+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12192 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Salmonella export gate and rod refined in focussed C1 map | |||||||||||||||
 Map data Map data | post processed volume | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | bacterial flagellum rod bacterial flagellum export gate basal body / PROTEIN TRANSPORT | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationbacterial-type flagellum basal body, distal rod / bacterial-type flagellum basal body, rod / bacterial-type flagellum organization / bacterial-type flagellum basal body / bacterial-type flagellum-dependent swarming motility / bacterial-type flagellum / cytoskeletal motor activity / bacterial-type flagellum assembly / protein secretion / bacterial-type flagellum-dependent cell motility ...bacterial-type flagellum basal body, distal rod / bacterial-type flagellum basal body, rod / bacterial-type flagellum organization / bacterial-type flagellum basal body / bacterial-type flagellum-dependent swarming motility / bacterial-type flagellum / cytoskeletal motor activity / bacterial-type flagellum assembly / protein secretion / bacterial-type flagellum-dependent cell motility / protein targeting / structural molecule activity / plasma membrane Similarity search - Function | |||||||||||||||
| Biological species |  Salmonella enterica subsp. enterica serovar Typhi (bacteria) / Salmonella enterica subsp. enterica serovar Typhi (bacteria) /  Salmonella typhi (bacteria) Salmonella typhi (bacteria) | |||||||||||||||
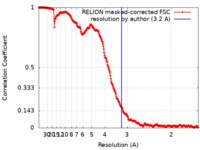
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||||||||
 Authors Authors | Johnson S / Furlong E | |||||||||||||||
| Funding support |  United Kingdom, 4 items United Kingdom, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Microbiol / Year: 2021 Journal: Nat Microbiol / Year: 2021Title: Molecular structure of the intact bacterial flagellar basal body. Authors: Steven Johnson / Emily J Furlong / Justin C Deme / Ashley L Nord / Joseph J E Caesar / Fabienne F V Chevance / Richard M Berry / Kelly T Hughes / Susan M Lea /    Abstract: The bacterial flagellum is a macromolecular protein complex that enables motility in many species. Bacterial flagella self-assemble a strong, multicomponent drive shaft that couples rotation in the ...The bacterial flagellum is a macromolecular protein complex that enables motility in many species. Bacterial flagella self-assemble a strong, multicomponent drive shaft that couples rotation in the inner membrane to the micrometre-long flagellar filament that powers bacterial swimming in viscous fluids. Here, we present structures of the intact Salmonella flagellar basal body, encompassing the inner membrane rotor, drive shaft and outer-membrane bushing, solved using cryo-electron microscopy to resolutions of 2.2-3.7 Å. The structures reveal molecular details of how 173 protein molecules of 13 different types assemble into a complex spanning two membranes and a cell wall. The helical drive shaft at one end is intricately interwoven with the rotor component with both the export gate complex and the proximal rod forming interactions with the MS-ring. At the other end, the drive shaft distal rod passes through the LP-ring bushing complex, which functions as a molecular bearing anchored in the outer membrane through interactions with the lipopolysaccharide. The in situ structure of a protein complex capping the drive shaft provides molecular insights into the assembly process of this molecular machine. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12192.map.gz emd_12192.map.gz | 92.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12192-v30.xml emd-12192-v30.xml emd-12192.xml emd-12192.xml | 24.8 KB 24.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12192_fsc.xml emd_12192_fsc.xml | 27 KB | Display |  FSC data file FSC data file |
| Images |  emd_12192.png emd_12192.png | 24.1 KB | ||
| Masks |  emd_12192_msk_1.map emd_12192_msk_1.map | 1.7 GB |  Mask map Mask map | |
