[English] 日本語
 Yorodumi
Yorodumi- EMDB-0145: CryoEM structure of the MDA5-dsRNA filament with 93 degree twist ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0145 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of the MDA5-dsRNA filament with 93 degree twist and without nucleotide | |||||||||
 Map data Map data | cryoEM reconstruction map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Protein-RNA complex / helical filament / ATPase / innate immune receptor / IMMUNE SYSTEM | |||||||||
| Function / homology |  Function and homology information Function and homology informationMDA-5 signaling pathway / positive regulation of response to cytokine stimulus / Ub-specific processing proteases / mRNA transcription / negative regulation of viral genome replication / type I interferon-mediated signaling pathway / cellular response to exogenous dsRNA / pattern recognition receptor activity / protein complex oligomerization / positive regulation of interferon-alpha production ...MDA-5 signaling pathway / positive regulation of response to cytokine stimulus / Ub-specific processing proteases / mRNA transcription / negative regulation of viral genome replication / type I interferon-mediated signaling pathway / cellular response to exogenous dsRNA / pattern recognition receptor activity / protein complex oligomerization / positive regulation of interferon-alpha production / protein sumoylation / ribonucleoprotein complex binding / positive regulation of interferon-beta production / antiviral innate immune response / cellular response to virus / positive regulation of interleukin-6 production / response to virus / positive regulation of tumor necrosis factor production / double-stranded RNA binding / defense response to virus / RNA helicase activity / single-stranded RNA binding / RNA helicase / protein domain specific binding / innate immune response / ATP hydrolysis activity / mitochondrion / DNA binding / RNA binding / zinc ion binding / ATP binding / identical protein binding / nucleus / cytoplasm Similarity search - Function | |||||||||
| Biological species |   Pseudomonas savastanoi pv. phaseolicola (bacteria) / Pseudomonas savastanoi pv. phaseolicola (bacteria) /  Pseudomonas phage phi6 (virus) Pseudomonas phage phi6 (virus) | |||||||||
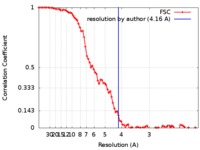
| Method | helical reconstruction / cryo EM / Resolution: 4.16 Å | |||||||||
 Authors Authors | Yu Q / Qu K | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2018 Journal: Mol Cell / Year: 2018Title: Cryo-EM Structures of MDA5-dsRNA Filaments at Different Stages of ATP Hydrolysis. Authors: Qin Yu / Kun Qu / Yorgo Modis /  Abstract: Double-stranded RNA (dsRNA) is a potent proinflammatory signature of viral infection. Long cytosolic dsRNA is recognized by MDA5. The cooperative assembly of MDA5 into helical filaments on dsRNA ...Double-stranded RNA (dsRNA) is a potent proinflammatory signature of viral infection. Long cytosolic dsRNA is recognized by MDA5. The cooperative assembly of MDA5 into helical filaments on dsRNA nucleates the assembly of a multiprotein type I interferon signaling platform. Here, we determined cryoelectron microscopy (cryo-EM) structures of MDA5-dsRNA filaments with different helical twists and bound nucleotide analogs at resolutions sufficient to build and refine atomic models. The structures identify the filament-forming interfaces, which encode the dsRNA binding cooperativity and length specificity of MDA5. The predominantly hydrophobic interface contacts confer flexibility, reflected in the variable helical twist within filaments. Mutation of filament-forming residues can result in loss or gain of signaling activity. Each MDA5 molecule spans 14 or 15 RNA base pairs, depending on the twist. Variations in twist also correlate with variations in the occupancy and type of nucleotide in the active site, providing insights on how ATP hydrolysis contributes to MDA5-dsRNA recognition. