+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Dopamine Receptor D2R-Gi-Rotigotine complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Dopamine / Dopamine receptor / GPCR / D2R / Gi / Rotigotine / Polypharmacology Parkinson's disease / Restless legs syndrome / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of dopamine receptor signaling pathway / positive regulation of dopamine uptake involved in synaptic transmission / negative regulation of dephosphorylation / positive regulation of glial cell-derived neurotrophic factor production / acid secretion / nervous system process involved in regulation of systemic arterial blood pressure / regulation of synapse structural plasticity / dopamine neurotransmitter receptor activity, coupled via Gi/Go / response to histamine / negative regulation of circadian sleep/wake cycle, sleep ...negative regulation of dopamine receptor signaling pathway / positive regulation of dopamine uptake involved in synaptic transmission / negative regulation of dephosphorylation / positive regulation of glial cell-derived neurotrophic factor production / acid secretion / nervous system process involved in regulation of systemic arterial blood pressure / regulation of synapse structural plasticity / dopamine neurotransmitter receptor activity, coupled via Gi/Go / response to histamine / negative regulation of circadian sleep/wake cycle, sleep / regulation of locomotion involved in locomotory behavior / neuron-neuron synaptic transmission / negative regulation of cellular response to hypoxia / adenylate cyclase-inhibiting dopamine receptor signaling pathway / response to inactivity / regulation of potassium ion transport / orbitofrontal cortex development / negative regulation of dopamine secretion / adenohypophysis development / cerebral cortex GABAergic interneuron migration / negative regulation of neuron migration / hyaloid vascular plexus regression / Dopamine receptors / branching morphogenesis of a nerve / regulation of dopamine uptake involved in synaptic transmission / dopamine binding / positive regulation of growth hormone secretion / phospholipase C-activating dopamine receptor signaling pathway / heterotrimeric G-protein binding / peristalsis / beta-arrestin-dependent dopamine receptor signaling pathway / G protein-coupled receptor complex / grooming behavior / positive regulation of renal sodium excretion / drinking behavior / striatum development / auditory behavior / positive regulation of G protein-coupled receptor signaling pathway / dopaminergic synapse / behavioral response to ethanol / positive regulation of multicellular organism growth / non-motile cilium / G protein-coupled receptor internalization / response to iron ion / adult walking behavior / negative regulation of synaptic transmission, glutamatergic / arachidonate secretion / positive regulation of urine volume / positive regulation of neuroblast proliferation / ciliary membrane / regulation of synaptic transmission, GABAergic / positive regulation of cytokinesis / response to morphine / negative regulation of cytosolic calcium ion concentration / pigmentation / temperature homeostasis / dopamine metabolic process / dopamine uptake involved in synaptic transmission / regulation of dopamine secretion / neuroblast proliferation / positive regulation of receptor internalization / lateral plasma membrane / negative regulation of protein secretion / associative learning / response to light stimulus / cellular response to ethanol / potassium channel regulator activity / endocytic vesicle / G-protein alpha-subunit binding / response to axon injury / sperm flagellum / postsynaptic modulation of chemical synaptic transmission / long-term memory / prepulse inhibition / regulation of sodium ion transport / cellular response to retinoic acid / negative regulation of blood pressure / release of sequestered calcium ion into cytosol / adenylate cyclase inhibitor activity / behavioral response