| Filedesc metadata |  emd-12192.cif.gz emd-12192.cif.gz | 6.3 KB | ||
| Others |  emd_12192_additional_1.map.gz emd_12192_additional_1.map.gz emd_12192_half_map_1.map.gz emd_12192_half_map_1.map.gz emd_12192_half_map_2.map.gz emd_12192_half_map_2.map.gz | 1.4 GB 1.4 GB 1.4 GB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12192 http://ftp.pdbj.org/pub/emdb/structures/EMD-12192 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12192 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12192 | HTTPS FTP |
-Related structure data
| Related structure data |  7binMC  7bglC  7bhqC  7bj2C  7bk0C  7nvgC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12192.map.gz / Format: CCP4 / Size: 1.7 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12192.map.gz / Format: CCP4 / Size: 1.7 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | post processed volume | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.832 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_12192_msk_1.map emd_12192_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
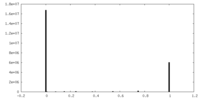
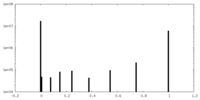
| Density Histograms |
-Additional map: refinement volume
| File | emd_12192_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | refinement volume | ||||||||||||
| Projections & Slices |
| ||||||||||||
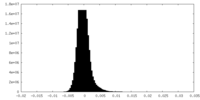
| Density Histograms |
-Half map: half map 2
| File | emd_12192_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
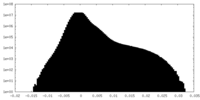
| Density Histograms |
-Half map: half map 1
| File | emd_12192_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
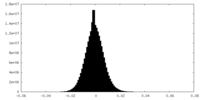
| Density Histograms |
- Sample components
Sample components
-Entire : Flagellar LP ring
| Entire | Name: Flagellar LP ring |
|---|---|
| Components |
|
-Supramolecule #1: Flagellar LP ring
| Supramolecule | Name: Flagellar LP ring / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhi (bacteria) Salmonella enterica subsp. enterica serovar Typhi (bacteria) |
-Macromolecule #1: Flagellar biosynthetic protein FliP
| Macromolecule | Name: Flagellar biosynthetic protein FliP / type: protein_or_peptide / ID: 1 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella typhi (bacteria) Salmonella typhi (bacteria) |
| Molecular weight | Theoretical: 26.801086 KDa |
| Sequence | String: MRRLLFLSLA GLWLFSPAAA AQLPGLISQP LAGGGQSWSL SVQTLVFITS LTFLPAILLM MTSFTRIIIV FGLLRNALGT PSAPPNQVL LGLALFLTFF IMSPVIDKIY VDAYQPFSEQ KISMQEALDK GAQPLRAFML RQTREADLAL FARLANSGPL Q GPEAVPMR ...String: MRRLLFLSLA GLWLFSPAAA AQLPGLISQP LAGGGQSWSL SVQTLVFITS LTFLPAILLM MTSFTRIIIV FGLLRNALGT PSAPPNQVL LGLALFLTFF IMSPVIDKIY VDAYQPFSEQ KISMQEALDK GAQPLRAFML RQTREADLAL FARLANSGPL Q GPEAVPMR ILLPAYVTSE LKTAFQIGFT IFIPFLIIDL VIASVLMALG MMMVPPATIA LPFKLMLFVL VDGWQLLMGS LA QSFYS UniProtKB: Flagellar biosynthetic protein FliP |
-Macromolecule #2: Flagellar biosynthetic protein FliR
| Macromolecule | Name: Flagellar biosynthetic protein FliR / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhi (bacteria) Salmonella enterica subsp. enterica serovar Typhi (bacteria) |
| Molecular weight | Theoretical: 28.938865 KDa |
| Sequence | String: MIQVTSEQWL YWLHLYFWPL LRVLALISTA PILSERAIPK RVKLGLGIMI TLVIAPSLPA NDTPLFSIAA LWLAMQQILI GIALGFTMQ FAFAAVRTAG EFIGLQMGLS FATFVDPGSH LNMPVLARIM DMLAMLLFLT FNGHLWLISL LVDTFHTLPI G SNPVNSNA ...String: MIQVTSEQWL YWLHLYFWPL LRVLALISTA PILSERAIPK RVKLGLGIMI TLVIAPSLPA NDTPLFSIAA LWLAMQQILI GIALGFTMQ FAFAAVRTAG EFIGLQMGLS FATFVDPGSH LNMPVLARIM DMLAMLLFLT FNGHLWLISL LVDTFHTLPI G SNPVNSNA FMALARAGGL IFLNGLMLAL PVITLLLTLN LALGLLNRMA PQLSIFVIGF PLTLTVGIML MAALMPLIAP FC EHLFSEI FNLLADIVSE MPINNNP UniProtKB: Flagellar biosynthetic protein FliR |
-Macromolecule #3: Flagellar biosynthetic protein FliQ
| Macromolecule | Name: Flagellar biosynthetic protein FliQ / type: protein_or_peptide / ID: 3 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhi (bacteria) Salmonella enterica subsp. enterica serovar Typhi (bacteria) |
