[English] 日本語
 Yorodumi
Yorodumi- EMDB-10735: Cryo-EM structure of trimeric human STEAP1 bound to three Fab120.... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10735 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of trimeric human STEAP1 bound to three Fab120.545 fragments | |||||||||
 Map data Map data | Unsharpened cryo-EM map of homotrimeric STEAP1 bound to three Fab-fragments of antibody mAb120.545. The constant region of the Fabs is masked out. | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationcell-cell junction / endosome membrane / endosome / heme binding / membrane / metal ion binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /   | |||||||||
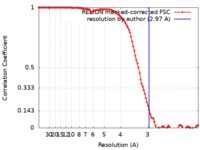
| Method | single particle reconstruction / cryo EM / Resolution: 2.97 Å | |||||||||
 Authors Authors | Oosterheert W / Gros P | |||||||||
| Funding support |  Netherlands, 1 items Netherlands, 1 items
| |||||||||
 Citation Citation |  Journal: J Biol Chem / Year: 2020 Journal: J Biol Chem / Year: 2020Title: Cryo-electron microscopy structure and potential enzymatic function of human six-transmembrane epithelial antigen of the prostate 1 (STEAP1). Authors: Wout Oosterheert / Piet Gros /  Abstract: Six-transmembrane epithelial antigen of the prostate 1 (STEAP1) is an integral membrane protein that is highly up-regulated on the cell surface of several human cancers, making it a promising ...Six-transmembrane epithelial antigen of the prostate 1 (STEAP1) is an integral membrane protein that is highly up-regulated on the cell surface of several human cancers, making it a promising therapeutic target to manage these diseases. It shares sequence homology with three enzymes (STEAP2-STEAP4) that catalyze the NADPH-dependent reduction of iron(III). However, STEAP1 lacks an intracellular NADPH-binding domain and does not exhibit cellular ferric reductase activity. Thus, both the molecular function of STEAP1 and its role in cancer progression remain elusive. Here, we present a ∼3.0-Å cryo-EM structure of trimeric human STEAP1 bound to three antigen-binding fragments (Fabs) of the clinically used antibody mAb120.545. The structure revealed that STEAP1 adopts a reductase-like conformation and interacts with the Fabs through its extracellular helices. Enzymatic assays in human cells revealed that STEAP1 promotes iron(III) reduction when fused to the intracellular NADPH-binding domain of its family member STEAP4, suggesting that STEAP1 functions as a ferric reductase in STEAP heterotrimers. Our work provides a foundation for deciphering the molecular mechanisms of STEAP1 and may be useful in the design of new therapeutic strategies to target STEAP1 in cancer. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10735.map.gz emd_10735.map.gz | 4.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10735-v30.xml emd-10735-v30.xml emd-10735.xml emd-10735.xml | 32.2 KB 32.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_10735_fsc.xml emd_10735_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_10735.png emd_10735.png | 107.2 KB | ||
| Masks |  emd_10735_msk_1.map emd_10735_msk_1.map | 103 MB |  Mask map Mask map | |
| Others |  emd_10735_additional_1.map.gz emd_10735_additional_1.map.gz emd_10735_additional_2.map.gz emd_10735_additional_2.map.gz emd_10735_half_map_1.map.gz emd_10735_half_map_1.map.gz emd_10735_half_map_2.map.gz emd_10735_half_map_2.map.gz | 79.9 MB 5.7 MB 80.6 MB 80.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10735 http://ftp.pdbj.org/pub/emdb/structures/EMD-10735 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10735 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10735 | HTTPS FTP |
-Validation report
| Summary document |  emd_10735_validation.pdf.gz emd_10735_validation.pdf.gz | 382.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_10735_full_validation.pdf.gz emd_10735_full_validation.pdf.gz | 381.4 KB | Display | |
| Data in XML |  emd_10735_validation.xml.gz emd_10735_validation.xml.gz | 16.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10735 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10735 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10735 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10735 | HTTPS FTP |
-Related structure data
| Related structure data |  6y9bMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_10735.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10735.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened cryo-EM map of homotrimeric STEAP1 bound to three Fab-fragments of antibody mAb120.545. The constant region of the Fabs is masked out. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0285 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_10735_msk_1.map emd_10735_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Unsharpened, unmasked cryo-EM map of homotrimeric STEAP1 bound...
| File | emd_10735_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened, unmasked cryo-EM map of homotrimeric STEAP1 bound to three Fab-fragments of antibody mAb120.545. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Sharpened cryo-EM map of homotrimeric STEAP1 bound to...
| File | emd_10735_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened cryo-EM map of homotrimeric STEAP1 bound to three Fab-fragments of antibody mAb120.545. The constant region of the Fabs is masked out. The map is autosharpened (B = -69 Angstrom) in Relion. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 1 of homotrimeric STEAP1 bound to...
| File | emd_10735_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 of homotrimeric STEAP1 bound to three Fab-fragments of antibody mAb120.545. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 2 of homotrimeric STEAP1 bound to...
| File | emd_10735_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 of homotrimeric STEAP1 bound to three Fab-fragments of antibody mAb120.545. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Trimeric human six-transmembrane epithelial antigen of the prosta...
+Supramolecule #1: Trimeric human six-transmembrane epithelial antigen of the prosta...
+Supramolecule #2: Fab-fragment 1 of antibody mAb120.545
+Supramolecule #3: Homo-trimeric STEAP1, chain a, b and c.
+Supramolecule #4: Fab Fragment 2 of antibody mAb120.545
+Supramolecule #5: Fab-fragment 3 of antibody mAb120.545
+Macromolecule #1: Metalloreductase STEAP1
+Macromolecule #2: Fab120.545 light chain
+Macromolecule #3: Fab120.545 heavy chain
+Macromolecule #4: PROTOPORPHYRIN IX CONTAINING FE
+Macromolecule #5: 1,2-DIMYRISTOYL-SN-GLYCERO-3-PHOSPHATE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5.0 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.8 Component:
Details: Protein sample was subjected to size-exclusion chromatography in 20 mM Tris pH 7.8, 200 mM NaCl, 0.08% (w/v) digitonin. 1 mM FAD (final concentration) was added to the sample before vitrification. | |||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE | |||||||||||||||
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK IV / Details: blot 4 seconds, blotforce 0. | |||||||||||||||
| Details | STEAP1-Fab120.545 complex purified in digitonin. The sample was monodisperse. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Details | Data was collected on a 200 kV Talos Arctica, located at Utrecht University, the Netherlands. |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Frames/image: 1-26 / Number grids imaged: 1 / Number real images: 5325 / Average exposure time: 6.5 sec. / Average electron dose: 49.5 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated defocus max: 3.5 µm / Calibrated defocus min: 0.4 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 130000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z
Z Y
Y X
X