH SELISKMSTR LERIANFLRI YRRQASELGP LIDFFNYNNQ SHLADFLED YIDFAINEPD LLRPVVIAPQ FSRQMLDRKL LLGNVPKQMT CYIREYHVDR VIKKLDEMCD LDSFFLFLHG R AGSGKSVI ASQALSKSDQ LIGINYDSIV WLKDSGTAPK STFDLFTDIL LMLKSEDDLL NFPSVEHVTS VVLKRMICNA LI DRPNTLF VFDDVVQEET IRWAQELRLR CLVTTRDVEI SNAASQTCEF IEVTSLEIDE CYDFLEAYGM PMPVGEKEED VLN KTIELS SGNPATLMMF FKSCEPKTFE KMAQLNNKLE SRGLVGVECI TPYSYKSLAM ALQRCVEVLS DEDRSALAFA VVMP PGVDI PVKLWSCVIP VDICSNEEEQ LDDEVADRLK RLSKRGALLS GKRMPVLTFK IDHIIHMFLK HVVDAQTIAN GISIL EQRL LEIGNNNVSV PERHIPSHFQ KFRRSSASEM YPKTTEETVI RPEDFPKFMQ LHQKFYDSLK NFACC UniProtKB: Cell death protein 4 |
-Macromolecule #2: Cell death protein 3
| Macromolecule | Name: Cell death protein 3 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: caspase-7 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 56.693902 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MMRQDRRSLL ERNIMMFSSH LKVDEILEVL IAKQVLNSDN GDMINSCGTV REKRREIVKA VQRRGDVAFD AFYDALRSTG HEGLAEVLE PLARSVDSNA VEFECPMSPA SHRRSRALSP AGYTSPTRVH RDSVSSVSSF TSYQDIYSRA RSRSRSRALH S SDRHNYSS ...String: MMRQDRRSLL ERNIMMFSSH LKVDEILEVL IAKQVLNSDN GDMINSCGTV REKRREIVKA VQRRGDVAFD AFYDALRSTG HEGLAEVLE PLARSVDSNA VEFECPMSPA SHRRSRALSP AGYTSPTRVH RDSVSSVSSF TSYQDIYSRA RSRSRSRALH S SDRHNYSS PPVNAFPSQP SSANSSFTGC SSLGYSSSRN RSFSKASGPT QYIFHEEDMN FVDAPTISRV FDEKTMYRNF SS PRGMCLI INNEHFEQMP TRNGTKADKD NLTNLFRCMG YTVICKDNLT GRGMLLTIRD FAKHESHGDS AILVILSHGE ENV IIGVDD IPISTHEIYD LLNAANAPRL ANKPKIVFVQ ACRGERRDNG FPVLDSVDGV PAFLRRGWDN RDGPLFNFLG CVRP QVQQV WRKKPSQADI LIAYATTAQY VSWRNSARGS WFIQAVCEVF STHAKDMDVV ELLTEVNKKV ACGFQTSQGS NILKQ MPEM TSRLLKKFYF WPEARNSAV UniProtKB: Cell death protein 3 |
-Macromolecule #3: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 3 / Number of copies: 2 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #4: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 4 / Number of copies: 2 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)