+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12296 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Ovine b0,+AT-rBAT heterodimer | |||||||||
 Map data Map data | RELION postprocessed map | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationbroad specificity neutral L-amino acid:basic L-amino acid antiporter activity / L-cystine transmembrane transporter activity / brush border membrane / carbohydrate metabolic process / protein heterodimerization activity / apical plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
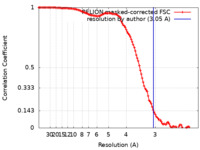
| Method | single particle reconstruction / cryo EM / Resolution: 3.05 Å | |||||||||
 Authors Authors | Lee Y / Kuehlbrandt W | |||||||||
| Funding support |  Germany, 2 items Germany, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Ca-mediated higher-order assembly of heterodimers in amino acid transport system b biogenesis and cystinuria. Authors: Yongchan Lee / Pattama Wiriyasermkul / Pornparn Kongpracha / Satomi Moriyama / Deryck J Mills / Werner Kühlbrandt / Shushi Nagamori /   Abstract: Cystinuria is a genetic disorder characterized by overexcretion of dibasic amino acids and cystine, causing recurrent kidney stones and kidney failure. Mutations of the regulatory glycoprotein rBAT ...Cystinuria is a genetic disorder characterized by overexcretion of dibasic amino acids and cystine, causing recurrent kidney stones and kidney failure. Mutations of the regulatory glycoprotein rBAT and the amino acid transporter bAT, which constitute system b, are linked to type I and non-type I cystinuria respectively and they exhibit distinct phenotypes due to protein trafficking defects or catalytic inactivation. Here, using electron cryo-microscopy and biochemistry, we discover that Ca mediates higher-order assembly of system b. Ca stabilizes the interface between two rBAT molecules, leading to super-dimerization of bAT-rBAT, which in turn facilitates N-glycan maturation and protein trafficking. A cystinuria mutant T216M and mutations of the Ca site of rBAT cause the loss of higher-order assemblies, resulting in protein trapping at the ER and the loss of function. These results provide the molecular basis of system b biogenesis and type I cystinuria and serve as a guide to develop new therapeutic strategies against it. More broadly, our findings reveal an unprecedented link between transporter oligomeric assembly and protein-trafficking diseases. #1:  Journal: Biorxiv / Year: 2021 Journal: Biorxiv / Year: 2021Title: Ca 2+ -mediated higher-order assembly of b 0,+ AT-rBAT is a key step for system b 0,+ biogenesis and cystinuria Authors: Lee Y / Wiriyasermkul P / Moriyama S / Mills DJ / Kuhlbrandt W / Nagamori S | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12296.map.gz emd_12296.map.gz | 10.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12296-v30.xml emd-12296-v30.xml emd-12296.xml emd-12296.xml | 17.9 KB 17.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12296_fsc.xml emd_12296_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_12296.png emd_12296.png | 98.8 KB | ||
| Masks |  emd_12296_msk_1.map emd_12296_msk_1.map | 125 MB |  Mask map Mask map | |
| Others |  emd_12296_half_map_1.map.gz emd_12296_half_map_1.map.gz emd_12296_half_map_2.map.gz emd_12296_half_map_2.map.gz | 98.3 MB 98.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12296 http://ftp.pdbj.org/pub/emdb/structures/EMD-12296 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12296 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12296 | HTTPS FTP |
-Validation report
| Summary document |  emd_12296_validation.pdf.gz emd_12296_validation.pdf.gz | 549.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_12296_full_validation.pdf.gz emd_12296_full_validation.pdf.gz | 548.6 KB | Display | |
| Data in XML |  emd_12296_validation.xml.gz emd_12296_validation.xml.gz | 18.5 KB | Display | |
| Data in CIF |  emd_12296_validation.cif.gz emd_12296_validation.cif.gz | 24.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12296 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12296 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12296 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12296 | HTTPS FTP |
-Related structure data
