+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-23014 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | GluK2/K5 with 6-Cyano-7-nitroquinoxaline-2,3-dione (CNQX) | |||||||||
 Map data Map data | GluK2/K5 with CNQX (full-length map) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Kainate receptor / ionotropic glutamate receptor / membrane protein / ligand-gated ion channel / SIGNALING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of synaptic vesicle fusion to presynaptic active zone membrane / protein retention in ER lumen / mossy fiber rosette / detection of cold stimulus involved in thermoception / Activation of Na-permeable kainate receptors / Activation of Ca-permeable Kainate Receptor / kainate selective glutamate receptor complex / regulation of short-term neuronal synaptic plasticity / ubiquitin conjugating enzyme binding / glutamate receptor activity ...regulation of synaptic vesicle fusion to presynaptic active zone membrane / protein retention in ER lumen / mossy fiber rosette / detection of cold stimulus involved in thermoception / Activation of Na-permeable kainate receptors / Activation of Ca-permeable Kainate Receptor / kainate selective glutamate receptor complex / regulation of short-term neuronal synaptic plasticity / ubiquitin conjugating enzyme binding / glutamate receptor activity / negative regulation of synaptic transmission, glutamatergic / regulation of JNK cascade / receptor clustering / inhibitory postsynaptic potential / kainate selective glutamate receptor activity / extracellularly glutamate-gated ion channel activity / modulation of excitatory postsynaptic potential / ionotropic glutamate receptor complex / positive regulation of synaptic transmission / behavioral fear response / neuronal action potential / glutamate-gated receptor activity / glutamate-gated calcium ion channel activity / establishment of localization in cell / dendrite cytoplasm / ligand-gated monoatomic ion channel activity involved in regulation of presynaptic membrane potential / presynaptic modulation of chemical synaptic transmission / bioluminescence / hippocampal mossy fiber to CA3 synapse / SNARE binding / generation of precursor metabolites and energy / PDZ domain binding / transmitter-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / cellular response to glucose stimulus / synaptic transmission, glutamatergic / regulation of membrane potential / excitatory postsynaptic potential / intracellular protein transport / SH3 domain binding / regulation of long-term neuronal synaptic plasticity / postsynaptic density membrane / modulation of chemical synaptic transmission / intracellular calcium ion homeostasis / terminal bouton / positive regulation of neuron apoptotic process / neuron apoptotic process / scaffold protein binding / presynaptic membrane / chemical synaptic transmission / negative regulation of neuron apoptotic process / perikaryon / postsynaptic membrane / postsynaptic density / axon / neuronal cell body / ubiquitin protein ligase binding / synapse / dendrite / glutamatergic synapse / endoplasmic reticulum / nucleoplasm / membrane / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |   | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 5.3 Å | |||||||||
 Authors Authors | Khanra N / Brown PMGE | |||||||||
