+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12183 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Salmonella LP ring 26 mer refined in C26 map | |||||||||||||||
 Map data Map data | refinement map | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | bacterial flagellum LP ring salmonella / MEMBRANE PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationbacterial-type flagellum basal body, distal rod, L ring / bacterial-type flagellum basal body, distal rod, P ring / cytoskeletal motor activity / bacterial-type flagellum-dependent cell motility / cell outer membrane / outer membrane-bounded periplasmic space / structural molecule activity Similarity search - Function | |||||||||||||||
| Biological species |  Salmonella enterica subsp. enterica serovar Typhi (bacteria) / Salmonella enterica subsp. enterica serovar Typhi (bacteria) /  Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) (bacteria) Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) (bacteria) | |||||||||||||||
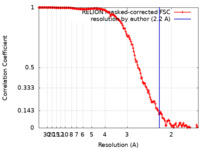
| Method | single particle reconstruction / cryo EM / Resolution: 2.2 Å | |||||||||||||||
 Authors Authors | Johnson S / Furlong E | |||||||||||||||
| Funding support |  United Kingdom, 4 items United Kingdom, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Microbiol / Year: 2021 Journal: Nat Microbiol / Year: 2021Title: Molecular structure of the intact bacterial flagellar basal body. Authors: Steven Johnson / Emily J Furlong / Justin C Deme / Ashley L Nord / Joseph J E Caesar / Fabienne F V Chevance / Richard M Berry / Kelly T Hughes / Susan M Lea /    Abstract: The bacterial flagellum is a macromolecular protein complex that enables motility in many species. Bacterial flagella self-assemble a strong, multicomponent drive shaft that couples rotation in the ...The bacterial flagellum is a macromolecular protein complex that enables motility in many species. Bacterial flagella self-assemble a strong, multicomponent drive shaft that couples rotation in the inner membrane to the micrometre-long flagellar filament that powers bacterial swimming in viscous fluids. Here, we present structures of the intact Salmonella flagellar basal body, encompassing the inner membrane rotor, drive shaft and outer-membrane bushing, solved using cryo-electron microscopy to resolutions of 2.2-3.7 Å. The structures reveal molecular details of how 173 protein molecules of 13 different types assemble into a complex spanning two membranes and a cell wall. The helical drive shaft at one end is intricately interwoven with the rotor component with both the export gate complex and the proximal rod forming interactions with the MS-ring. At the other end, the drive shaft distal rod passes through the LP-ring bushing complex, which functions as a molecular bearing anchored in the outer membrane through interactions with the lipopolysaccharide. The in situ structure of a protein complex capping the drive shaft provides molecular insights into the assembly process of this molecular machine. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12183.map.gz emd_12183.map.gz | 241.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12183-v30.xml emd-12183-v30.xml emd-12183.xml emd-12183.xml | 19.7 KB 19.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12183_fsc.xml emd_12183_fsc.xml | 15.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_12183.png emd_12183.png | 77.1 KB | ||
| Filedesc metadata |  emd-12183.cif.gz emd-12183.cif.gz | 5.7 KB | ||
| Others |  emd_12183_additional_1.map.gz emd_12183_additional_1.map.gz emd_12183_half_map_1.map.gz emd_12183_half_map_1.map.gz emd_12183_half_map_2.map.gz emd_12183_half_map_2.map.gz | 34.2 MB 242.7 MB 242.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12183 http://ftp.pdbj.org/pub/emdb/structures/EMD-12183 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12183 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12183 | HTTPS FTP |
-Related structure data
| Related structure data |  7bglMC  7bhqC  7binC  7bj2C  7bk0C  7nvgC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12183.map.gz / Format: CCP4 / Size: 307.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12183.map.gz / Format: CCP4 / Size: 307.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | refinement map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.832 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: post processed
