[English] 日本語
 Yorodumi
Yorodumi- EMDB-22975: Lethocerus Myosin II complete coiled-coil domain resolved in its ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22975 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Lethocerus Myosin II complete coiled-coil domain resolved in its native environment | |||||||||
 Map data Map data | One myosin molecule segmented out of the helically extended reconstruction | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | striated muscle / asynchronous flight muscle / MOTOR PROTEIN | |||||||||
| Biological species |  Lethocerus indicus (insect) Lethocerus indicus (insect) | |||||||||
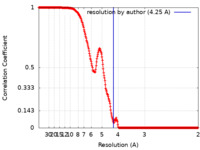
| Method | single particle reconstruction / cryo EM / Resolution: 4.25 Å | |||||||||
 Authors Authors | Rahmani H / Hu Z / Daneshparvar N / Taylor D / Irving T / Edwards RJ / Taylor KA | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2021 Journal: Proc Natl Acad Sci U S A / Year: 2021Title: The myosin II coiled-coil domain atomic structure in its native environment. Authors: Hamidreza Rahmani / Wen Ma / Zhongjun Hu / Nadia Daneshparvar / Dianne W Taylor / J Andrew McCammon / Thomas C Irving / Robert J Edwards / Kenneth A Taylor /  Abstract: The atomic structure of the complete myosin tail within thick filaments isolated from flight muscle is described and compared to crystal structures of recombinant, human cardiac myosin tail segments. ...The atomic structure of the complete myosin tail within thick filaments isolated from flight muscle is described and compared to crystal structures of recombinant, human cardiac myosin tail segments. Overall, the agreement is good with three exceptions: the proximal S2, in which the filament has heads attached but the crystal structure doesn't, and skip regions 2 and 4. At the head-tail junction, the tail α-helices are asymmetrically structured encompassing well-defined unfolding of 12 residues for one myosin tail, ∼4 residues of the other, and different degrees of α-helix unwinding for both tail α-helices, thereby providing an atomic resolution description of coiled-coil "uncoiling" at the head-tail junction. Asymmetry is observed in the nonhelical C termini; one C-terminal segment is intercalated between ribbons of myosin tails, the other apparently terminating at Skip 4 of another myosin tail. Between skip residues, crystal and filament structures agree well. Skips 1 and 3 also agree well and show the expected α-helix unwinding and coiled-coil untwisting in response to skip residue insertion. Skips 2 and 4 are different. Skip 2 is accommodated in an unusual manner through an increase in α-helix radius and corresponding reduction in rise/residue. Skip 4 remains helical in one chain, with the other chain unfolded, apparently influenced by the acidic myosin C terminus. The atomic model may shed some light on thick filament mechanosensing and is a step in understanding the complex roles that thick filaments of all species undergo during muscle contraction. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22975.map.gz emd_22975.map.gz | 1.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22975-v30.xml emd-22975-v30.xml emd-22975.xml emd-22975.xml | 13.4 KB 13.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_22975_fsc.xml emd_22975_fsc.xml | 26.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_22975.png emd_22975.png | 249 KB | ||