to cocaine / positive regulation of protein localization to cell cortex / T cell migration / positive regulation of relaxation of smooth muscle / Adenylate cyclase inhibitory pathway / synapse assembly / D2 dopamine receptor binding / axonogenesis / epithelial cell proliferation / regulation of heart rate / negative regulation of innate immune response / adenylate cyclase-inhibiting serotonin receptor signaling pathway / G protein-coupled serotonin receptor binding / ionotropic glutamate receptor binding / axon terminus / acrosomal vesicle / negative regulation of cell migration / presynaptic modulation of chemical synaptic transmission / cellular response to forskolin / negative regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction / regulation of mitotic spindle organization Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Xu P / Huang S / Zhuang Y / Mao C / Zhang Y / Wang Y / Li H / Jiang Y / Xu HE | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Res / Year: 2023 Journal: Cell Res / Year: 2023Title: Structural genomics of the human dopamine receptor system. Authors: Peiyu Xu / Sijie Huang / Brian E Krumm / Youwen Zhuang / Chunyou Mao / Yumu Zhang / Yue Wang / Xi-Ping Huang / Yong-Feng Liu / Xinheng He / Huadong Li / Wanchao Yin / Yi Jiang / Yan Zhang / ...Authors: Peiyu Xu / Sijie Huang / Brian E Krumm / Youwen Zhuang / Chunyou Mao / Yumu Zhang / Yue Wang / Xi-Ping Huang / Yong-Feng Liu / Xinheng He / Huadong Li / Wanchao Yin / Yi Jiang / Yan Zhang / Bryan L Roth / H Eric Xu /   Abstract: The dopaminergic system, including five dopamine receptors (D1R to D5R), plays essential roles in the central nervous system (CNS); and ligands that activate dopamine receptors have been used to ...The dopaminergic system, including five dopamine receptors (D1R to D5R), plays essential roles in the central nervous system (CNS); and ligands that activate dopamine receptors have been used to treat many neuropsychiatric disorders, including Parkinson's Disease (PD) and schizophrenia. Here, we report cryo-EM structures of all five subtypes of human dopamine receptors in complex with G protein and bound to the pan-agonist, rotigotine, which is used to treat PD and restless legs syndrome. The structures reveal the basis of rotigotine recognition in different dopamine receptors. Structural analysis together with functional assays illuminate determinants of ligand polypharmacology and selectivity. The structures also uncover the mechanisms of dopamine receptor activation, unique structural features among the five receptor subtypes, and the basis of G protein coupling specificity. Our work provides a comprehensive set of structural templates for the rational design of specific ligands to treat CNS diseases targeting the dopaminergic system. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35684.map.gz emd_35684.map.gz | 17.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35684-v30.xml emd-35684-v30.xml emd-35684.xml emd-35684.xml | 19.7 KB 19.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_35684.png emd_35684.png | 60.8 KB | ||
| Filedesc metadata |  emd-35684.cif.gz emd-35684.cif.gz | 6.7 KB | ||
| Others |  emd_35684_half_map_1.map.gz emd_35684_half_map_1.map.gz emd_35684_half_map_2.map.gz emd_35684_half_map_2.map.gz | 20.6 MB 20.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35684 http://ftp.pdbj.org/pub/emdb/structures/EMD-35684 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35684 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35684 | HTTPS FTP |
-Related structure data
| Related structure data |  8irsMC  8irrC  8irtC  8iruC  8irvC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_35684.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35684.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.045 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_35684_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_35684_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Dopamine Receptor D2R-Gi-Rotigotine complex
