[English] 日本語
 Yorodumi
Yorodumi- EMDB-9645: Cryo-EM structure of a human post-catalytic spliceosome (P comple... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9645 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of a human post-catalytic spliceosome (P complex) at 3.0 angstrom | |||||||||
 Map data Map data | The 3.0 angstrom map of the human P complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Human Post-catalytic Spliceosome / SPLICING | |||||||||
| Function / homology |  Function and homology information Function and homology informationsecond spliceosomal transesterification activity / exon-exon junction subcomplex mago-y14 / negative regulation of selenocysteine incorporation / cellular response to selenite ion / regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay / exon-exon junction complex / selenocysteine insertion sequence binding / pre-mRNA 3'-splice site binding / regulation of translation at postsynapse, modulating synaptic transmission / regulation of retinoic acid receptor signaling pathway ...second spliceosomal transesterification activity / exon-exon junction subcomplex mago-y14 / negative regulation of selenocysteine incorporation / cellular response to selenite ion / regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay / exon-exon junction complex / selenocysteine insertion sequence binding / pre-mRNA 3'-splice site binding / regulation of translation at postsynapse, modulating synaptic transmission / regulation of retinoic acid receptor signaling pathway / post-mRNA release spliceosomal complex / renal system process / U2 snRNP binding / U7 snRNA binding / histone pre-mRNA DCP binding / generation of catalytic spliceosome for first transesterification step / U7 snRNP / 3'-5' RNA helicase activity / Z-decay: degradation of maternal mRNAs by zygotically expressed factors / cis assembly of pre-catalytic spliceosome / histone pre-mRNA 3'end processing complex / regulation of vitamin D receptor signaling pathway / regulation of mRNA processing / SLBP independent Processing of Histone Pre-mRNAs / SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs / spliceosome conformational change to release U4 (or U4atac) and U1 (or U11) / Deadenylation of mRNA / negative regulation of excitatory postsynaptic potential / intracellular mRNA localization / embryonic cranial skeleton morphogenesis / alternative mRNA splicing, via spliceosome / protein methylation / poly(A) binding / 7-methylguanosine cap hypermethylation / U12-type spliceosomal complex / embryonic brain development / nuclear retinoic acid receptor binding / U1 snRNP binding / methylosome / U2-type catalytic step 1 spliceosome / ATP-dependent activity, acting on RNA / regulation of mRNA splicing, via spliceosome / pre-mRNA binding / C2H2 zinc finger domain binding / pICln-Sm protein complex / positive regulation of mRNA splicing, via spliceosome / mRNA 3'-end processing / RNA splicing, via transesterification reactions / sno(s)RNA-containing ribonucleoprotein complex / M-decay: degradation of maternal mRNAs by maternally stored factors / small nuclear ribonucleoprotein complex / SMN-Sm protein complex / spliceosomal tri-snRNP complex / P granule / positive regulation of vitamin D receptor signaling pathway / snRNP binding / commitment complex / host-mediated activation of viral transcription / U2-type precatalytic spliceosome / oocyte development / mRNA cis splicing, via spliceosome / telomerase holoenzyme complex / Regulation of gene expression in late stage (branching morphogenesis) pancreatic bud precursor cells / Transport of Mature mRNA derived from an Intron-Containing Transcript / Notch binding / RUNX3 regulates NOTCH signaling / telomerase RNA binding / U2-type prespliceosome assembly / U2-type catalytic step 2 spliceosome / U2-type spliceosomal complex / nuclear vitamin D receptor binding / NOTCH4 Intracellular Domain Regulates Transcription / U1 snRNP / U2 snRNP / RNA Polymerase II Transcription Termination / U4 snRNP / NOTCH3 Intracellular Domain Regulates Transcription / nuclear-transcribed mRNA catabolic process, nonsense-mediated decay / U2-type prespliceosome / protein peptidyl-prolyl isomerization / positive regulation of neurogenesis / inner cell mass cell proliferation / K63-linked polyubiquitin modification-dependent protein binding / nuclear androgen receptor binding / cyclosporin A binding / ubiquitin-ubiquitin ligase activity / precatalytic spliceosome / Notch-HLH transcription pathway / Formation of paraxial mesoderm / lipid biosynthetic process / WD40-repeat domain binding / regulation of alternative mRNA splicing, via spliceosome / positive regulation of transforming growth factor beta receptor signaling pathway / mRNA 3'-splice site recognition / SMAD binding / mitotic G2 DNA damage checkpoint signaling / mRNA Splicing - Minor Pathway / protein kinase inhibitor activity / spliceosomal complex assembly / exploration behavior Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Zhang X / Zhan X | |||||||||
| Funding support |  China, 2 items China, 2 items
| |||||||||