| Filedesc metadata |  emd-22975.cif.gz emd-22975.cif.gz | 6.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22975 http://ftp.pdbj.org/pub/emdb/structures/EMD-22975 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22975 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22975 | HTTPS FTP |
-Validation report
| Summary document |  emd_22975_validation.pdf.gz emd_22975_validation.pdf.gz | 309.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_22975_full_validation.pdf.gz emd_22975_full_validation.pdf.gz | 309.3 KB | Display | |
| Data in XML |  emd_22975_validation.xml.gz emd_22975_validation.xml.gz | 14.9 KB | Display | |
| Data in CIF |  emd_22975_validation.cif.gz emd_22975_validation.cif.gz | 22.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22975 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22975 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22975 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22975 | HTTPS FTP |
-Related structure data
| Related structure data |  7kogMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| EM raw data |  EMPIAR-10675 (Title: The Myosin II Coiled Coil Domain Atomic Structure in its Native Environment EMPIAR-10675 (Title: The Myosin II Coiled Coil Domain Atomic Structure in its Native EnvironmentData size: 18.8 TB / Data #1: Frame Stacks 1/3 [micrographs - multiframe] / Data #2: Frame Stacks 2/3 [micrographs - multiframe] / Data #3: Frame Stacks 3/3 [micrographs - multiframe] Data #4: Micrographs before frames alignment [micrographs - single frame] Data #5: Micrographs: frame-aligned and dose-weighted [micrographs - single frame] Data #6: Particle Stack [picked particles - single frame - unprocessed]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_22975.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22975.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | One myosin molecule segmented out of the helically extended reconstruction | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.98 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Lethocerus flight muscle myosin filament
| Entire | Name: Lethocerus flight muscle myosin filament |
|---|---|
| Components |
|
-Supramolecule #1: Lethocerus flight muscle myosin filament
| Supramolecule | Name: Lethocerus flight muscle myosin filament / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all Details: The sample is a bipolar helical structure, with helical repeat 145 Angstrom and helical turn 33.98 degree. The sample has C4 symmetry. The map contains 6 unique features: myosin molecule ...Details: The sample is a bipolar helical structure, with helical repeat 145 Angstrom and helical turn 33.98 degree. The sample has C4 symmetry. The map contains 6 unique features: myosin molecule with completely resolved rods, 4 resolved non-myosin densities among the myosin rods and an annular region inside of annulus occupied by myosin rods that most likely contains paramyosin. The 4 non-myosin densities may contain parts of the proteins myofilin and flightin. |
|---|---|
