[English] 日本語
 Yorodumi
Yorodumi- EMDB-11606: Cryo-EM map of the large glutamate dehydrogenase composed of 180 ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11606 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM map of the large glutamate dehydrogenase composed of 180 kDa subunits from Mycobacterium smegmatis (open conformation) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | large glutamate dehydrogenase Mycobacterium metabolism / OXIDOREDUCTASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationglutamate dehydrogenase / L-glutamate dehydrogenase (NAD+) activity / L-glutamate catabolic process / L-aspartate:2-oxoglutarate transaminase activity Similarity search - Function | |||||||||
| Biological species |  Mycolicibacterium smegmatis MC2 155 (bacteria) Mycolicibacterium smegmatis MC2 155 (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.19 Å | |||||||||
 Authors Authors | Lazaro M / Melero R | |||||||||
| Funding support |  Spain, 1 items Spain, 1 items
| |||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2021 Journal: Commun Biol / Year: 2021Title: 3D architecture and structural flexibility revealed in the subfamily of large glutamate dehydrogenases by a mycobacterial enzyme. Authors: Melisa Lázaro / Roberto Melero / Charlotte Huet / Jorge P López-Alonso / Sandra Delgado / Alexandra Dodu / Eduardo M Bruch / Luciano A Abriata / Pedro M Alzari / Mikel Valle / María-Natalia Lisa /     Abstract: Glutamate dehydrogenases (GDHs) are widespread metabolic enzymes that play key roles in nitrogen homeostasis. Large glutamate dehydrogenases composed of 180 kDa subunits (L-GDHs) contain long N- ...Glutamate dehydrogenases (GDHs) are widespread metabolic enzymes that play key roles in nitrogen homeostasis. Large glutamate dehydrogenases composed of 180 kDa subunits (L-GDHs) contain long N- and C-terminal segments flanking the catalytic core. Despite the relevance of L-GDHs in bacterial physiology, the lack of structural data for these enzymes has limited the progress of functional studies. Here we show that the mycobacterial L-GDH (mL-GDH) adopts a quaternary structure that is radically different from that of related low molecular weight enzymes. Intersubunit contacts in mL-GDH involve a C-terminal domain that we propose as a new fold and a flexible N-terminal segment comprising ACT-like and PAS-type domains that could act as metabolic sensors for allosteric regulation. These findings uncover unique aspects of the structure-function relationship in the subfamily of L-GDHs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11606.map.gz emd_11606.map.gz | 13.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11606-v30.xml emd-11606-v30.xml emd-11606.xml emd-11606.xml | 25 KB 25 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11606_fsc.xml emd_11606_fsc.xml | 13.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_11606.png emd_11606.png | 236.6 KB | ||
| Masks |  emd_11606_msk_1.map emd_11606_msk_1.map | 209.3 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-11606.cif.gz emd-11606.cif.gz | 8.4 KB | ||
| Others |  emd_11606_half_map_1.map.gz emd_11606_half_map_1.map.gz emd_11606_half_map_2.map.gz emd_11606_half_map_2.map.gz | 164.2 MB 164.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11606 http://ftp.pdbj.org/pub/emdb/structures/EMD-11606 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11606 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11606 | HTTPS FTP |
-Related structure data
| Related structure data |  7a1dMC  7jsrC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_11606.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11606.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
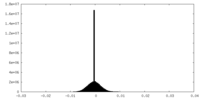
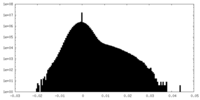
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_11606_msk_1.map emd_11606_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
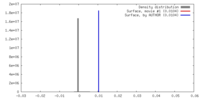
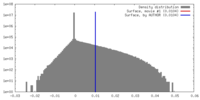
| Density Histograms |
-Half map: #2
| File | emd_11606_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_11606_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM map of the large glutamate dehydrogenase composed of 180 ...
| Entire | Name: Cryo-EM map of the large glutamate dehydrogenase composed of 180 kDa subunits from Mycobacterium smegmatis (open conformation) |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM map of the large glutamate dehydrogenase composed of 180 ...
