+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2989 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The structure of the COPI coat linkage IV | |||||||||
 Map data Map data | Reconstruction of the COPI coat linkage IV | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | COPI / coatomer / coated vesicles | |||||||||
| Function / homology |  Function and homology information Function and homology informationcerebellar Purkinje cell layer maturation / protein localization to cell leading edge / Synthesis of PIPs at the plasma membrane / protein localization to axon / VxPx cargo-targeting to cilium / Synthesis of PIPs at the Golgi membrane / Intra-Golgi traffic / trans-Golgi Network Vesicle Budding / protein localization to Golgi membrane / Golgi localization ...cerebellar Purkinje cell layer maturation / protein localization to cell leading edge / Synthesis of PIPs at the plasma membrane / protein localization to axon / VxPx cargo-targeting to cilium / Synthesis of PIPs at the Golgi membrane / Intra-Golgi traffic / trans-Golgi Network Vesicle Budding / protein localization to Golgi membrane / Golgi localization / COPI-coated vesicle / pancreatic juice secretion / regulation of Golgi organization / organelle membrane contact site / COPI vesicle coat / COPI-dependent Golgi-to-ER retrograde traffic / COPI-mediated anterograde transport / COPI-mediated anterograde transport / Golgi vesicle transport / COPI-dependent Golgi-to-ER retrograde traffic / positive regulation of mitochondrial fusion / organelle transport along microtubule / regulation of fatty acid metabolic process / establishment of Golgi localization / intra-Golgi vesicle-mediated transport / Golgi to plasma membrane transport / retrograde vesicle-mediated transport, Golgi to endoplasmic reticulum / pigmentation / Golgi-associated vesicle / positive regulation of mitochondrial fission / endoplasmic reticulum-Golgi intermediate compartment / protein secretion / endoplasmic reticulum to Golgi vesicle-mediated transport / vesicle-mediated transport / Neutrophil degranulation / adult locomotory behavior / small monomeric GTPase / establishment of localization in cell / macroautophagy / protein kinase C binding / intracellular protein transport / hormone activity / protein transport / growth cone / Golgi membrane / axon / GTPase activity / mRNA binding / neuronal cell body / GTP binding / structural molecule activity / Golgi apparatus / endoplasmic reticulum / extracellular space / nucleoplasm / plasma membrane / cytosol / cytoplasm Similarity search - Function | |||||||||
| Biological species |   | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 21.0 Å | |||||||||
 Authors Authors | Dodonova SO / Diestelkoetter-Bachert P / von Appen A / Hagen WJH / Beck R / Beck M / Wieland F / Briggs JAG | |||||||||
 Citation Citation |  Journal: Science / Year: 2015 Journal: Science / Year: 2015Title: VESICULAR TRANSPORT. A structure of the COPI coat and the role of coat proteins in membrane vesicle assembly. Authors: S O Dodonova / P Diestelkoetter-Bachert / A von Appen / W J H Hagen / R Beck / M Beck / F Wieland / J A G Briggs /  Abstract: Transport of material within cells is mediated by trafficking vesicles that bud from one cellular compartment and fuse with another. Formation of a trafficking vesicle is driven by membrane coats ...Transport of material within cells is mediated by trafficking vesicles that bud from one cellular compartment and fuse with another. Formation of a trafficking vesicle is driven by membrane coats that localize cargo and polymerize into cages to bend the membrane. Although extensive structural information is available for components of these coats, the heterogeneity of trafficking vesicles has prevented an understanding of how complete membrane coats assemble on the membrane. We combined cryo-electron tomography, subtomogram averaging, and cross-linking mass spectrometry to derive a complete model of the assembled coat protein complex I (COPI) coat involved in traffic between the Golgi and the endoplasmic reticulum. The highly interconnected COPI coat structure contradicted the current "adaptor-and-cage" understanding of coated vesicle formation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2989.map.gz emd_2989.map.gz | 28.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2989-v30.xml emd-2989-v30.xml emd-2989.xml emd-2989.xml | 13 KB 13 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_2989_fsc.xml emd_2989_fsc.xml | 8.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_2989.jpg emd_2989.jpg | 187.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2989 http://ftp.pdbj.org/pub/emdb/structures/EMD-2989 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2989 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2989 | HTTPS FTP |
