+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-22685 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Cryo-EM structure of a chromatosome containing human linker histone H1.10 | |||||||||
 マップデータ マップデータ | sharpened map of H1.10 chromatosome for structure model building | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | chromatosome / nucleosome / linker histones / Single-chain antibody / charge-charge interaction / chromatin / NUCLEAR PROTEIN / NUCLEAR PROTEIN-DNA complex | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報negative regulation of DNA recombination / chromosome condensation / nucleosomal DNA binding / negative regulation of tumor necrosis factor-mediated signaling pathway / negative regulation of megakaryocyte differentiation / protein localization to CENP-A containing chromatin / Chromatin modifying enzymes / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / Packaging Of Telomere Ends ...negative regulation of DNA recombination / chromosome condensation / nucleosomal DNA binding / negative regulation of tumor necrosis factor-mediated signaling pathway / negative regulation of megakaryocyte differentiation / protein localization to CENP-A containing chromatin / Chromatin modifying enzymes / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / Packaging Of Telomere Ends / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / Deposition of new CENPA-containing nucleosomes at the centromere / epigenetic regulation of gene expression / telomere organization / Interleukin-7 signaling / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / RNA Polymerase I Promoter Opening / Inhibition of DNA recombination at telomere / Assembly of the ORC complex at the origin of replication / Meiotic synapsis / SUMOylation of chromatin organization proteins / Regulation of endogenous retroelements by the Human Silencing Hub (HUSH) complex / DNA methylation / Condensation of Prophase Chromosomes / Chromatin modifications during the maternal to zygotic transition (MZT) / SIRT1 negatively regulates rRNA expression / HCMV Late Events / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / PRC2 methylates histones and DNA / innate immune response in mucosa / Regulation of endogenous retroelements by KRAB-ZFP proteins / Defective pyroptosis / HDACs deacetylate histones / Regulation of endogenous retroelements by Piwi-interacting RNAs (piRNAs) / Nonhomologous End-Joining (NHEJ) / RNA Polymerase I Promoter Escape / lipopolysaccharide binding / Transcriptional regulation by small RNAs / Formation of the beta-catenin:TCF transactivating complex / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / HDMs demethylate histones / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / G2/M DNA damage checkpoint / Negative Regulation of CDH1 Gene Transcription / NoRC negatively regulates rRNA expression / PKMTs methylate histone lysines / B-WICH complex positively regulates rRNA expression / DNA Damage/Telomere Stress Induced Senescence / Pre-NOTCH Transcription and Translation / Meiotic recombination / Activation of anterior HOX genes in hindbrain development during early embryogenesis / Metalloprotease DUBs / Transcriptional regulation of granulopoiesis / RMTs methylate histone arginines / HCMV Early Events / structural constituent of chromatin / UCH proteinases / nucleosome / antimicrobial humoral immune response mediated by antimicrobial peptide / heterochromatin formation / nucleosome assembly / E3 ubiquitin ligases