[English] 日本語
 Yorodumi
Yorodumi- EMDB-9375: Stabilized beta-arrestin 1-V2T subcomplex of a GPCR-G protein-bet... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9375 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Stabilized beta-arrestin 1-V2T subcomplex of a GPCR-G protein-beta-arrestin mega-complex | |||||||||
 Map data Map data | Stabilized beta-arrestin 1-V2T subcomplex of a GPCR-G protein-beta-arrestin mega-complex | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationMAP2K and MAPK activation / Activation of SMO / Golgi Associated Vesicle Biogenesis / Lysosome Vesicle Biogenesis / AP-2 adaptor complex binding / clathrin heavy chain binding / clathrin coat of coated pit / Ub-specific processing proteases / desensitization of G protein-coupled receptor signaling pathway / Cargo recognition for clathrin-mediated endocytosis ...MAP2K and MAPK activation / Activation of SMO / Golgi Associated Vesicle Biogenesis / Lysosome Vesicle Biogenesis / AP-2 adaptor complex binding / clathrin heavy chain binding / clathrin coat of coated pit / Ub-specific processing proteases / desensitization of G protein-coupled receptor signaling pathway / Cargo recognition for clathrin-mediated endocytosis / inositol hexakisphosphate binding / Clathrin-mediated endocytosis / clathrin-dependent endocytosis / G protein-coupled receptor internalization / acetylcholine receptor binding / Thrombin signalling through proteinase activated receptors (PARs) / G alpha (s) signalling events / clathrin binding / negative regulation of Notch signaling pathway / pseudopodium / small molecule binding / positive regulation of receptor internalization / phosphatidylinositol-3,4,5-trisphosphate binding / visual perception / G protein-coupled receptor binding / receptor internalization / protein transport / ubiquitin-dependent protein catabolic process / cytoplasmic vesicle / molecular adaptor activity / positive regulation of ERK1 and ERK2 cascade / positive regulation of protein phosphorylation / signal transduction / nucleus / plasma membrane / cytosol / cytoplasm Similarity search - Function | |||||||||
| Biological species |    Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.0 Å | |||||||||
 Authors Authors | Nguyen AH / Thomsen ARB / Cahill TJ / des Georges A / Lefkowitz RJ | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2019 Journal: Nat Struct Mol Biol / Year: 2019Title: Structure of an endosomal signaling GPCR-G protein-β-arrestin megacomplex. Authors: Anthony H Nguyen / Alex R B Thomsen / Thomas J Cahill / Rick Huang / Li-Yin Huang / Tara Marcink / Oliver B Clarke / Søren Heissel / Ali Masoudi / Danya Ben-Hail / Fadi Samaan / Venkata P ...Authors: Anthony H Nguyen / Alex R B Thomsen / Thomas J Cahill / Rick Huang / Li-Yin Huang / Tara Marcink / Oliver B Clarke / Søren Heissel / Ali Masoudi / Danya Ben-Hail / Fadi Samaan / Venkata P Dandey / Yong Zi Tan / Chuan Hong / Jacob P Mahoney / Sarah Triest / John Little / Xin Chen / Roger Sunahara / Jan Steyaert / Henrik Molina / Zhiheng Yu / Amedee des Georges / Robert J Lefkowitz /    Abstract: Classically, G-protein-coupled receptors (GPCRs) are thought to activate G protein from the plasma membrane and are subsequently desensitized by β-arrestin (β-arr). However, some GPCRs continue to ...Classically, G-protein-coupled receptors (GPCRs) are thought to activate G protein from the plasma membrane and are subsequently desensitized by β-arrestin (β-arr). However, some GPCRs continue to signal through G protein from internalized compartments, mediated by a GPCR-G protein-β-arr 'megaplex'. Nevertheless, the molecular architecture of the megaplex remains unknown. Here, we present its cryo-electron microscopy structure, which shows simultaneous engagement of human G protein and bovine β-arr to the core and phosphorylated tail, respectively, of a single active human chimeric β-adrenergic receptor with the C-terminal tail of the arginine vasopressin type 2 receptor (βVR). All three components adopt their canonical active conformations, suggesting that a single megaplex GPCR is capable of simultaneously activating G protein and β-arr. Our findings provide a structural basis for GPCR-mediated sustained internalized G protein signaling. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9375.map.gz emd_9375.map.gz | 32.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9375-v30.xml emd-9375-v30.xml emd-9375.xml emd-9375.xml | 18.3 KB 18.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9375.png emd_9375.png | 151.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9375 http://ftp.pdbj.org/pub/emdb/structures/EMD-9375 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9375 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9375 | HTTPS FTP |
-Validation report
| Summary document |  emd_9375_validation.pdf.gz emd_9375_validation.pdf.gz | 396.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_9375_full_validation.pdf.gz emd_9375_full_validation.pdf.gz | 395.7 KB | Display | |
| Data in XML |  emd_9375_validation.xml.gz emd_9375_validation.xml.gz | 6 KB | Display | |
| Data in CIF |  emd_9375_validation.cif.gz emd_9375_validation.cif.gz | 6.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9375 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9375 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9375 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9375 | HTTPS FTP |
-Related structure data
| Related structure data |  6ni2MC  9376C  9377C  6ni3C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9375.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9375.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Stabilized beta-arrestin 1-V2T subcomplex of a GPCR-G protein-beta-arrestin mega-complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
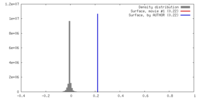
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Stabilized beta-arrestin 1-V2T subcomplex of a GPCR-G protein-bet...
+Supramolecule #1: Stabilized beta-arrestin 1-V2T subcomplex of a GPCR-G protein-bet...
+Supramolecule #2: Nanobody 32
+Supramolecule #3: Beta-arrestin-1
+Supramolecule #4: Fab30
+Supramolecule #5: Vasopressin 2 Tail
+Macromolecule #1: Nanobody 32
+Macromolecule #2: Beta-arrestin-1
+Macromolecule #3: Fab30 Heavy Chain
+Macromolecule #4: Fab30 Light Chain
+Macromolecule #5: Vasopressin V2 receptor
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Details: unidentified |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 104.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 4.0 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC / Number images used: 230021 |
|---|---|
| Initial angle assignment | Type: OTHER / Software - Name: cryoSPARC |
| Final angle assignment | Type: OTHER / Software - Name: cryoSPARC |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)