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0145.map.gz emd_0145.map.gz | 5.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0145-v30.xml emd-0145-v30.xml emd-0145.xml emd-0145.xml | 32.2 KB 32.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_0145_fsc.xml emd_0145_fsc.xml | 9.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_0145.png emd_0145.png | 166.9 KB | ||
| Masks |  emd_0145_msk_1.map emd_0145_msk_1.map | 47.6 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-0145.cif.gz emd-0145.cif.gz | 8.6 KB | ||
| Others |  emd_0145_half_map_1.map.gz emd_0145_half_map_1.map.gz emd_0145_half_map_2.map.gz emd_0145_half_map_2.map.gz | 11.5 MB 11.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0145 http://ftp.pdbj.org/pub/emdb/structures/EMD-0145 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0145 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0145 | HTTPS FTP |
-Related structure data
| Related structure data |  6h66MC  0012C  0023C  0024C  0143C  4338C  4340C  4341C  6g19C  6g1sC  6g1xC  6gjzC  6gkhC  6gkmC  6h61C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10210 (Title: mouse MDA5-dsRNA filaments / Data size: 499.5 EMPIAR-10210 (Title: mouse MDA5-dsRNA filaments / Data size: 499.5 Data #1: Unaligned multi-frame movies of mouse MDA5-dsRNA filaments without nucleotide [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0145.map.gz / Format: CCP4 / Size: 47.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0145.map.gz / Format: CCP4 / Size: 47.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryoEM reconstruction map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_0145_msk_1.map emd_0145_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
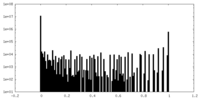
| Density Histograms |
-Half map: Half map 1
| File | emd_0145_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
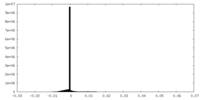
| Density Histograms |
-Half map: Half map 2
| File | emd_0145_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
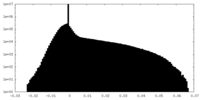
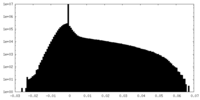
| Density Histograms |
- Sample components
Sample components
-Entire : MDA5-dsRNA helical filament without nucleotide
| Entire | Name: MDA5-dsRNA helical filament without nucleotide |
|---|---|
| Components |
|
-Supramolecule #1: MDA5-dsRNA helical filament without nucleotide
| Supramolecule | Name: MDA5-dsRNA helical filament without nucleotide / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 / Details: Filaments formed without nucleotide |
|---|---|
| Molecular weight | Theoretical: 2.23 kDa/nm |
-Supramolecule #2: MDA5 (IFIH1)
| Supramolecule | Name: MDA5 (IFIH1) / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 Details: RIG-I-like Superfamily 2 (SF2) RNA helicase consisting of Hel1, Hel2, Hel2i, and pincer domains, followed by a C-terminal domain |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: Double-stranded RNA from bacteriophage Phi6, generated by n vitro...