ubiquitinate target proteins / antibacterial humoral response / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / HATs acetylate histones / chromosome / Factors involved in megakaryocyte development and platelet production / RUNX1 regulates transcription of genes involved in differentiation of HSCs / MLL4 and MLL3 complexes regulate expression of PPARG target genes in adipogenesis and hepatic steatosis / chromatin organization / Processing of DNA double-strand break ends / Senescence-Associated Secretory Phenotype (SASP) / double-stranded DNA binding / Oxidative Stress Induced Senescence / Estrogen-dependent gene expression / killing of cells of another organism / defense response to Gram-negative bacterium / chromosome, telomeric region / Ub-specific processing proteases / defense response to Gram-positive bacterium / cadherin binding / Amyloid fiber formation / protein heterodimerization activity / negative regulation of cell population proliferation / nucleolus / protein-containing complex / : / DNA binding / RNA binding / extracellular exosome / extracellular region / nucleoplasm / membrane / nucleus / cytosol 類似検索 - 分子機能 | |||||||||
| 生物種 |  human (ヒト) / human (ヒト) /  Homo sapiens (ヒト) / Homo sapiens (ヒト) /  | |||||||||
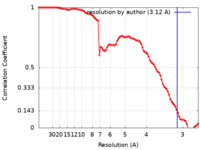
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.12 Å | |||||||||
 データ登録者 データ登録者 | Zhou B-R / Bai Y | |||||||||
| 資金援助 |  米国, 1件 米国, 1件
| |||||||||
 引用 引用 |  ジャーナル: Mol Cell / 年: 2021 ジャーナル: Mol Cell / 年: 2021タイトル: Distinct Structures and Dynamics of Chromatosomes with Different Human Linker Histone Isoforms. 著者: Bing-Rui Zhou / Hanqiao Feng / Seyit Kale / Tara Fox / Htet Khant / Natalia de Val / Rodolfo Ghirlando / Anna R Panchenko / Yawen Bai /    要旨: The repeating structural unit of metazoan chromatin is the chromatosome, a nucleosome bound to a linker histone, H1. There are 11 human H1 isoforms with diverse cellular functions, but how they ...The repeating structural unit of metazoan chromatin is the chromatosome, a nucleosome bound to a linker histone, H1. There are 11 human H1 isoforms with diverse cellular functions, but how they interact with the nucleosome remains elusive. Here, we determined the cryoelectron microscopy (cryo-EM) structures of chromatosomes containing 197 bp DNA and three different human H1 isoforms, respectively. The globular domains of all three H1 isoforms bound to the nucleosome dyad. However, the flanking/linker DNAs displayed substantial distinct dynamic conformations. Nuclear magnetic resonance (NMR) and H1 tail-swapping cryo-EM experiments revealed that the C-terminal tails of the H1 isoforms mainly controlled the flanking DNA orientations. We also observed partial ordering of the core histone H2A C-terminal and H3 N-terminal tails in the chromatosomes. Our results provide insights into the structures and dynamics of the chromatosomes and have implications for the structure and function of chromatin. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_22685.map.gz emd_22685.map.gz | 59.3 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-22685-v30.xml emd-22685-v30.xml emd-22685.xml emd-22685.xml | 22 KB 22 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_22685_fsc.xml emd_22685_fsc.xml | 8.9 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_22685.png emd_22685.png | 99.5 KB | ||
