+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9036 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Full-length S. pombe Mdn1 in the presence of ATP and Rbin-1 | |||||||||
 Map data Map data | Full-length S. pombe Mdn1 in the presence of ATP and Rbin-1, primary map | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationrixosome complex / ribosome biogenesis / ribosomal large subunit assembly / calcium ion binding / nucleolus / ATP hydrolysis activity / nucleoplasm / ATP binding / nucleus Similarity search - Function | |||||||||
| Biological species |  | |||||||||
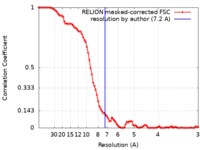
| Method | single particle reconstruction / cryo EM / Resolution: 7.2 Å | |||||||||
 Authors Authors | Chen Z / Suzuki H / Wang AC / DiMaio F / Walz T / Kapoor TM | |||||||||
 Citation Citation |  Journal: Cell / Year: 2018 Journal: Cell / Year: 2018Title: Structural Insights into Mdn1, an Essential AAA Protein Required for Ribosome Biogenesis. Authors: Zhen Chen / Hiroshi Suzuki / Yuki Kobayashi / Ashley C Wang / Frank DiMaio / Shigehiro A Kawashima / Thomas Walz / Tarun M Kapoor /   Abstract: Mdn1 is an essential AAA (ATPase associated with various activities) protein that removes assembly factors from distinct precursors of the ribosomal 60S subunit. However, Mdn1's large size (∼5,000 ...Mdn1 is an essential AAA (ATPase associated with various activities) protein that removes assembly factors from distinct precursors of the ribosomal 60S subunit. However, Mdn1's large size (∼5,000 amino acid [aa]) and its limited homology to other well-studied proteins have restricted our understanding of its remodeling function. Here, we present structures for S. pombe Mdn1 in the presence of AMPPNP at up to ∼4 Å or ATP plus Rbin-1, a chemical inhibitor, at ∼8 Å resolution. These data reveal that Mdn1's MIDAS domain is tethered to its ring-shaped AAA domain through an ∼20 nm long structured linker and a flexible ∼500 aa Asp/Glu-rich motif. We find that the MIDAS domain, which also binds other ribosome-assembly factors, docks onto the AAA ring in a nucleotide state-specific manner. Together, our findings reveal how conformational changes in the AAA ring can be directly transmitted to the MIDAS domain and thereby drive the targeted release of assembly factors from ribosomal 60S-subunit precursors. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9036.map.gz emd_9036.map.gz | 6.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9036-v30.xml emd-9036-v30.xml emd-9036.xml emd-9036.xml | 12.7 KB 12.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_9036_fsc.xml emd_9036_fsc.xml | 8.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_9036.png emd_9036.png | 75 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9036 http://ftp.pdbj.org/pub/emdb/structures/EMD-9036 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9036 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9036 | HTTPS FTP |
-Related structure data
| Related structure data |  6orbMC  9032C  9033C  9034C  9035C  6or5C  6or6C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9036.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9036.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Full-length S. pombe Mdn1 in the presence of ATP and Rbin-1, primary map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.5 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
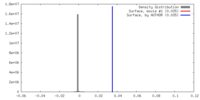
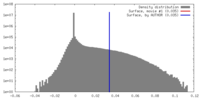
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Full-length Mdn1 in the presence of ATP and Rbin-1
| Entire | Name: Full-length Mdn1 in the presence of ATP and Rbin-1 |
|---|---|
| Components |
|
-Supramolecule #1: Full-length Mdn1 in the presence of ATP and Rbin-1
| Supramolecule | Name: Full-length Mdn1 in the presence of ATP and Rbin-1 / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 540 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.15 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - #0 - Film type ID: 1 / Support film - #0 - Material: CARBON / Support film - #0 - topology: HOLEY ARRAY / Support film - #1 - Film type ID: 2 / Support film - #1 - Material: CARBON / Support film - #1 - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 80 % / Chamber temperature: 295 K / Instrument: GATAN CRYOPLUNGE 3 |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Frames/image: 1-50 / Number real images: 3887 / Average exposure time: 15.0 sec. / Average electron dose: 66.7 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 28000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)