 Citation Citation |  Journal: Elife / Year: 2021 Journal: Elife / Year: 2021Title: Architecture and structural dynamics of the heteromeric GluK2/K5 kainate receptor. Authors: Nandish Khanra / Patricia Mge Brown / Amanda M Perozzo / Derek Bowie / Joel R Meyerson /   Abstract: Kainate receptors (KARs) are L-glutamate-gated ion channels that regulate synaptic transmission and modulate neuronal circuits. KARs have strict assembly rules and primarily function as heteromeric ...Kainate receptors (KARs) are L-glutamate-gated ion channels that regulate synaptic transmission and modulate neuronal circuits. KARs have strict assembly rules and primarily function as heteromeric receptors in the brain. A longstanding question is how KAR heteromer subunits organize and coordinate together to fulfill their signature physiological roles. Here we report structures of the GluK2/GluK5 heteromer in apo, antagonist-bound, and desensitized states. The receptor assembles with two copies of each subunit, ligand binding domains arranged as two heterodimers and GluK5 subunits proximal to the channel. Strikingly, during desensitization, GluK2, but not GluK5, subunits undergo major structural rearrangements to facilitate channel closure. We show how the large conformational differences between antagonist-bound and desensitized states are mediated by the linkers connecting the pore helices to the ligand binding domains. This work presents the first KAR heteromer structure, reveals how its subunits are organized, and resolves how the heteromer can accommodate functionally distinct closed channel structures. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23014.map.gz emd_23014.map.gz | 117.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23014-v30.xml emd-23014-v30.xml emd-23014.xml emd-23014.xml | 17.4 KB 17.4 KB | Display Display |  EMDB header EMDB header |
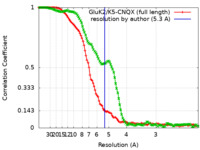
| FSC (resolution estimation) |  emd_23014_fsc.xml emd_23014_fsc.xml emd_23014_fsc_2.xml emd_23014_fsc_2.xml | 11.5 KB 11.2 KB | Display Display |  FSC data file FSC data file |
| Images |  emd_23014.png emd_23014.png | 57.4 KB | ||
| Filedesc metadata |  emd-23014.cif.gz emd-23014.cif.gz | 6.6 KB | ||
| Others |  emd_23014_additional_1.map.gz emd_23014_additional_1.map.gz emd_23014_additional_2.map.gz emd_23014_additional_2.map.gz | 98.6 MB 96.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23014 http://ftp.pdbj.org/pub/emdb/structures/EMD-23014 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23014 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23014 | HTTPS FTP |
-Related structure data
| Related structure data |  7ks0MC  7ks3C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10659 (Title: GluK2/K5 with 6-Cyano-7-nitroquinoxaline-2,3-dione (CNQX) EMPIAR-10659 (Title: GluK2/K5 with 6-Cyano-7-nitroquinoxaline-2,3-dione (CNQX)Data size: 5.9 TB Data #1: GluK2/K5 with 6-Cyano-7-nitroquinoxaline-2,3-dione (CNQX) [micrographs - multiframe] Data #2: GluK2/K5 with 6-Cyano-7-nitroquinoxaline-2,3-dione (CNQX) [micrographs - multiframe] Data #3: GluK2/K5 with 6-Cyano-7-nitroquinoxaline-2,3-dione (CNQX) [micrographs - multiframe] Data #4: GluK2/K5 with 6-Cyano-7-nitroquinoxaline-2,3-dione (CNQX) [picked particles - single frame - processed] Data #5: GluK2/K5 with 6-Cyano-7-nitroquinoxaline-2,3-dione (CNQX) [picked particles - single frame - processed]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_23014.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23014.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | GluK2/K5 with CNQX (full-length map) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.096 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: GluK2/K5 with CNQX (ATD map)
| File | emd_23014_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | GluK2/K5 with CNQX (ATD map) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: GluK2/K5 with CNQX (LBD-TMD map)
| File | emd_23014_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | GluK2/K5 with CNQX (LBD-TMD map) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : GluK2/K5 with 6-Cyano-7-nitroquinoxaline-2,3-dione (CNQX)
| Entire | Name: GluK2/K5 with 6-Cyano-7-nitroquinoxaline-2,3-dione (CNQX) |
|---|---|
| Components |
|
-Supramolecule #1: GluK2/K5 with 6-Cyano-7-nitroquinoxaline-2,3-dione (CNQX)