| Related structure data |  7nf6MC  7nf7C  7nf8C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12296.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12296.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | RELION postprocessed map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.09856 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
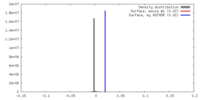
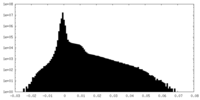
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_12296_msk_1.map emd_12296_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
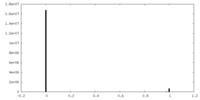
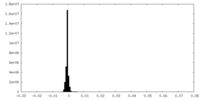
| Density Histograms |
-Half map: Half-map 2
| File | emd_12296_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
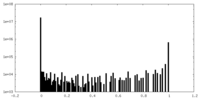
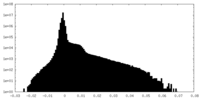
| Density Histograms |
-Half map: Half-map 1
| File | emd_12296_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
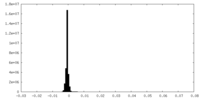
| Density Histograms |
- Sample components
Sample components
-Entire : b0,+AT-rBAT heterodimer
| Entire | Name: b0,+AT-rBAT heterodimer |
|---|---|
| Components |
|
-Supramolecule #1: b0,+AT-rBAT heterodimer
| Supramolecule | Name: b0,+AT-rBAT heterodimer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 135 kDa/nm |
-Macromolecule #1: B(0,+)-type amino acid transporter 1
| Macromolecule | Name: B(0,+)-type amino acid transporter 1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 56.318047 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MEETSLRKRR GDEKSIQSSE PKTTSLQKEL GLFSGTCIIV GTIIGSGIFI SPKSVLSNME AVGPCLIIWA MCGVLATLGA LCFAELGTM ITKSGGEYPY LMEAFGPIPA YLFSWSSLFV IKPSSFAIIC LSFSEYVCAP FYSGCSPPQV VIKTLAAAAI L LISTVNAL ...String: MEETSLRKRR GDEKSIQSSE PKTTSLQKEL GLFSGTCIIV GTIIGSGIFI SPKSVLSNME AVGPCLIIWA MCGVLATLGA LCFAELGTM ITKSGGEYPY LMEAFGPIPA YLFSWSSLFV IKPSSFAIIC LSFSEYVCAP FYSGCSPPQV VIKTLAAAAI L LISTVNAL SVRLGSYVQN VFTAAKMVIV VIIIISGLVL LAQGNTRNFE NSFEGASLSV GSISLALYNG LWAYDGWNQL NY ITEELRN PFRNLPLAII IGIPLVTGCY ILMNVSYFTV MTATELLQSQ AVAVTFGDRV LYPASWIVPL FVAFSTIGAA NGS CFTAGR LVFVAGREGH MLKVLSYISV KRLTPAPAIM FHSMIAIIYI IPGDINSLVN YFSFAAWLFY GLTITGLIVM RFTR KELKR PIKVPIFIPI LVTLLSVFLV LAPIISAPAW EYLYCVLFML SGLVFYFLFV HYKFGWAQKI SKPLTMHLQM LMEVV PPEE APEWSHPQFE KGGGSGGGSG GSAWSHPQFE K |
-Macromolecule #2: neutral and basic amino acid transport protein rBAT
| Macromolecule | Name: neutral and basic amino acid transport protein rBAT / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 78.754281 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GSAEDKSKRD SIGLNAKEGQ TNNGFVQNED ILETDLDPSS PAAGPQHNTV DILGPGEPDV KDVRPYAGMP KEVLFQFSGQ ARYRIPREV LFWLTVASVL LLIAATIAII AISPKCLDWW QAGPMYQIYP RSFRDSNKDG DGDLKGIQDK LDYITTLNIK T VWITSFYK ...String: GSAEDKSKRD SIGLNAKEGQ TNNGFVQNED ILETDLDPSS PAAGPQHNTV DILGPGEPDV KDVRPYAGMP KEVLFQFSGQ ARYRIPREV LFWLTVASVL LLIAATIAII AISPKCLDWW QAGPMYQIYP RSFRDSNKDG DGDLKGIQDK LDYITTLNIK T VWITSFYK SSLKDFRHAV EDFQEIDPIF GTMKDFENLV AAIHDKGLKL IIDFIPNHTS DKHAWFQWSR NRTGKYTDYY IW HDCNYEN GTTIPPNNWL SVYGNSSWHF DEVRKQCYFH QFMKEQPDLN FRNPDVQEEI KEIIQFWLSK GVDGFSFNAL QYL LEAKHL RDEAQVNKTQ IPDTVTHYSQ LHHDFTTTQV GMHDIVRSFR QTMNQYSREP GRYRFMGTEA HGESITETMV YYGL PFIQE ADFPFNSYLS KLDKPSGNSV SEVITSWLEN MPEGKWPNWM TGGPDNVRLT SRLGEKYVNI MNMLVFTLPG TPITY YGEE IGMRNILAAN LNENYDTGTL FSKSPMQWDN SSNAGFSEGN HTWLPTSSDY HTVNVDVQKT QPRSALKLYQ ELSLLH ANE LLLSRGWFCY LRNDNHSIMY TRELDGINKV FLMVLNFGES SLLNLKEMIS NIPTRVRIRL STSSAYSGRE VDTHAVT LA SGEGLILEYN TGNLLHRQTA FKDRCFVSNR ACYSRVLNIL YSLC |
-Macromolecule #4: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 4 / Number of copies: 2 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Macromolecule #5: 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine
| Macromolecule | Name: 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine / type: ligand / ID: 5 / Number of copies: 1 / Formula: LBN |
|---|---|
| Molecular weight | Theoretical: 760.076 Da |
| Chemical component information |  ChemComp-LBN: |
-Macromolecule #6: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 6 / Number of copies: 1 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #7: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 7 / Number of copies: 4 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)