| Filedesc metadata |  emd-22685.cif.gz emd-22685.cif.gz | 7.2 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22685 http://ftp.pdbj.org/pub/emdb/structures/EMD-22685 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22685 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22685 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  7k60MC  7k5xC  7k5yC  7k61C  7k63C M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | |
| 電子顕微鏡画像生データ |  EMPIAR-10575 (タイトル: Single-particle cryo-EM micrographs of a chromatosome containing human linker histone H1.10 EMPIAR-10575 (タイトル: Single-particle cryo-EM micrographs of a chromatosome containing human linker histone H1.10Data size: 278.2 Data #1: Unaligned multi-frames micrograph of a chomatosome containing human linker histone H1.10 [micrographs - multiframe]) |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_22685.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_22685.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | sharpened map of H1.10 chromatosome for structure model building | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.34 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
- 試料の構成要素
試料の構成要素
-全体 : chromatosome containing human linker histone H1.10
| 全体 | 名称: chromatosome containing human linker histone H1.10 |
|---|---|
| 要素 |
|
-超分子 #1: chromatosome containing human linker histone H1.10
| 超分子 | 名称: chromatosome containing human linker histone H1.10 / タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: #1-#8 |
|---|---|
| 由来(天然) | 生物種:  human (ヒト) human (ヒト) |
-分子 #1: Histone H3.1
| 分子 | 名称: Histone H3.1 / タイプ: protein_or_peptide / ID: 1 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 15.437167 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MARTKQTARK STGGKAPRKQ LATKAARKSA PATGGVKKPH RYRPGTVALR EIRRYQKSTE LLIRKLPFQR LVREIAQDFK TDLRFQSSA VMALQEACEA YLVGLFEDTN LCAIHAKRVT IMPKDIQLAR RIRGERA UniProtKB: Histone H3.1 |
-分子 #2: Histone H4
| 分子 | 名称: Histone H4 / タイプ: protein_or_peptide / ID: 2 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 11.394426 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MSGRGKGGKG LGKGGAKRHR KVLRDNIQGI TKPAIRRLAR RGGVKRISGL IYEETRGVLK VFLENVIRDA VTYTEHAKRK TVTAMDVVY ALKRQGRTLY GFGG UniProtKB: Histone H4 |
-分子 #3: Histone H2A type 1-B/E
| 分子 | 名称: Histone H2A type 1-B/E / タイプ: protein_or_peptide / ID: 3 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 14.165551 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MSGRGKQGGK ARAKAKTRSS RAGLQFPVGR VHRLLRKGNY SERVGAGAPV YLAAVLEYLT AEILELAGNA ARDNKKTRII PRHLQLAIR NDEELNKLLG RVTIAQGGVL PNIQAVLLPK KTESHHKAKG K UniProtKB: Histone H2A type 1-B/E |
-分子 #4: Histone H2B type 1-J
| 分子 | 名称: Histone H2B type 1-J / タイプ: protein_or_peptide / ID: 4 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 13.935239 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MPEPAKSAPA PKKGSKKAVT KAQKKDGKKR KRSRKESYSI YVYKVLKQVH PDTGISSKAM GIMNSFVNDI FERIAGEASR LAHYNKRST ITSREIQTAV RLLLPGELAK HAVSEGTKAV TKYTSAK UniProtKB: Histone H2B type 1-J |
-分子 #7: scFv20
| 分子 | 名称: scFv20 / タイプ: protein_or_peptide / ID: 7 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  |
| 分子量 | 理論値: 29.34542 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MKSSHHHHHH ENLYFQSNAM DIKMTQSPSS MHASLGERVT ITCKASQDIR SYLSWYQQKP WKSPKTLIYY ATSLADGVPS RFSGSGSGQ DFSLTINNLE SDDTATYYCL QHGESPYTFG SGTKLEIKRA GGGGSGGGGS GGGGSGGGGS MEVQLQQSGP E LVEPGTSV ...文字列: MKSSHHHHHH ENLYFQSNAM DIKMTQSPSS MHASLGERVT ITCKASQDIR SYLSWYQQKP WKSPKTLIYY ATSLADGVPS RFSGSGSGQ DFSLTINNLE SDDTATYYCL QHGESPYTFG SGTKLEIKRA GGGGSGGGGS GGGGSGGGGS MEVQLQQSGP E LVEPGTSV KMPCKASGYT FTSYTIQWVK QTPRQGLEWI GYIYPYNAGT KYNEKFKGKA TLTSDKSSST VYMELSSLTS ED SAVYYCA RKSSRLRSTL DYWGQGTSVT VSS |
