[English] 日本語
 Yorodumi
Yorodumi- EMDB-0651: Electron cryo-microscopy of the eukaryotic translation initiation... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0651 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Electron cryo-microscopy of the eukaryotic translation initiation factor 2B bound to eukaryotic translation initiation factor 2 from Homo sapiens | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | eukaryotic translation initiation factor 2B / eukaryotic translation initiation factor 2 / TRANSLATION | |||||||||
| Function / homology |  Function and homology information Function and homology informationtranslation initiation ternary complex / regulation of translation in response to endoplasmic reticulum stress / glial limiting end-foot / HRI-mediated signaling / eukaryotic translation initiation factor 2B complex / response to manganese-induced endoplasmic reticulum stress / Cellular response to mitochondrial stress / positive regulation of type B pancreatic cell apoptotic process / Response of EIF2AK1 (HRI) to heme deficiency / Recycling of eIF2:GDP ...translation initiation ternary complex / regulation of translation in response to endoplasmic reticulum stress / glial limiting end-foot / HRI-mediated signaling / eukaryotic translation initiation factor 2B complex / response to manganese-induced endoplasmic reticulum stress / Cellular response to mitochondrial stress / positive regulation of type B pancreatic cell apoptotic process / Response of EIF2AK1 (HRI) to heme deficiency / Recycling of eIF2:GDP / methionyl-initiator methionine tRNA binding / negative regulation of translational initiation in response to stress / PERK-mediated unfolded protein response / PERK regulates gene expression / response to kainic acid / eukaryotic translation initiation factor 2 complex / regulation of translational initiation in response to stress / translation factor activity, RNA binding / astrocyte development / formation of translation preinitiation complex / eukaryotic 48S preinitiation complex / astrocyte differentiation / oligodendrocyte development / regulation of translational initiation / protein-synthesizing GTPase / cytoplasmic translational initiation / Formation of the ternary complex, and subsequently, the 43S complex / Ribosomal scanning and start codon recognition / Translation initiation complex formation / positive regulation of translational initiation / Response of EIF2AK4 (GCN2) to amino acid deficiency / GTP hydrolysis and joining of the 60S ribosomal subunit / response to glucose / L13a-mediated translational silencing of Ceruloplasmin expression / ovarian follicle development / mitophagy / myelination / translation initiation factor binding / translation initiation factor activity / guanyl-nucleotide exchange factor activity / response to endoplasmic reticulum stress / cellular response to amino acid starvation / stress granule assembly / central nervous system development / hippocampus development / translational initiation / PKR-mediated signaling / ABC-family protein mediated transport / response to peptide hormone / cytoplasmic stress granule / cellular response to UV / regulation of translation / T cell receptor signaling pathway / cellular response to heat / response to heat / ribosome binding / cellular response to oxidative stress / positive regulation of apoptotic process / cadherin binding / GTPase activity / synapse / GTP binding / mitochondrion / RNA binding / extracellular exosome / ATP binding / membrane / identical protein binding / nucleus / plasma membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.03 Å | |||||||||
 Authors Authors | Nguyen HC / Kenner LR | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2019 Journal: Science / Year: 2019Title: eIF2B-catalyzed nucleotide exchange and phosphoregulation by the integrated stress response. Authors: Lillian R Kenner / Aditya A Anand / Henry C Nguyen / Alexander G Myasnikov / Carolin J Klose / Lea A McGeever / Jordan C Tsai / Lakshmi E Miller-Vedam / Peter Walter / Adam Frost /   Abstract: The integrated stress response (ISR) tunes the rate of protein synthesis. Control is exerted by phosphorylation of the general translation initiation factor eIF2. eIF2 is a guanosine triphosphatase ...The integrated stress response (ISR) tunes the rate of protein synthesis. Control is exerted by phosphorylation of the general translation initiation factor eIF2. eIF2 is a guanosine triphosphatase that becomes activated by eIF2B, a two-fold symmetric and heterodecameric complex that functions as eIF2's dedicated nucleotide exchange factor. Phosphorylation converts eIF2 from a substrate into an inhibitor of eIF2B. We report cryo-electron microscopy structures of eIF2 bound to eIF2B in the dephosphorylated state. The structures reveal that the eIF2B decamer is a static platform upon which one or two flexible eIF2 trimers bind and align with eIF2B's bipartite catalytic centers to catalyze nucleotide exchange. Phosphorylation refolds eIF2α, allowing it to contact eIF2B at a different interface and, we surmise, thereby sequestering it into a nonproductive complex. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0651.map.gz emd_0651.map.gz | 398.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0651-v30.xml emd-0651-v30.xml emd-0651.xml emd-0651.xml | 23.9 KB 23.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_0651.png emd_0651.png | 423.1 KB | ||
| Filedesc metadata |  emd-0651.cif.gz emd-0651.cif.gz | 8.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0651 http://ftp.pdbj.org/pub/emdb/structures/EMD-0651 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0651 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0651 | HTTPS FTP |
-Related structure data
| Related structure data |  6o85MC  0649C  0664C  6o81C  6o9zC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0651.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0651.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.822 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : eIF2 bound to eIF2B
+Supramolecule #1: eIF2 bound to eIF2B
+Macromolecule #1: Translation initiation factor eIF-2B subunit epsilon
+Macromolecule #2: Translation initiation factor eIF-2B subunit beta
+Macromolecule #3: Translation initiation factor eIF-2B subunit delta
+Macromolecule #4: Translation initiation factor eIF-2B subunit alpha
+Macromolecule #5: Translation initiation factor eIF-2B subunit gamma
+Macromolecule #6: Eukaryotic translation initiation factor 2 subunit 1
+Macromolecule #7: Eukaryotic translation initiation factor 2 subunit 3
+Macromolecule #8: Eukaryotic translation initiation factor 2 subunit beta
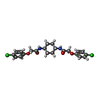
+Macromolecule #9: 2-(4-chloranylphenoxy)-~{N}-[4-[2-(4-chloranylphenoxy)ethanoylami...
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 80.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)