| Supramolecule | Name: Cryo-EM map of the large glutamate dehydrogenase composed of 180 kDa subunits from Mycobacterium smegmatis (open conformation) type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Mycolicibacterium smegmatis MC2 155 (bacteria) Mycolicibacterium smegmatis MC2 155 (bacteria) |
| Molecular weight | Theoretical: 705 KDa |
-Macromolecule #1: NAD-specific glutamate dehydrogenase
| Macromolecule | Name: NAD-specific glutamate dehydrogenase / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO / EC number: glutamate dehydrogenase |
|---|---|
| Source (natural) | Organism:  Mycolicibacterium smegmatis MC2 155 (bacteria) / Strain: ATCC 700084 / mc(2)155 Mycolicibacterium smegmatis MC2 155 (bacteria) / Strain: ATCC 700084 / mc(2)155 |
| Molecular weight | Theoretical: 176.341859 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MHHHHHHENL YFQGAASMIR RLSVAFLSTY RGPQADAPGV TSTGPLAVAA HDDLVSDDLV AAHYRLASMR APGETKAAVY PGDAGSGAA LQIVTDQAPM LVDSVTVLLH RHGIAYTAIM NPVFRVRRGL DGELLDVRPA AEAAPGDGAD ECWILVPITA A ADGEALTE ...String: MHHHHHHENL YFQGAASMIR RLSVAFLSTY RGPQADAPGV TSTGPLAVAA HDDLVSDDLV AAHYRLASMR APGETKAAVY PGDAGSGAA LQIVTDQAPM LVDSVTVLLH RHGIAYTAIM NPVFRVRRGL DGELLDVRPA AEAAPGDGAD ECWILVPITA A ADGEALTE ATRLVPGILA EARQIGLDSG AMIAALHGLA NDLATDLEGH FPNAERKEVA ALLRWLADGH FVLLGYQQCV VG DGNAEVD PASRLGVLRL RNDVLPPLTD SDDLLVLAQA TMPSYLRYGA YPYIVVVRES PGASRVIEHR FVGLFTVAAM NAN ALEIPL ISRRVEEALA MAHRDPSHPG QLLRDIIQTI PRPELFALSS KQLLEMALAV VDLGSRRRTL LFLRADHLAH FVSC LVYLP RDRYTTAVRL EMQDILVREL GGAGIDYSAR VSESPWAVVH FTVRLPEGTA ADSVDTSLEN ESRIQDLLTE ATRNW GDRM ISAAAAASIS PAALEHYAHA FPEDYKQAFA PQDAIADISL IEALQDDSVK LVLADTAEDR VWKLTWYLGG HSASLS ELL PMLQSMGVVV LEERPFTLRR TDGLPVWIYQ FKISPHPSIP HAPDAEAQRD TAQRFADAVT AIWHGRVEID RFNELVM RA GLTWQQVVVL RAYAKYLRQA GFPYSQSHIE SVLNENPHTT RSLIDLFEAL FDPSQETDGR RDAQGAAAAV AADIDALV S LDTDRVLRAF ANLIEATLRT NYFVARPDSA RARNVLAFKL NPLVIKELPL PRPKFEIFVY SPRVEGVHLR FGFVARGGL RWSDRREDFR TEILGLVKAQ AVKNAVIVPV GAKGGFVVKR PPTLTGDAAA DREATRAEGV ECYRLFISGL LDVTDNVDKA TGAVVTPPE VVRRDGEDAY LVVAADKGTA TFSDIANEVA KSYGFWLGDA FASGGSIGYD HKAMGITAKG AWESVKRHFR E MGVDTQTQ DFTVVGIGDM SGDVFGNGML LSKHIRLVAA FDHRDIFLDP NPDAGRSWDE RKRLFDLPRS SWADYDKSLI SE GGGVYSR QQKSIPISPQ VRTALGLDAD VEELTPPALI KAILKAPVDL LWNGGIGTYI KAETEADADV GDRANDQIRV CGN QVRAKV IGEGGNLGVT ALGRIEFDLA GGRINTDALD NSAGVDCSDH EVNIKILIDS AVTAGKVTPE ERTELLLSMT DEVG ELVLA DNRDQNDLMG TSRANAASLL SVHARMIKDL VDNRGLNREL EALPSEKEIR RRADAGIGLT SPELATLMAH VKLAL KDDV LASDLPDQEV FASRLPYYFP TRLREELHGE IRSHQLRREI ITTMLVNDLV DTAGISYAYR ITEDVGVGPV DAVRSY VAI NAIFGIGDVW RRIRAAGDAG VPTSVTDRMT LDLRRLVDRA GRWLLNYRPQ PLAVGAEINR FGAKVAALTP RMSEWLR GD DKAIVSKEAG DFASHGVPED LAYHIATGLY QYSLLDVIDI ADIVDREPDE VADTYFALMD HLGADALLTA VSRLSRDD R WHSLARLAIR DDIYGSLRAL CFDVLAVGEP DENGEEKIAE WETTNSSRVT RARRTLTEIY KDGEQDLATL SVAARQIRS MTRTSGTGTT G UniProtKB: NAD-specific glutamate dehydrogenase |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL |
|---|---|
| Buffer | pH: 6 |
| Grid | Model: Quantifoil / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Frames/image: 1-20 / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated defocus max: 3.2600000000000002 µm / Calibrated defocus min: 0.67 µm / Calibrated magnification: 47170 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.2600000000000002 µm / Nominal defocus min: 0.67 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Model fitting into the cryo-EM map was performed using the programs UCSF Chimera (Pettersen et al., 2004), Namdinator (Kidmose et al., 2019), phenix.real_space_refine (Afonine et al., 2018) and Coot (Emsley, Lohkamp, Scott, & Cowtan, 2010). Residues 500-1588 from the crystal structure of Se-Met mL-GDH180 (PDB code 7JSR) were fitted into the cryo-EM map. Se-methionine residues were replaced by methionine residues using Coot (Emsley et al., 2010) and the model was finally refined employing phenix.real_space_refine (Afonine et al., 2018) with NCS and secondary structure restraints. |
|---|---|
| Refinement | Protocol: OTHER |
| Output model |  PDB-7a1d: |
 Movie
Movie Controller
Controller
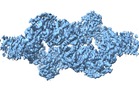










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)