-分子 #8: Histone H1.10
| 分子 | 名称: Histone H1.10 / タイプ: protein_or_peptide / ID: 8 / コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 22.543301 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MSVELEEALP VTTAEGMAKK VTKAGGSAAL SPSKKRKNSK KKNQPGKYSQ LVVETIRRLG ERNGSSLAKI YTEAKKVPWF DQQNGRTYL KYSIKALVQN DTLLQVKGTG ANGSFKLNRK KLEGGGERRG APAAATAPAP TAHKAKKAAP GAAGSRRADK K PARGQKPE ...文字列: MSVELEEALP VTTAEGMAKK VTKAGGSAAL SPSKKRKNSK KKNQPGKYSQ LVVETIRRLG ERNGSSLAKI YTEAKKVPWF DQQNGRTYL KYSIKALVQN DTLLQVKGTG ANGSFKLNRK KLEGGGERRG APAAATAPAP TAHKAKKAAP GAAGSRRADK K PARGQKPE QRSHKKGAGA KKDKGGKAKK TAAAGGKKVK KAAKPSVPKV PKGRK UniProtKB: Histone H1.10 |
-分子 #5: DNA (197-MER)
| 分子 | 名称: DNA (197-MER) / タイプ: dna / ID: 5 / コピー数: 1 / 分類: DNA |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 60.510484 KDa |
| 配列 | 文字列: (DG)(DG)(DG)(DC)(DT)(DG)(DG)(DA)(DC)(DC) (DC)(DT)(DA)(DT)(DA)(DC)(DG)(DC)(DG)(DG) (DC)(DC)(DG)(DC)(DC)(DC)(DT)(DG)(DG) (DA)(DG)(DA)(DA)(DT)(DC)(DC)(DC)(DG)(DG) (DT) (DG)(DC)(DC)(DG)(DA) ...文字列: (DG)(DG)(DG)(DC)(DT)(DG)(DG)(DA)(DC)(DC) (DC)(DT)(DA)(DT)(DA)(DC)(DG)(DC)(DG)(DG) (DC)(DC)(DG)(DC)(DC)(DC)(DT)(DG)(DG) (DA)(DG)(DA)(DA)(DT)(DC)(DC)(DC)(DG)(DG) (DT) (DG)(DC)(DC)(DG)(DA)(DG)(DG)(DC) (DC)(DG)(DC)(DT)(DC)(DA)(DA)(DT)(DT)(DG) (DG)(DT) (DC)(DG)(DT)(DA)(DG)(DA)(DC) (DA)(DG)(DC)(DT)(DC)(DT)(DA)(DG)(DC)(DA) (DC)(DC)(DG) (DC)(DT)(DT)(DA)(DA)(DA) (DC)(DG)(DC)(DA)(DC)(DG)(DT)(DA)(DC)(DG) (DC)(DG)(DC)(DT) (DG)(DT)(DC)(DC)(DC) (DC)(DC)(DG)(DC)(DG)(DT)(DT)(DT)(DT)(DA) (DA)(DC)(DC)(DG)(DC) (DC)(DA)(DA)(DG) (DG)(DG)(DG)(DA)(DT)(DT)(DA)(DC)(DT)(DC) (DC)(DC)(DT)(DA)(DG)(DT) (DC)(DT)(DC) (DC)(DA)(DG)(DG)(DC)(DA)(DC)(DG)(DT)(DG) (DT)(DC)(DA)(DG)(DA)(DT)(DA) (DT)(DA) (DT)(DA)(DC)(DA)(DT)(DC)(DC)(DT)(DG)(DT) (DG)(DC)(DA)(DT)(DG)(DT)(DA)(DT) (DT) (DG)(DA)(DA)(DC)(DA)(DG)(DC)(DG)(DA)(DC) (DC)(DA)(DC)(DC)(DC)(DC) |
-分子 #6: DNA (197-MER)
| 分子 | 名称: DNA (197-MER) / タイプ: dna / ID: 6 / コピー数: 1 / 分類: DNA |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 61.141883 KDa |
| 配列 | 文字列: (DG)(DG)(DG)(DG)(DT)(DG)(DG)(DT)(DC)(DG) (DC)(DT)(DG)(DT)(DT)(DC)(DA)(DA)(DT)(DA) (DC)(DA)(DT)(DG)(DC)(DA)(DC)(DA)(DG) (DG)(DA)(DT)(DG)(DT)(DA)(DT)(DA)(DT)(DA) (DT) (DC)(DT)(DG)(DA)(DC) ...文字列: (DG)(DG)(DG)(DG)(DT)(DG)(DG)(DT)(DC)(DG) (DC)(DT)(DG)(DT)(DT)(DC)(DA)(DA)(DT)(DA) (DC)(DA)(DT)(DG)(DC)(DA)(DC)(DA)(DG) (DG)(DA)(DT)(DG)(DT)(DA)(DT)(DA)(DT)(DA) (DT) (DC)(DT)(DG)(DA)(DC)(DA)(DC)(DG) (DT)(DG)(DC)(DC)(DT)(DG)(DG)(DA)(DG)(DA) (DC)(DT) (DA)(DG)(DG)(DG)(DA)(DG)(DT) (DA)(DA)(DT)(DC)(DC)(DC)(DC)(DT)(DT)(DG) (DG)(DC)(DG) (DG)(DT)(DT)(DA)(DA)(DA) (DA)(DC)(DG)(DC)(DG)(DG)(DG)(DG)(DG)(DA) (DC)(DA)(DG)(DC) (DG)(DC)(DG)(DT)(DA) (DC)(DG)(DT)(DG)(DC)(DG)(DT)(DT)(DT)(DA) (DA)(DG)(DC)(DG)(DG) (DT)(DG)(DC)(DT) (DA)(DG)(DA)(DG)(DC)(DT)(DG)(DT)(DC)(DT) (DA)(DC)(DG)(DA)(DC)(DC) (DA)(DA)(DT) (DT)(DG)(DA)(DG)(DC)(DG)(DG)(DC)(DC)(DT) (DC)(DG)(DG)(DC)(DA)(DC)(DC) (DG)(DG) (DG)(DA)(DT)(DT)(DC)(DT)(DC)(DC)(DA)(DG) (DG)(DG)(DC)(DG)(DG)(DC)(DC)(DG) (DC) (DG)(DT)(DA)(DT)(DA)(DG)(DG)(DG)(DT)(DC) (DC)(DA)(DG)(DC)(DC)(DC) |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.4 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 平均電子線量: 40.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: OTHER / 撮影モード: OTHER |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー






















 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)