| Supramolecule | Name: GluK2/K5 with 6-Cyano-7-nitroquinoxaline-2,3-dione (CNQX) type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Glutamate receptor ionotropic, kainate 5,Green fluorescent protei...
| Macromolecule | Name: Glutamate receptor ionotropic, kainate 5,Green fluorescent protein chimera type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 123.676812 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MPAELLLLLI VAFANPSCQV LSSLRMAAIL DDQTVCGRGE RLALALAREQ INGIIEVPAK ARVEVDIFEL QRDSQYETTD TMCQILPKG VVSVLGPSSS PASASTVSHI CGEKEIPHIK VGPEETPRLQ YLRFASVSLY PSNEDVSLAV SRILKSFNYP S ASLICAKA ...String: MPAELLLLLI VAFANPSCQV LSSLRMAAIL DDQTVCGRGE RLALALAREQ INGIIEVPAK ARVEVDIFEL QRDSQYETTD TMCQILPKG VVSVLGPSSS PASASTVSHI CGEKEIPHIK VGPEETPRLQ YLRFASVSLY PSNEDVSLAV SRILKSFNYP S ASLICAKA ECLLRLEELV RGFLISKETL SVRMLDDSRD PTPLLKEIRD DKVSTIIIDA NASISHLVLR KASELGMTSA FY KYILTTM DFPILHLDGI VEDSSNILGF SMFNTSHPFY PEFVRSLNMS WRENCEASTY PGPALSAALM FDAVHVVVSA VRE LNRSQE IGVKPLACTS ANIWPHGTSL MNYLRMVEYD GLTGRVEFNS KGQRTNYTLR ILEKSRQGHR EIGVWYSNRT LAMN ATTLD INLSQTLANK TLVVTTILEN PYVMRRPNFQ ALSGNERFEG FCVDMLRELA ELLRFRYRLR LVEDGLYGAP EPNGS WTGM VGELINRKAD LAVAAFTITA EREKVIDFSK PFMTLGISIL YRVHMGRKPG YFSFLDPFSP AVWLFMLLAY LAVSVV LFL AARLSPYEWY NPHPSLRARP HILENQYTLG NSLWFPVGGF MQQGSEIMPR ALSTRIVSGV WWAFTLIIIS SYTANLA AF LTVQRMEVPV ESADDLADQT NIEYGTIHAG STMTFFQNSR YQTYQRMWNY MQSKQPSVFV KSTEEGIARV LNSRYAFL L ESTMNEYHRR LNCNLTQIGG LLDTKGYGIG MPLGSPFRDE ITLAILQLQE NNRLEILKRK WWEGGRCPKE EDHRAKGLG MENIGGIFVV LIAGLIIAVF VAVMEFIYKS RAEAKRMKGL VPRGSAAAAM VSKGEELFTG VVPILVELDG DVNGHKFSVS GEGEGDATY GKLTLKFICT TGKLPVPWPT LVTTLTYGVQ CFSRYPDHMK QHDFFKSAMP EGYVQERTIF FKDDGNYKTR A EVKFEGDT LVNRIELKGI DFKEDGNILG HKLEYNYNSH NVYIMADKQK NGIKVNFKIR HNIEDGSVQL ADHYQQNTPI GD GPVLLPD NHYLSTQSKL SKDPNEKRDH MVLLEFVTAA GITLGMDELY KSGLRTETSQ VAPA UniProtKB: Glutamate receptor ionotropic, kainate 5, Green fluorescent protein |
-Macromolecule #2: Glutamate receptor ionotropic, kainate 2
| Macromolecule | Name: Glutamate receptor ionotropic, kainate 2 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 105.981617 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MKIISPVLSN LVFSRSIKVL LCLLWIGYSQ GTTHVLRFGG IFEYVESGPM GAEELAFRFA VNTINRNRTL LPNTTLTYDT QKINLYDSF EASKKACDQL SLGVAAIFGP SHSSSANAVQ SICNALGVPH IQTRWKHQVS DNKDSFYVSL YPDFSSLSRA I LDLVQFFK ...String: MKIISPVLSN LVFSRSIKVL LCLLWIGYSQ GTTHVLRFGG IFEYVESGPM GAEELAFRFA VNTINRNRTL LPNTTLTYDT QKINLYDSF EASKKACDQL SLGVAAIFGP SHSSSANAVQ SICNALGVPH IQTRWKHQVS DNKDSFYVSL YPDFSSLSRA I LDLVQFFK WKTVTVVYDD STGLIRLQEL IKAPSRYNLR LKIRQLPADT KDAKPLLKEM KRGKEFHVIF DCSHEMAAGI LK QALAMGM MTEYYHYIFT TLDLFALDVE PYRYSGVNMT GFRILNTENT QVSSIIEKWS MERLQAPPKP DSGLLDGFMT TDA ALMYDA VHVVSVAVQQ FPQMTVSSLQ CNRHKPWRFG TRFMSLIKEA HWEGLTGRIT FNKTNGLRTD FDLDVISLKE EGLE KIGTW DPASGLNMTE SQKGKPANIT DSLSNRSLIV TTILEEPYVL FKKSDKPLYG NDRFEGYCID LLRELSTILG FTYEI RLVE DGKYGAQDDV NGQWNGMVRE LIDHKADLAV APLAITYVRE KVIDFSKPFM TLGISILYRK PNGTNPGVFS FLNPLS PDI WMYVLLAYLG VSVVLFVIAR FSPYEWYNPH PSNPDSDVVE NNFTLLNSFW FGVGALMQQG SELMPKALST RIVGGIW WF FTLIIISSYT ANLAAFLTVE RMESPIDSAD DLAKQTKIEY GAVEDGATMT FFKKSKISTY DKMWAFMSSR RQSVLVKS N EEGIQRVLTS DYAFLMESTT IEFVTQRNCN LTQIGGLIDS KGYGVGTPMG SPYRDKITIA ILQLQEEGKL HMMKEKWWR GNGCPEEESK EASALGVQNI GGIFIVLAAG LVLSVFVAVG EFLYKSKKNA QLEKRSFCSA MVEELRMSLK CQRRLKHKPQ APVIVKTEE VINMHTFNDR RLPGKETMAS GLRSAWSHPQ FEKGGGSGGG SGGGSWSHPQ FEK UniProtKB: Glutamate receptor ionotropic, kainate 2 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3.7 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Details: Functionalized with PEG-thiol | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV | |||||||||
| Details | 1 mM 6-Cyano-7-nitroquinoxaline-2,3-dione |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 51.5 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)