| Supramolecule | Name: Double-stranded RNA from bacteriophage Phi6, generated by n vitro translation type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:  Pseudomonas savastanoi pv. phaseolicola (bacteria) Pseudomonas savastanoi pv. phaseolicola (bacteria) |
-Macromolecule #1: Interferon-induced helicase C domain-containing protein 1
| Macromolecule | Name: Interferon-induced helicase C domain-containing protein 1 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: RNA helicase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 114.214477 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSIVCSAEDS FRNLILFFRP RLKMYIQVEP VLDHLIFLSA ETKEQILKKI NTCGNTSAAE LLLSTLEQGQ WPLGWTQMFV EALEHSGNP LAARYVKPTL TDLPSPSSET AHDECLHLLT LLQPTLVDKL LINDVLDTCF EKGLLTVEDR NRISAAGNSG N ESGVRELL ...String: MSIVCSAEDS FRNLILFFRP RLKMYIQVEP VLDHLIFLSA ETKEQILKKI NTCGNTSAAE LLLSTLEQGQ WPLGWTQMFV EALEHSGNP LAARYVKPTL TDLPSPSSET AHDECLHLLT LLQPTLVDKL LINDVLDTCF EKGLLTVEDR NRISAAGNSG N ESGVRELL RRIVQKENWF STFLDVLRQT GNDALFQELT GGGCPEDNTD LANSSHRDGP AANECLLPAV DESSLETEAW NV DDILPEA SCTDSSVTTE SDTSLAEGSV SCFDESLGHN SNMGRDSGTM GSDSDESVIQ TKRVSPEPEL QLRPYQMEVA QPA LDGKNI IICLPTGSGK TRVAVYITKD HLDKKKQASE SGKVIVLVNK VMLAEQLFRK EFNPYLKKWY RIIGLSGDTQ LKIS FPEVV KSYDVIISTA QILENSLLNL ESGDDDGVQL SDFSLIIIDE CHHTNKEAVY NNIMRRYLKQ KLRNNDLKKQ NKPAI PLPQ ILGLTASPGV GAAKKQSEAE KHILNICANL DAFTIKTVKE NLGQLKHQIK EPCKKFVIAD DTRENPFKEK LLEIMA SIQ TYCQKSPMSD FGTQHYEQWA IQMEKKAAKD GNRKDRVCAE HLRKYNEALQ INDTIRMIDA YSHLETFYTD EKEKKFA VL NDSKKSLKLD ETDEFLMNLF FDNKKMLKKL AENPKYENEK LIKLRNTILE QFTRSEESSR GIIFTKTRQS TYALSQWI M ENAKFAEVGV KAHHLIGAGH SSEVKPMTQT EQKEVISKFR TGEINLLIAT TVAEEGLDIK ECNIVIRYGL VTNEIAMVQ ARGRARADES TYVLVTSSGS GVTEREIVND FREKMMYKAI NRVQNMKPEE YAHKILELQV QSILEKKMKV KRSIAKQYND NPSLITLLC KNCSMLVCSG ENIHVIEKMH HVNMTPEFKG LYIVRENKAL QKKFADYQTN GEIICKCGQA WGTMMVHKGL D LPCLKIRN FVVNFKNNSP KKQYKKWVEL PIRFPDLDYS EYCLYSDED UniProtKB: Interferon-induced helicase C domain-containing protein 1 |
-Macromolecule #2: RNA (5'-R(P*UP*CP*CP*AP*UP*GP*CP*GP*CP*AP*UP*GP*AP*CP*G)-3')
| Macromolecule | Name: RNA (5'-R(P*UP*CP*CP*AP*UP*GP*CP*GP*CP*AP*UP*GP*AP*CP*G)-3') type: rna / ID: 2 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  Pseudomonas phage phi6 (virus) Pseudomonas phage phi6 (virus) |
| Molecular weight | Theoretical: 4.767889 KDa |
| Sequence | String: UCCAUGCGCA UGACG |
-Macromolecule #3: RNA (5'-R(P*CP*GP*UP*CP*AP*UP*GP*CP*GP*CP*AP*UP*GP*GP*A)-3')
| Macromolecule | Name: RNA (5'-R(P*CP*GP*UP*CP*AP*UP*GP*CP*GP*CP*AP*UP*GP*GP*A)-3') type: rna / ID: 3 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  Pseudomonas phage phi6 (virus) Pseudomonas phage phi6 (virus) |
| Molecular weight | Theoretical: 4.807914 KDa |
| Sequence | String: CGUCAUGCGC AUGGA |
-Macromolecule #4: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 4 / Number of copies: 1 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.7 Component:
| ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV | ||||||||||||
| Details | Samples were diluted twofold from 1 mg/ml to 0.5 mg/ml immediately prior to plunge freezing |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Average exposure time: 1.0 sec. / Average electron dose: 27.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 75000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: -2.7 µm / Nominal defocus min: -1.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Overall B value: 175 / Target criteria: Cross-correlation coefficient |
| Output model |  PDB-6h66: |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)