-Validation report
| Summary document |  emd_2989_validation.pdf.gz emd_2989_validation.pdf.gz | 256.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_2989_full_validation.pdf.gz emd_2989_full_validation.pdf.gz | 255.5 KB | Display | |
| Data in XML |  emd_2989_validation.xml.gz emd_2989_validation.xml.gz | 10 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2989 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2989 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2989 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2989 | HTTPS FTP |
-Related structure data
| Related structure data |  5a1yMC  2985C  2986C  2987C  2988C  5a1uC  5a1vC  5a1wC  5a1xC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_2989.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2989.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
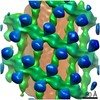
| Annotation | Reconstruction of the COPI coat linkage IV | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.019 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : COPI coat linkage IV on the membrane
| Entire | Name: COPI coat linkage IV on the membrane |
|---|---|
| Components |
|
-Supramolecule #1000: COPI coat linkage IV on the membrane
| Supramolecule | Name: COPI coat linkage IV on the membrane / type: sample / ID: 1000 Details: The COPI coated vesicles were formed in an in vitro reconstituted budding reaction, which was plunge-frozen without further purification. Number unique components: 2 |
|---|
-Macromolecule #1: Coat protein 1
| Macromolecule | Name: Coat protein 1 / type: protein_or_peptide / ID: 1 / Name.synonym: COPI / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 560 KDa |
| Recombinant expression | Organism:  |
| Sequence | InterPro: Coatomer subunit alpha, Coatomer beta' subunit (COPB2), Coatomer, epsilon subunit, Coatomer beta subunit (COPB1), Coatomer delta subunit, Coatomer gamma subunit, AP complex, mu/sigma subunit |
-Macromolecule #2: ADP-ribosylation factor 1
| Macromolecule | Name: ADP-ribosylation factor 1 / type: protein_or_peptide / ID: 2 / Name.synonym: Arf1 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 20 KDa |
| Recombinant expression | Organism:  |
| Sequence | UniProtKB: ADP-ribosylation factor 1 / InterPro: Small GTPase superfamily, ARF type |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 / Details: 50 mM HEPES, 50 mM KAc, 1mM MgCl2 |
|---|---|
| Grid | Details: C-Flat Multihole 3C-50 grids glow discharged |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 85 % / Instrument: HOMEMADE PLUNGER Method: The grids were glow discharged for 1 min at 20 mA. 5 ul of 10 nm fiducial gold was added to 40 ul of reaction mix. The reaction mix with the in vitro formed coated vesicles was applied to a ...Method: The grids were glow discharged for 1 min at 20 mA. 5 ul of 10 nm fiducial gold was added to 40 ul of reaction mix. The reaction mix with the in vitro formed coated vesicles was applied to a grid. The grid was blotted for 12 seconds at room temperature before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 89 K / Max: 91 K / Average: 90 K |
| Specialist optics | Energy filter - Name: GATAN GIF 2002 / Energy filter - Lower energy threshold: 0.0 eV / Energy filter - Upper energy threshold: 20.0 eV |
| Date | Feb 7, 2014 |
| Image recording | Category: CCD / Film or detector model: GATAN MULTISCAN / Average electron dose: 45 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 42000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Tilt series - Axis1 - Min angle: -45 ° / Tilt series - Axis1 - Max angle: 60 ° |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - #0 - Chain ID: A / Chain - #1 - Chain ID: B |
|---|---|
| Software | Name:  Chimera Chimera |
| Details | The crystal structure (chains A and B) was fitted into the EM map using global search option in Chimera software. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: cross-correlation |
| Output model |  PDB-5a1y: |
 Movie
Movie Controller
Controller






















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)























