[English] 日本語
 Yorodumi
Yorodumi- EMDB-13931: High-resolution structure of the Anaphase-promoting complex/cyclo... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-13931 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | High-resolution structure of the Anaphase-promoting complex/cyclosome (APC/C) bound to co-activator Cdh1 | |||||||||
 Map data Map data | Overall map (sharpened with deepEMhancer) of the Anaphase-promoting complex/cyclosome (APC/C) bound to co-factor Cdh1 at 2.9 A resolution. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | APC/C / cyclosome / Cdc20 / Cdh1 / ubiquitination / Emi1 / mitosis / Cell cycle | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of DNA endoreduplication / positive regulation of biomineral tissue development / negative regulation of meiotic nuclear division / positive regulation of anaphase-promoting complex-dependent catabolic process / negative regulation of mitotic metaphase/anaphase transition / positive regulation of mesenchymal stem cell migration / regulation of meiotic nuclear division / Mitotic Metaphase/Anaphase Transition / positive regulation of synapse maturation / Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase ...negative regulation of DNA endoreduplication / positive regulation of biomineral tissue development / negative regulation of meiotic nuclear division / positive regulation of anaphase-promoting complex-dependent catabolic process / negative regulation of mitotic metaphase/anaphase transition / positive regulation of mesenchymal stem cell migration / regulation of meiotic nuclear division / Mitotic Metaphase/Anaphase Transition / positive regulation of synapse maturation / Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase / regulation of mitotic cell cycle spindle assembly checkpoint / vesicle organization / protein branched polyubiquitination / Inactivation of APC/C via direct inhibition of the APC/C complex / APC/C:Cdc20 mediated degradation of mitotic proteins / Phosphorylation of Emi1 / anaphase-promoting complex / Aberrant regulation of mitotic exit in cancer due to RB1 defects / regulation of meiotic cell cycle / anaphase-promoting complex-dependent catabolic process / metaphase/anaphase transition of mitotic cell cycle / lens fiber cell differentiation / positive regulation of synaptic plasticity / regulation of exit from mitosis / anaphase-promoting complex binding / Phosphorylation of the APC/C / negative regulation of ubiquitin-protein transferase activity / positive regulation of mitotic metaphase/anaphase transition / positive regulation of ubiquitin protein ligase activity / ubiquitin ligase activator activity / spindle assembly involved in female meiosis I / protein K11-linked ubiquitination / meiotic spindle / enzyme-substrate adaptor activity / positive regulation of dendrite morphogenesis / regulation of mitotic metaphase/anaphase transition / oocyte maturation / ubiquitin-ubiquitin ligase activity / negative regulation of ubiquitin protein ligase activity / regulation of mitotic nuclear division / mitotic metaphase chromosome alignment / molecular function inhibitor activity / G1/S-Specific Transcription / regulation of DNA replication / mitotic G2 DNA damage checkpoint signaling / negative regulation of cellular senescence / microtubule polymerization / Regulation of APC/C activators between G1/S and early anaphase / cullin family protein binding / ubiquitin ligase inhibitor activity / Transcriptional Regulation by VENTX / ubiquitin-like ligase-substrate adaptor activity / positive regulation of axon extension / protein K48-linked ubiquitination / positive regulation of osteoblast differentiation / heterochromatin / Cyclin A:Cdk2-associated events at S phase entry / regulation of mitotic cell cycle / positive regulation of G2/M transition of mitotic cell cycle / APC/C:Cdc20 mediated degradation of Cyclin B / APC-Cdc20 mediated degradation of Nek2A / nuclear periphery / Autodegradation of Cdh1 by Cdh1:APC/C / APC/C:Cdc20 mediated degradation of Securin / SCF-beta-TrCP mediated degradation of Emi1 / Assembly of the pre-replicative complex / Cdc20:Phospho-APC/C mediated degradation of Cyclin A / brain development / APC/C:Cdh1 mediated degradation of Cdc20 and other APC/C:Cdh1 targeted proteins in late mitosis/early G1 / mitotic spindle / CDK-mediated phosphorylation and removal of Cdc6 / kinetochore / spindle / ubiquitin-protein transferase activity / Separation of Sister Chromatids / microtubule cytoskeleton / ubiquitin protein ligase activity / Antigen processing: Ubiquitination & Proteasome degradation / mitotic cell cycle / nervous system development / Senescence-Associated Secretory Phenotype (SASP) / ubiquitin-dependent protein catabolic process / protein phosphatase binding / nuclear membrane / molecular adaptor activity / cell differentiation / protein ubiquitination / cell division / negative regulation of gene expression / DNA repair / centrosome / DNA damage response / ubiquitin protein ligase binding / positive regulation of cell population proliferation / nucleolus / protein kinase binding / zinc ion binding / nucleoplasm / nucleus / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||