| File | emd_12183_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | post processed | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 1
| File | emd_12183_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2
| File | emd_12183_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Flagellar LP ring
| Entire | Name: Flagellar LP ring |
|---|---|
| Components |
|
-Supramolecule #1: Flagellar LP ring
| Supramolecule | Name: Flagellar LP ring / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhi (bacteria) Salmonella enterica subsp. enterica serovar Typhi (bacteria) |
-Macromolecule #1: Flagellar L-ring protein
| Macromolecule | Name: Flagellar L-ring protein / type: protein_or_peptide / ID: 1 / Number of copies: 26 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) (bacteria) Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) (bacteria) |
| Molecular weight | Theoretical: 24.726666 KDa |
| Sequence | String: MQKYALHAYP VMALMVATLT GCAWIPAKPL VQGATTAQPI PGPVPVANGS IFQSAQPINY GYQPLFEDRR PRNIGDTLTI VLQENVSAS KSSSANASRD GKTSFGFDTV PRYLQGLFGN SRADMEASGG NSFNGKGGAN ASNTFSGTLT VTVDQVLANG N LHVVGEKQ ...String: MQKYALHAYP VMALMVATLT GCAWIPAKPL VQGATTAQPI PGPVPVANGS IFQSAQPINY GYQPLFEDRR PRNIGDTLTI VLQENVSAS KSSSANASRD GKTSFGFDTV PRYLQGLFGN SRADMEASGG NSFNGKGGAN ASNTFSGTLT VTVDQVLANG N LHVVGEKQ IAINQGTEFI RFSGVVNPRT ISGSNSVPST QVADARIEYV GNGYINEAQN MGWLQRFFLN LSPM UniProtKB: Flagellar L-ring protein |
-Macromolecule #2: Flagellar P-ring protein
| Macromolecule | Name: Flagellar P-ring protein / type: protein_or_peptide / ID: 2 / Number of copies: 26 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) (bacteria) Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) (bacteria) |
| Molecular weight | Theoretical: 38.194176 KDa |
| Sequence | String: MFKALAGIVL ALVATLAHAE RIRDLTSVQG VRENSLIGYG LVVGLDGTGD QTTQTPFTTQ TLNNMLSQLG ITVPTGTNMQ LKNVAAVMV TASYPPFARQ GQTIDVVVSS MGNAKSLRGG TLLMTPLKGV DSQVYALAQG NILVGGAGAS AGGSSVQVNQ L NGGRITNG ...String: MFKALAGIVL ALVATLAHAE RIRDLTSVQG VRENSLIGYG LVVGLDGTGD QTTQTPFTTQ TLNNMLSQLG ITVPTGTNMQ LKNVAAVMV TASYPPFARQ GQTIDVVVSS MGNAKSLRGG TLLMTPLKGV DSQVYALAQG NILVGGAGAS AGGSSVQVNQ L NGGRITNG AIIERELPTQ FGAGNTINLQ LNDEDFTMAQ QITDAINRAR GYGSATALDA RTVQVRVPSG NSSQVRFLAD IQ NMEVNVT PQDAKVVINS RTGSVVMNRE VTLDSCAVAQ GNLSVTVNRQ LNVNQPNTPF GGGQTVVTPQ TQIDLRQSGG SLQ SVRSSA NLNSVVRALN ALGATPMDLM SILQSMQSAG CLRAKLEII UniProtKB: Flagellar P-ring protein |
-Macromolecule #3: YecR
| Macromolecule | Name: YecR / type: protein_or_peptide / ID: 3 / Number of copies: 26 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) (bacteria) Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) (bacteria) |
| Molecular weight | Theoretical: 12.194968 KDa |
| Sequence | String: MKSLIFTLSL LALTGCTITR QAQVSEASPI SGIVRLTYNQ PLFFTSRTDD YVSHGTATRE CQQMGYADAV SFGQPVGTCS IYAGSLCLN TRFTLSWQCR GVAVPQIMPL YY UniProtKB: Outer membrane lipoprotein |
-Macromolecule #4: (2~{R},4~{R},5~{R},6~{R})-6-[(1~{R})-1,2-bis(oxidanyl)ethyl]-4,5-...
| Macromolecule | Name: (2~{R},4~{R},5~{R},6~{R})-6-[(1~{R})-1,2-bis(oxidanyl)ethyl]-4,5-bis(oxidanyl)oxane-2-carboxylic acid type: ligand / ID: 4 / Number of copies: 26 / Formula: TLW |
|---|---|
| Molecular weight | Theoretical: 222.193 Da |
| Chemical component information |  ChemComp-TLW: |
-Macromolecule #5: [(3~{R})-1-[[(2~{R},3~{R},4~{R},5~{S},6~{R})-6-[[(2~{R},3~{R},4~{...
| Macromolecule | Name: [(3~{R})-1-[[(2~{R},3~{R},4~{R},5~{S},6~{R})-6-[[(2~{R},3~{R},4~{R},5~{S},6~{R})-3-[[(3~{R})-3-dodecanoyloxytetradecanoyl]amino]-6-(hydroxymethyl)-5-phosphonooxy-4-[(3~{R})-3- ...Name: [(3~{R})-1-[[(2~{R},3~{R},4~{R},5~{S},6~{R})-6-[[(2~{R},3~{R},4~{R},5~{S},6~{R})-3-[[(3~{R})-3-dodecanoyloxytetradecanoyl]amino]-6-(hydroxymethyl)-5-phosphonooxy-4-[(3~{R})-3-tetradecanoyloxytetradecanoyl]oxy-oxan-2-yl]oxymethyl]-5-oxidanyl-4-[(3~{R})-3-oxidanyltetradecanoyl]oxy-2-phosphonooxy-oxan-3-yl]amino]-1-oxidanylidene-tetradecan-3-yl] hexadecanoate type: ligand / ID: 5 / Number of copies: 26 / Formula: TQN |
|---|---|
| Molecular weight | Theoretical: 2.036774 KDa |
| Chemical component information |  ChemComp-TQN: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 59.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)