| Entire | Name: Dopamine Receptor D2R-Gi-Rotigotine complex |
|---|---|
| Components |
|
-Supramolecule #1: Dopamine Receptor D2R-Gi-Rotigotine complex
| Supramolecule | Name: Dopamine Receptor D2R-Gi-Rotigotine complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Guanine nucleotide-binding protein G(i) subunit alpha-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(i) subunit alpha-1 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.414047 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKNTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI ...String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKNTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI PTQQDVLRTR VKTTGIVETH FTFKDLHFKM FDVGAQRSER KKWIHCFEGV TAIIFCVALS DYDLVLAEDE EM NRMHASM KLFDSICNNK WFTDTSIILF LNKKDLFEEK IKKSPLTICY PEYAGSNTYE EAAAYIQCQF EDLNKRKDTK EIY THFTCS TDTKNVQFVF DAVTDVIIKN NLKDCGLF UniProtKB: Guanine nucleotide-binding protein G(i) subunit alpha-1 |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.915496 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MGSLLQSELD QLRQEAEQLK NQIRDARKAC ADATLSQITN NIDPVGRIQM RTRRTLRGHL AKIYAMHWGT DSRLLVSASQ DGKLIIWDS YTTNKVHAIP LRSSWVMTCA YAPSGNYVAC GGLDNICSIY NLKTREGNVR VSRELAGHTG YLSCCRFLDD N QIVTSSGD ...String: MGSLLQSELD QLRQEAEQLK NQIRDARKAC ADATLSQITN NIDPVGRIQM RTRRTLRGHL AKIYAMHWGT DSRLLVSASQ DGKLIIWDS YTTNKVHAIP LRSSWVMTCA YAPSGNYVAC GGLDNICSIY NLKTREGNVR VSRELAGHTG YLSCCRFLDD N QIVTSSGD TTCALWDIET GQQTTTFTGH TGDVMSLSLA PDTRLFVSGA CDASAKLWDV REGMCRQTFT GHESDINAIC FF PNGNAFA TGSDDATCRL FDLRADQELM TYSHDNIICG ITSVSFSKSG RLLLAGYDDF NCNVWDALKA DRAGVLAGHD NRV SCLGVT DDGMAVATGS WDSFLKIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: ScFv16
| Macromolecule | Name: ScFv16 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 26.466486 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS ...String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS NGNTYLYWFL QRPGQSPQLL IYRMSNLASG VPDRFSGSGS GTAFTLTISR LEAEDVGVYY CMQHLEYPLT FG AGTKLEL K |
-Macromolecule #4: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #5: Soluble cytochrome b562,D(2) dopamine receptor
| Macromolecule | Name: Soluble cytochrome b562,D(2) dopamine receptor / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 67.234578 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: DYKDDDDAKL QTMHHHHHHH HHHHHHHHAD LEDNWETLND NLKVIEKADN AAQVKDALTK MRAAALDAQK ATPPKLEDKS PDSPEMKDF RHGFDILVGQ IDDALKLANE GKVKEAQAAA EQLKTTRNAY IQKYLASENL YFQGGTMDPL NLSWYDDDLE R QNWSRPFN ...String: DYKDDDDAKL QTMHHHHHHH HHHHHHHHAD LEDNWETLND NLKVIEKADN AAQVKDALTK MRAAALDAQK ATPPKLEDKS PDSPEMKDF RHGFDILVGQ IDDALKLANE GKVKEAQAAA EQLKTTRNAY IQKYLASENL YFQGGTMDPL NLSWYDDDLE R QNWSRPFN GSDGKADRPH YNYYATLLTL LIAVIVFGNV LVCMAVSREK ALQTTTNYLI VSLAVADLLV ATLVMPWVVY LE VVGEWKF SRIHCDIFVT LDVMMCTASI LNLCAISIDR YTAVAMPMLY NTRYSSKRRV TVMISIVWVL SFTISCPLLF GLN NADQNE CIIANPAFVV YSSIVSFYVP FIVTLLVYIK IYIVLRRRRK RVNTKRSSRA FRAHLRAPLK GNCTHPEDMK LCTV IMKSN GSFPVNRRRV EAARRAQELE MEMLSSTSPP ERTRYSPIPP SHHQLTLPDP SHHGLHSTPD SPAKPEKNGH AKDHP KIAK IFEIQTMPNG KTRTSLKTMS RRKLSQQKEK KATQMLAIVL GVFIICWLPF FITHILNIHC DCNIPPVLYS AFTWLG YVN SAVNPIIYTT FNIEFRKAFL KILHC UniProtKB: Soluble cytochrome b562, D(2) dopamine receptor |
-Macromolecule #6: Rotigotine
| Macromolecule | Name: Rotigotine / type: ligand / ID: 6 / Number of copies: 1 / Formula: R5F |
|---|---|
| Molecular weight | Theoretical: 315.473 Da |
| Chemical component information | 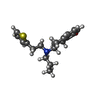 ChemComp-R5F: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: EMDB MAP EMDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.0 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 140237 |
| Initial angle assignment | Type: RANDOM ASSIGNMENT |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller






































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)