[English] 日本語
 Yorodumi
Yorodumi- EMDB-8902: Cryo-EM Structures of ASC and NLRC4 CARD Filaments Reveal a Unifi... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8902 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM Structures of ASC and NLRC4 CARD Filaments Reveal a Unified Mechanism of Nucleation and Activation of Caspase-1 | |||||||||
 Map data Map data | Helical reconstruction of ASC-CARD filament | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationNLRP6 inflammasome complex / myosin I binding / Pyrin domain binding / myeloid dendritic cell activation involved in immune response / positive regulation of antigen processing and presentation of peptide antigen via MHC class II / negative regulation of protein serine/threonine kinase activity / regulation of intrinsic apoptotic signaling pathway / myeloid dendritic cell activation / IkappaB kinase complex / The AIM2 inflammasome ...NLRP6 inflammasome complex / myosin I binding / Pyrin domain binding / myeloid dendritic cell activation involved in immune response / positive regulation of antigen processing and presentation of peptide antigen via MHC class II / negative regulation of protein serine/threonine kinase activity / regulation of intrinsic apoptotic signaling pathway / myeloid dendritic cell activation / IkappaB kinase complex / The AIM2 inflammasome / interleukin-6 receptor binding / AIM2 inflammasome complex / icosanoid biosynthetic process / NLRP1 inflammasome complex / macropinocytosis / canonical inflammasome complex / NLRP3 inflammasome complex assembly / BMP receptor binding / NLRP3 inflammasome complex / positive regulation of adaptive immune response / cysteine-type endopeptidase activator activity / CLEC7A/inflammasome pathway / negative regulation of interferon-beta production / osmosensory signaling pathway / regulation of tumor necrosis factor-mediated signaling pathway / positive regulation of extrinsic apoptotic signaling pathway / pattern recognition receptor signaling pathway / positive regulation of macrophage cytokine production / : / tropomyosin binding / pattern recognition receptor activity / positive regulation of release of cytochrome c from mitochondria / positive regulation of actin filament polymerization / intrinsic apoptotic signaling pathway by p53 class mediator / pyroptotic inflammatory response / positive regulation of activated T cell proliferation / positive regulation of interleukin-10 production / The NLRP3 inflammasome / cellular response to interleukin-1 / intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator / positive regulation of T cell migration / : / Purinergic signaling in leishmaniasis infection / positive regulation of chemokine production / positive regulation of defense response to virus by host / negative regulation of cytokine production involved in inflammatory response / activation of innate immune response / intrinsic apoptotic signaling pathway / positive regulation of phagocytosis / negative regulation of canonical NF-kappaB signal transduction / tumor necrosis factor-mediated signaling pathway / positive regulation of interleukin-1 beta production / positive regulation of interleukin-8 production / apoptotic signaling pathway / regulation of protein stability / positive regulation of non-canonical NF-kappaB signal transduction / : / protein homooligomerization / regulation of autophagy / positive regulation of JNK cascade / positive regulation of interleukin-6 production / positive regulation of type II interferon production / positive regulation of T cell activation / positive regulation of tumor necrosis factor production / positive regulation of inflammatory response / SARS-CoV-1 activates/modulates innate immune responses / azurophil granule lumen / cellular response to tumor necrosis factor / regulation of inflammatory response / cellular response to lipopolysaccharide / protease binding / secretory granule lumen / defense response to virus / defense response to Gram-negative bacterium / microtubule / transmembrane transporter binding / positive regulation of canonical NF-kappaB signal transduction / positive regulation of ERK1 and ERK2 cascade / protein dimerization activity / defense response to Gram-positive bacterium / positive regulation of apoptotic process / inflammatory response / Golgi membrane / innate immune response / neuronal cell body / apoptotic process / Neutrophil degranulation / nucleolus / enzyme binding / endoplasmic reticulum / signal transduction / protein homodimerization activity / protein-containing complex / mitochondrion / extracellular region / nucleoplasm / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.17 Å | |||||||||
 Authors Authors | Li Y / Fu T / Wu H | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2018 Journal: Proc Natl Acad Sci U S A / Year: 2018Title: Cryo-EM structures of ASC and NLRC4 CARD filaments reveal a unified mechanism of nucleation and activation of caspase-1. Authors: Yang Li / Tian-Min Fu / Alvin Lu / Kristen Witt / Jianbin Ruan / Chen Shen / Hao Wu /  Abstract: Canonical inflammasomes are cytosolic supramolecular complexes that activate caspase-1 upon sensing extrinsic microbial invasions and intrinsic sterile stress signals. During inflammasome assembly, ...Canonical inflammasomes are cytosolic supramolecular complexes that activate caspase-1 upon sensing extrinsic microbial invasions and intrinsic sterile stress signals. During inflammasome assembly, adaptor proteins ASC and NLRC4 recruit caspase-1 through homotypic caspase recruitment domain (CARD) interactions, leading to caspase-1 dimerization and activation. Activated caspase-1 processes proinflammatory cytokines and Gasdermin D to induce cytokine maturation and pyroptotic cell death. Here, we present cryo-electron microscopy (cryo-EM) structures of NLRC4 CARD and ASC CARD filaments mediated by conserved three types of asymmetric interactions (types I, II, and III). We find that the CARDs of these two adaptor proteins share a similar assembly pattern, which matches that of the caspase-1 CARD filament whose structure we defined previously. These data indicate a unified mechanism for downstream caspase-1 recruitment through CARD-CARD interactions by both adaptors. Using structure modeling, we further show that full-length NLRC4 assembles via two separate symmetries at its CARD and its nucleotide-binding domain (NBD), respectively. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8902.map.gz emd_8902.map.gz | 25.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8902-v30.xml emd-8902-v30.xml emd-8902.xml emd-8902.xml | 9.4 KB 9.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_8902.png emd_8902.png | 79.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8902 http://ftp.pdbj.org/pub/emdb/structures/EMD-8902 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8902 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8902 | HTTPS FTP |
-Related structure data
| Related structure data |  6n1hMC  8903C  6n1iC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_8902.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8902.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Helical reconstruction of ASC-CARD filament | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.32 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
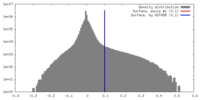
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Caspase recruitment domain of Apoptosis-associated speck-like pro...
| Entire | Name: Caspase recruitment domain of Apoptosis-associated speck-like protein containing a CARD |
|---|---|
| Components |
|
-Supramolecule #1: Caspase recruitment domain of Apoptosis-associated speck-like pro...
| Supramolecule | Name: Caspase recruitment domain of Apoptosis-associated speck-like protein containing a CARD type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism: |
-Macromolecule #1: Apoptosis-associated speck-like protein containing a CARD
| Macromolecule | Name: Apoptosis-associated speck-like protein containing a CARD type: protein_or_peptide / ID: 1 / Number of copies: 8 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 9.775143 KDa |
| Recombinant expression | Organism: |
| Sequence | String: LHFIDQHRAA LIARVTNVEW LLDALYGKVL TDEQYQAVRA EPTNPSKMRK LFSFTPAWNW TCKDLLLQAL RESQSYLVED LER |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 41.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 5.0 Å Applied symmetry - Helical parameters - Δ&Phi: -100.58 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Resolution.type: BY AUTHOR / Resolution: 3.17 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 264167 |
|---|---|
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller





















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)