 Authors Authors | Hoefler A / Yu J | |||||||||
| Funding support |  Switzerland, Switzerland,  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: High-resolution structure of the Anaphase-promoting complex (APC/C) bound to co-activator Cdh1 Authors: Hoefler A / Yu J / Chang L / Zhang Z / Yang J / Boland A / Barford D | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13931.map.gz emd_13931.map.gz | 158.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13931-v30.xml emd-13931-v30.xml emd-13931.xml emd-13931.xml | 45 KB 45 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_13931.png emd_13931.png | 64.2 KB | ||
| Filedesc metadata |  emd-13931.cif.gz emd-13931.cif.gz | 13 KB | ||
| Others |  emd_13931_additional_1.map.gz emd_13931_additional_1.map.gz emd_13931_half_map_1.map.gz emd_13931_half_map_1.map.gz emd_13931_half_map_2.map.gz emd_13931_half_map_2.map.gz | 89.1 MB 165.1 MB 165.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13931 http://ftp.pdbj.org/pub/emdb/structures/EMD-13931 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13931 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13931 | HTTPS FTP |
-Related structure data
| Related structure data |  7qe7  9gawMC  8pkpC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_13931.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13931.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Overall map (sharpened with deepEMhancer) of the Anaphase-promoting complex/cyclosome (APC/C) bound to co-factor Cdh1 at 2.9 A resolution. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Overall map (unsharpened) of the Anaphase-promoting complex/cyclosome (APC/C)...
| File | emd_13931_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Overall map (unsharpened) of the Anaphase-promoting complex/cyclosome (APC/C) bound to co-factor Cdh1 at 2.9 A resolution. | ||||||||||||
| Projections & Slices |
| ||||||||||||
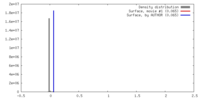
| Density Histograms |
-Half map: Half map B of the Anaphase-promoting complex/cyclosome (APC/C)...
| File | emd_13931_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B of the Anaphase-promoting complex/cyclosome (APC/C) bound to co-factor Cdh1 at 2.9 A resolution. | ||||||||||||
| Projections & Slices |
| ||||||||||||
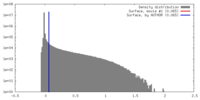
| Density Histograms |
-Half map: Half map A of the Anaphase-promoting complex/cyclosome (APC/C)...
| File | emd_13931_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A of the Anaphase-promoting complex/cyclosome (APC/C) bound to co-factor Cdh1 at 2.9 A resolution. | ||||||||||||
| Projections & Slices |
| ||||||||||||
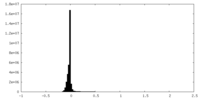
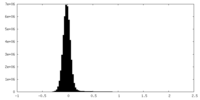
| Density Histograms |
- Sample components
Sample components
+Entire : Anaphase-promoting complex (APC/C) bound to co-activator Cdh1
+Supramolecule #1: Anaphase-promoting complex (APC/C) bound to co-activator Cdh1
+Macromolecule #1: Anaphase-promoting complex subunit 10
+Macromolecule #2: Anaphase-promoting complex subunit 15
+Macromolecule #3: Anaphase-promoting complex subunit 1
+Macromolecule #4: Anaphase-promoting complex subunit 2
+Macromolecule #5: Anaphase-promoting complex subunit 4
+Macromolecule #6: Anaphase-promoting complex subunit 5
+Macromolecule #7: F-box only protein 5
+Macromolecule #8: Cell division cycle protein 16 homolog
+Macromolecule #9: Anaphase-promoting complex subunit CDC26
+Macromolecule #10: Anaphase-promoting complex subunit 13
+Macromolecule #11: Anaphase-promoting complex subunit 16
+Macromolecule #12: Cell division cycle protein 27 homolog
+Macromolecule #13: Anaphase-promoting complex subunit 7
+Macromolecule #14: Cell division cycle protein 23 homolog
+Macromolecule #15: Fizzy-related protein homolog
+Macromolecule #16: Anaphase-promoting complex subunit 11
+Macromolecule #17: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| ||||||||||||
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 50 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 20 sec. / Pretreatment - Atmosphere: OTHER | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number real images: 8297 / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.5 µm |
| Sample stage | Specimen holder model: OTHER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Overall B value: 80 |
| Output model | 
PDB-7qe7:  PDB-9gaw: |
 Movie
Movie Controller
Controller






































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)






































 Trichoplusia ni (cabbage looper)
Trichoplusia ni (cabbage looper)