| Molecular weight | Theoretical: 9.606758 KDa |
| Sequence | String: MTPESVMMMG TEAMKVALAL AAPLLLVALI TGLIISILQA ATQINEMTLS FIPKIVAVFI AIIVAGPWML NLLLDYVRTL FSNLPYIIG UniProtKB: Flagellar biosynthetic protein FliQ |
-Macromolecule #4: Flagellar hook-basal body complex protein FliE
| Macromolecule | Name: Flagellar hook-basal body complex protein FliE / type: protein_or_peptide / ID: 4 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhi (bacteria) Salmonella enterica subsp. enterica serovar Typhi (bacteria)Strain: LT2 / SGSC1412 / ATCC 700720 |
| Molecular weight | Theoretical: 11.087662 KDa |
| Sequence | String: MAAIQGIEGV ISQLQATAMA ARGQDTHSQS TVSFAGQLHA ALDRISDRQA AARVQAEKFT LGEPGIALND VMADMQKASV SMQMGIQVR NKLVAAYQEV MSMQV UniProtKB: Flagellar hook-basal body complex protein FliE |
-Macromolecule #5: Flagellar basal body rod protein FlgB
| Macromolecule | Name: Flagellar basal body rod protein FlgB / type: protein_or_peptide / ID: 5 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhi (bacteria) Salmonella enterica subsp. enterica serovar Typhi (bacteria)Strain: LT2 / SGSC1412 / ATCC 700720 |
| Molecular weight | Theoretical: 15.145061 KDa |
| Sequence | String: MLDRLDAALR FQQEALNLRA QRQEILAANI ANADTPGYQA RDIDFASELK KVMVRGREET GGVALTLTSS HHIPAQAVSS PAVDLLYRV PDQPSLDGNT VDMDRERTQF ADNSLKYQMG LTVLGSQLKG MMNVLQGGN UniProtKB: Flagellar basal body rod protein FlgB |
-Macromolecule #6: Flagellar basal-body rod protein FlgC
| Macromolecule | Name: Flagellar basal-body rod protein FlgC / type: protein_or_peptide / ID: 6 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhi (bacteria) Salmonella enterica subsp. enterica serovar Typhi (bacteria)Strain: LT2 / SGSC1412 / ATCC 700720 |
| Molecular weight | Theoretical: 13.991889 KDa |
| Sequence | String: MALLNIFDIA GSALAAQSKR LNVAASNLAN ADSVTGPDGQ PYRAKQVVFQ VDAAPGQATG GVKVASVIES QAPEKLVYEP GNPLADANG YVKMPNVDVV GEMVNTMSAS RSYQANIEVL NTVKSMMLKT LTLGQ UniProtKB: Flagellar basal-body rod protein FlgC |
-Macromolecule #7: Flagellar basal-body rod protein FlgF
| Macromolecule | Name: Flagellar basal-body rod protein FlgF / type: protein_or_peptide / ID: 7 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhi (bacteria) Salmonella enterica subsp. enterica serovar Typhi (bacteria)Strain: LT2 / SGSC1412 / ATCC 700720 |
| Molecular weight | Theoretical: 26.121223 KDa |
| Sequence | String: MDHAIYTAMG AASQTLNQQA VTASNLANAS TPGFRAQLNA LRAVPVDGLS LATRTLVTAS TPGADMTPGQ LDYTSRPLDV ALQQDGWLV VQAADGAEGY TRNGNIQVGP TGQLTIQGHP VIGEGGPITV PEGSEITIAA DGTISALNPG DPPNTVAPVG R LKLVKAEG ...String: MDHAIYTAMG AASQTLNQQA VTASNLANAS TPGFRAQLNA LRAVPVDGLS LATRTLVTAS TPGADMTPGQ LDYTSRPLDV ALQQDGWLV VQAADGAEGY TRNGNIQVGP TGQLTIQGHP VIGEGGPITV PEGSEITIAA DGTISALNPG DPPNTVAPVG R LKLVKAEG NEVQRSDDGL FRLTAEAQAE RGAVLAADPS IRIMSGVLEG SNVKPVEAMT DMIANARRFE MQMKVITSVD EN EGRANQL LSMS UniProtKB: Flagellar basal-body rod protein FlgF |
-Macromolecule #8: Flagellar basal-body rod protein FlgG
| Macromolecule | Name: Flagellar basal-body rod protein FlgG / type: protein_or_peptide / ID: 8 / Number of copies: 24 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhi (bacteria) Salmonella enterica subsp. enterica serovar Typhi (bacteria)Strain: LT2 / SGSC1412 / ATCC 700720 |
| Molecular weight | Theoretical: 27.784807 KDa |
| Sequence | String: MISSLWIAKT GLDAQQTNMD VIANNLANVS TNGFKRQRAV FEDLLYQTIR QPGAQSSEQT TLPSGLQIGT GVRPVATERL HSQGNLSQT NNSKDVAIKG QGFFQVMLPD GTSAYTRDGS FQVDQNGQLV TAGGFQVQPA ITIPANALSI TIGRDGVVSV T QQGQAAPV ...String: MISSLWIAKT GLDAQQTNMD VIANNLANVS TNGFKRQRAV FEDLLYQTIR QPGAQSSEQT TLPSGLQIGT GVRPVATERL HSQGNLSQT NNSKDVAIKG QGFFQVMLPD GTSAYTRDGS FQVDQNGQLV TAGGFQVQPA ITIPANALSI TIGRDGVVSV T QQGQAAPV QVGQLNLTTF MNDTGLESIG ENLYIETQSS GAPNESTPGL NGAGLLYQGY VETSNVNVAE ELVNMIQVQR AY EINSKAV STTDQMLQKL TQL UniProtKB: Flagellar basal-body rod protein FlgG |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 59.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)