+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1364 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Locking and unlocking of ribosomal motions. | |||||||||
 Map data Map data | Cryo-EM map of E.coli 70S ribosome | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationribosome disassembly / translational elongation / translation elongation factor activity / GDP binding / ribosome binding / small ribosomal subunit / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / tRNA binding / rRNA binding ...ribosome disassembly / translational elongation / translation elongation factor activity / GDP binding / ribosome binding / small ribosomal subunit / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / tRNA binding / rRNA binding / structural constituent of ribosome / translation / GTPase activity / GTP binding / magnesium ion binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 10.9 Å | |||||||||
 Authors Authors | Mikel V / Andrey Z / Sengupta J / Rawat U / Ehrenberg M / Frank J | |||||||||
 Citation Citation |  Journal: Cell / Year: 2003 Journal: Cell / Year: 2003Title: Locking and unlocking of ribosomal motions. Authors: Mikel Valle / Andrey Zavialov / Jayati Sengupta / Urmila Rawat / Måns Ehrenberg / Joachim Frank /  Abstract: During the ribosomal translocation, the binding of elongation factor G (EF-G) to the pretranslocational ribosome leads to a ratchet-like rotation of the 30S subunit relative to the 50S subunit in the ...During the ribosomal translocation, the binding of elongation factor G (EF-G) to the pretranslocational ribosome leads to a ratchet-like rotation of the 30S subunit relative to the 50S subunit in the direction of the mRNA movement. By means of cryo-electron microscopy we observe that this rotation is accompanied by a 20 A movement of the L1 stalk of the 50S subunit, implying that this region is involved in the translocation of deacylated tRNAs from the P to the E site. These ribosomal motions can occur only when the P-site tRNA is deacylated. Prior to peptidyl-transfer to the A-site tRNA or peptide removal, the presence of the charged P-site tRNA locks the ribosome and prohibits both of these motions. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1364.map.gz emd_1364.map.gz | 7.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1364-v30.xml emd-1364-v30.xml emd-1364.xml emd-1364.xml | 7.9 KB 7.9 KB | Display Display |  EMDB header EMDB header |
| Images |  1364.gif 1364.gif | 21.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1364 http://ftp.pdbj.org/pub/emdb/structures/EMD-1364 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1364 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1364 | HTTPS FTP |
-Related structure data
| Related structure data |  1pn6MC  1pn7MC  1pn8MC  1362C  1363C  1365C  1366C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_1364.map.gz / Format: CCP4 / Size: 8.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1364.map.gz / Format: CCP4 / Size: 8.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM map of E.coli 70S ribosome | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.82 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
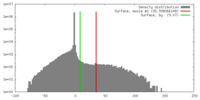
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Release Complex, Puromycin, EF-G, GDPNP
| Entire | Name: Release Complex, Puromycin, EF-G, GDPNP |
|---|---|
| Components |
|
-Supramolecule #1000: Release Complex, Puromycin, EF-G, GDPNP
| Supramolecule | Name: Release Complex, Puromycin, EF-G, GDPNP / type: sample / ID: 1000 / Number unique components: 1 |
|---|
-Macromolecule #1: EF-G
| Macromolecule | Name: EF-G / type: ligand / ID: 1 / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Vitrification | Cryogen name: ETHANE / Instrument: OTHER / Details: Rapid-freezing in liquid ethane |
|---|
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Temperature | Average: 93 K |
| Alignment procedure | Legacy - Electron beam tilt params: 0 |
| Date | Jul 11, 2003 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 14 µm |
| Tilt angle min | 0 |
| Tilt angle max | 0 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder: cryo transfer / Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Details: CTF correctionn of 3D map |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 10.9 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: SPIDER, package / Number images used: 1 |
 Movie
Movie Controller
Controller

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)