登録情報 データベース : EMDB / ID : EMD-16331タイトル RNA polymerase II pre-initiation complex with the proximal +1 nucleosome (PIC-Nuc10W) 複合体 : RNA polymerase II pre-initiation complex with the proximal +1 nucleosome複合体 : General transcription factor IIH複合体 : Mammalian RNA polymerase II複合体 : Nucleosome (protein)複合体 : Nucleosome (DNA)複合体 : General transcription factor IIB複合体 : TATA-box binding protein複合体 : General transcription factor IIF複合体 : General transcription factor IIA複合体 : General transcription factor IIEタンパク質・ペプチド : x 11種リガンド : x 3種 / / / 機能・相同性 分子機能 ドメイン・相同性 構成要素
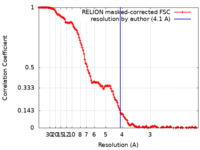
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / 生物種 Homo sapiens (ヒト) / Sus scrofa (ブタ) / Xenopus laevis (アフリカツメガエル) / 手法 / / 解像度 : 4.1 Å Abril-Garrido J / Dienemann C / Grabbe F / Velychko T / Lidschreiber M / Wang H / Cramer P 資金援助 Organization Grant number 国 German Research Foundation (DFG) European Research Council (ERC) European Union
ジャーナル : Mol Cell / 年 : 2023タイトル : Structural basis of transcription reduction by a promoter-proximal +1 nucleosome.著者 : Julio Abril-Garrido / Christian Dienemann / Frauke Grabbe / Taras Velychko / Michael Lidschreiber / Haibo Wang / Patrick Cramer / 要旨 : At active human genes, the +1 nucleosome is located downstream of the RNA polymerase II (RNA Pol II) pre-initiation complex (PIC). However, at inactive genes, the +1 nucleosome is found further ... At active human genes, the +1 nucleosome is located downstream of the RNA polymerase II (RNA Pol II) pre-initiation complex (PIC). However, at inactive genes, the +1 nucleosome is found further upstream, at a promoter-proximal location. Here, we establish a model system to show that a promoter-proximal +1 nucleosome can reduce RNA synthesis in vivo and in vitro, and we analyze its structural basis. We find that the PIC assembles normally when the edge of the +1 nucleosome is located 18 base pairs (bp) downstream of the transcription start site (TSS). However, when the nucleosome edge is located further upstream, only 10 bp downstream of the TSS, the PIC adopts an inhibited state. The transcription factor IIH (TFIIH) shows a closed conformation and its subunit XPB contacts DNA with only one of its two ATPase lobes, inconsistent with DNA opening. These results provide a mechanism for nucleosome-dependent regulation of transcription initiation. 履歴 登録 2022年12月14日 - ヘッダ(付随情報) 公開 2023年5月3日 - マップ公開 2023年5月3日 - 更新 2024年7月24日 - 現状 2024年7月24日 処理サイト : PDBe / 状態 : 公開
すべて表示 表示を減らす
 データを開く
データを開く 基本情報
基本情報
 マップデータ
マップデータ 試料
試料 キーワード
キーワード 機能・相同性情報
機能・相同性情報 Homo sapiens (ヒト) /
Homo sapiens (ヒト) / 
 unidentified adenovirus (ウイルス)
unidentified adenovirus (ウイルス) データ登録者
データ登録者 ドイツ, European Union, 2件
ドイツ, European Union, 2件  引用
引用 ジャーナル: Mol Cell / 年: 2023
ジャーナル: Mol Cell / 年: 2023
 構造の表示
構造の表示 ダウンロードとリンク
ダウンロードとリンク emd_16331.map.gz
emd_16331.map.gz EMDBマップデータ形式
EMDBマップデータ形式 emd-16331-v30.xml
emd-16331-v30.xml emd-16331.xml
emd-16331.xml EMDBヘッダ
EMDBヘッダ emd_16331_fsc.xml
emd_16331_fsc.xml FSCデータファイル
FSCデータファイル emd_16331.png
emd_16331.png emd_16331_msk_1.map
emd_16331_msk_1.map マスクマップ
マスクマップ emd-16331.cif.gz
emd-16331.cif.gz emd_16331_half_map_1.map.gz
emd_16331_half_map_1.map.gz emd_16331_half_map_2.map.gz
emd_16331_half_map_2.map.gz http://ftp.pdbj.org/pub/emdb/structures/EMD-16331
http://ftp.pdbj.org/pub/emdb/structures/EMD-16331 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16331
ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16331


 F&H 検索
F&H 検索 リンク
リンク EMDB (EBI/PDBe) /
EMDB (EBI/PDBe) /  EMDataResource
EMDataResource マップ
マップ ダウンロード / ファイル: emd_16331.map.gz / 形式: CCP4 / 大きさ: 244.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
ダウンロード / ファイル: emd_16331.map.gz / 形式: CCP4 / 大きさ: 244.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) emd_16331_msk_1.map
emd_16331_msk_1.map 試料の構成要素
試料の構成要素 解析
解析 試料調製
試料調製 電子顕微鏡法
電子顕微鏡法 FIELD EMISSION GUN
FIELD EMISSION GUN
 画像解析
画像解析
 ムービー
ムービー コントローラー
コントローラー















































 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)












































 Trichoplusia ni (イラクサキンウワバ)
Trichoplusia ni (イラクサキンウワバ)