 Citation Citation |  Journal: Cell Res / Year: 2019 Journal: Cell Res / Year: 2019Title: Structures of the human spliceosomes before and after release of the ligated exon. Authors: Xiaofeng Zhang / Xiechao Zhan / Chuangye Yan / Wenyu Zhang / Dongliang Liu / Jianlin Lei / Yigong Shi /  Abstract: Pre-mRNA splicing is executed by the spliceosome, which has eight major functional states each with distinct composition. Five of these eight human spliceosomal complexes, all preceding exon ...Pre-mRNA splicing is executed by the spliceosome, which has eight major functional states each with distinct composition. Five of these eight human spliceosomal complexes, all preceding exon ligation, have been structurally characterized. In this study, we report the cryo-electron microscopy structures of the human post-catalytic spliceosome (P complex) and intron lariat spliceosome (ILS) at average resolutions of 3.0 and 2.9 Å, respectively. In the P complex, the ligated exon remains anchored to loop I of U5 small nuclear RNA, and the 3'-splice site is recognized by the junction between the 5'-splice site and the branch point sequence. The ATPase/helicase Prp22, along with the ligated exon and eight other proteins, are dissociated in the P-to-ILS transition. Intriguingly, the ILS complex exists in two distinct conformations, one with the ATPase/helicase Prp43 and one without. Comparison of these three late-stage human spliceosomes reveals mechanistic insights into exon release and spliceosome disassembly. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9645.map.gz emd_9645.map.gz | 226.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9645-v30.xml emd-9645-v30.xml emd-9645.xml emd-9645.xml | 68.5 KB 68.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9645.png emd_9645.png | 89.7 KB | ||
| Filedesc metadata |  emd-9645.cif.gz emd-9645.cif.gz | 21.6 KB | ||
| Others |  emd_9645_additional.map.gz emd_9645_additional.map.gz | 193.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9645 http://ftp.pdbj.org/pub/emdb/structures/EMD-9645 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9645 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9645 | HTTPS FTP |
-Related structure data
| Related structure data |  6iczMC  9646C  9647C  6id0C  6id1C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9645.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9645.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The 3.0 angstrom map of the human P complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.338 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
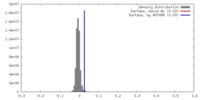
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: The 4.7 angstrom map of the human P complex
| File | emd_9645_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The 4.7 angstrom map of the human P complex | ||||||||||||
| Projections & Slices |
| ||||||||||||
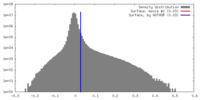
| Density Histograms |
- Sample components
Sample components
+Entire : Human Post_catalytic Spliceosome
+Supramolecule #1: Human Post_catalytic Spliceosome
+Macromolecule #1: Protein mago nashi homolog 2
+Macromolecule #2: RNA-binding protein 8A
+Macromolecule #3: Eukaryotic initiation factor 4A-III
+Macromolecule #4: Protein CASC3
+Macromolecule #5: Pre-mRNA-processing-splicing factor 8
+Macromolecule #7: 116 kDa U5 small nuclear ribonucleoprotein component
+Macromolecule #8: U5 small nuclear ribonucleoprotein 40 kDa protein
+Macromolecule #12: Crooked neck-like protein 1
+Macromolecule #13: Cell division cycle 5-like protein
+Macromolecule #14: Pre-mRNA-splicing factor SYF2
+Macromolecule #15: Protein BUD31 homolog
+Macromolecule #16: Pre-mRNA-splicing factor RBM22
+Macromolecule #17: Spliceosome-associated protein CWC15 homolog
+Macromolecule #18: SNW domain-containing protein 1
+Macromolecule #19: Peptidyl-prolyl cis-trans isomerase-like 1
+Macromolecule #20: Pleiotropic regulator 1
+Macromolecule #21: Serine/arginine repetitive matrix protein 2
+Macromolecule #22: Pre-mRNA-splicing factor CWC22 homolog
+Macromolecule #23: Pre-mRNA-processing factor 17
+Macromolecule #24: PRKR-interacting protein 1
+Macromolecule #25: Pre-mRNA-splicing factor SLU7
+Macromolecule #26: Pre-mRNA-splicing factor SYF1
+Macromolecule #27: Peptidyl-prolyl cis-trans isomerase E
+Macromolecule #28: RNA helicase aquarius
+Macromolecule #29: Small nuclear ribonucleoprotein Sm D3
+Macromolecule #30: Small nuclear ribonucleoprotein-associated protein
+Macromolecule #31: Small nuclear ribonucleoprotein Sm D1
+Macromolecule #32: Small nuclear ribonucleoprotein Sm D2
+Macromolecule #33: Small nuclear ribonucleoprotein F
+Macromolecule #34: Small nuclear ribonucleoprotein E
+Macromolecule #35: Small nuclear ribonucleoprotein G
+Macromolecule #36: U2 small nuclear ribonucleoprotein A'
+Macromolecule #37: U2 small nuclear ribonucleoprotein B''
+Macromolecule #38: ATP-dependent RNA helicase DHX8
+Macromolecule #39: Pre-mRNA-splicing factor SPF27
+Macromolecule #40: Pre-mRNA-processing factor 19
+Macromolecule #41: U5 small nuclear ribonucleoprotein 200 kDa helicase
+Macromolecule #6: U5snRNA
+Macromolecule #9: U6snRNA
+Macromolecule #10: pre-mRNA
+Macromolecule #11: U2snRNA
+Macromolecule #42: INOSITOL HEXAKISPHOSPHATE
+Macromolecule #43: GUANOSINE-5'-TRIPHOSPHATE
+Macromolecule #44: MAGNESIUM ION
+Macromolecule #45: ZINC ION
+Macromolecule #46: ADENOSINE-5'-TRIPHOSPHATE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.9 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.0 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 143320 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller










































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)