| Source (natural) | Organism:  Lethocerus indicus (insect) / Organ: myocyte / Tissue: dorsal longitudinal indirect flight muscle / Organelle: Sarcomere / Location in cell: myofibril Lethocerus indicus (insect) / Organ: myocyte / Tissue: dorsal longitudinal indirect flight muscle / Organelle: Sarcomere / Location in cell: myofibril |
-Macromolecule #1: Myosin heavy chain isoform Mhc_X1
| Macromolecule | Name: Myosin heavy chain isoform Mhc_X1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Lethocerus indicus (insect) Lethocerus indicus (insect) |
| Molecular weight | Theoretical: 225.227562 KDa |
| Sequence | String: MPGSKPTKTE EEDDPTPYLF VSLEQKRIDQ TKPYDAKKAC WVPDEHEGFV QGEIRGTKGD IVSVHLPNGE TKDFKKDQVG QVNPPKFEK CEDMSNLTYL NDASVLYNLK QRYYNKLIYT YSGLFCVAIN PYKRFPVYTM RCAKLYRGKR RNEVPPHIFA I SDGAYVNM ...String: MPGSKPTKTE EEDDPTPYLF VSLEQKRIDQ TKPYDAKKAC WVPDEHEGFV QGEIRGTKGD IVSVHLPNGE TKDFKKDQVG QVNPPKFEK CEDMSNLTYL NDASVLYNLK QRYYNKLIYT YSGLFCVAIN PYKRFPVYTM RCAKLYRGKR RNEVPPHIFA I SDGAYVNM LTNKENQSML ITGESGAGKT ENTKKVIAYF ATVGASTKKE EAASAASQKK GTLEDQVVQT NPVLEAFGNA KT VRNDNSS RFGKFIRIHF GPSGKLAGAD IETYLLEKAR VISQQSLERS YHIFYQVMSG AVPGVKELCL LSNDIYEYNY VSQ GKVTIP SVDDGEEFQA TDQAFDVLGF TQEEKDDIYK ITASVMHMGC MKFKQRGREE QAEADGTAEG ERVAKLLGLE AADL YKNLL KPRIKVGNEF VTQGRNLNQV IYSVGALSKG VFDRLFKFLV KKCNETLDTK QKRQHFIGVL DIAGFEIFDF NGFEQ LCIN FTNEKLQQFF NHHMFVLEQE EYTREGITWA FIDFGMDLVA CIDLIEKPMG ILSILEEESM FPKATDKTFE EKLMNN HLG KSPNFQKPKP PKPGCQAAHF AISHYAGVVS YNLTGWLEKN KDPLNDTVVD QFKKGSNKLL VEIFADHPGQ SGAPEAG GG GKGGRGKKGG GFATVSSSYK EQLNNLMTTL KSTQPHFVRC IIPNELKQPG LIDSHLVMHQ LTCNGVLEGI RICRKGFP N RMVYPDFKLR YMILAPATMA AEPDPKKAAD KCLKEVGLES ETYRIGHTKV FFRAGVLGQL EEMRDERLSK IIGWMQSHI RGYLARKQFK KYQDQRLSLQ VVQRNLRKYM ALRTWPWWKM WTKVKPLLNV ANVEEEMRKL EELVATTQAA LEKEEKARKE VEALNAKLI QEKTDLLRNL EGEKGSISSI QEKAAKLQAQ KSDLESQLMD TQERLQQEED NRNQMFQQKK KLEQEVGGLK K DIEDLELS LQKSDQDKAS KDHQIRNLND EIAHQDELIN KLNKEKKMQG EHTQKTAEEL QASEDKVNHL TKVKAKLEQT LD ELEDSLE REKKLRGDVE KAKRKVEGDL KLTQEAVADL ERNKKELEQT IQRKDKEIAS LTAKLEDEQS IVSKTQKQIK ELQ SRIEEL EEEVEAERQA RGKAEKQRAD LARELEELGE RLEEAGGATS AQIELNKKRE AEMSKLRRDL EESNIQHEST LANL RKKHN DAVSEMGEQI DQLNKLKTKA EHDRTHVQND LNNTRHALDQ MCREKAATEK IAKQLQHQVN EIQGKLDEAN RTLND FDSA KKKLSIENSD LLRQLEEAES QVSQLSKIKV SLTTQLEDTK RLADEEARER ATLLGKFRNL EHDLDNIREQ LEEEAE GKA DIQRQLSKAN AEAQLWRTKY ESEGVARAEE LEEAKRKLQA RLAEAEETIE SLNQKVIALE KTKQRLATEV EDLQLEV DR ATAIANAAEK KAKAIDKIIG EWKLKVDDLA AELDASQKEC RNYSTELFRL KGAYEEAQEQ LEAVRRENKN LADEVKDL L DQIGEGGRNI HEIEKQKKRL EVEKDELQAA LEEAEAALEQ EENKVLRSQL ELSQVRQEID RRIQEKEEEF ENTRKNHQR ALDSMQASLE AEAKGKAEAL RMKKKLEADI NELEIALDHA NKANSEAQKT IKKYQQQLKD VQTALEEEQR ARDDAREQLG ISERRANAL QNELEESRTL LEQADRGRRQ AEQELGDAHE QINELAAQAT SASAAKRKLE GELQTLHADL DELLNEAKNS E EKAKKAMV DAARLADELR AEQDHAQTQE KLRKALETQI KELQIRLDEA ETNALKGGKK AIAKLEQRVR ELENELDGEQ RR HADAQKN LRKSERRIKE LSFQADEDRK NHERMQDLVD KLQQKIKTYK RQIEEAEEIA ALNLAKFRKA QQELEEAEER ADL AEQAIA KFRTKGGRAG SAARAMSPVA HRPPVKHPLD GSTFPPRFDL HEDMM |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 6.8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: DIRECT ELECTRON DE-64 (8k x 8k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 8192 pixel / Digitization - Dimensions - Height: 8192 pixel / Digitization - Frames/image: 1-34 / Number grids imaged: 1 / Number real images: 3507 / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller