+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7542 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the nvTRPM2 channel in complex with Ca2+ | |||||||||
 Map data Map data | Structure of the nvTRPM2 channel in complex with Ca2+ | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Ion channel / TRPM2 / Ca2+ binding / Nematostella vectensis / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationADP-ribose diphosphatase activity / ligand-gated calcium channel activity / ligand-gated sodium channel activity / sodium ion transmembrane transport / calcium ion transmembrane transport / metal ion binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Nematostella vectensis (starlet sea anemone) Nematostella vectensis (starlet sea anemone) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.07 Å | |||||||||
 Authors Authors | Zhang Z / Toth B | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2018 Journal: Elife / Year: 2018Title: Structure of a TRPM2 channel in complex with Ca explains unique gating regulation. Authors: Zhe Zhang / Balázs Tóth / Andras Szollosi / Jue Chen / László Csanády /   Abstract: Transient receptor potential melastatin 2 (TRPM2) is a Ca-permeable cation channel required for immune cell activation, insulin secretion, and body heat control. TRPM2 is activated by cytosolic Ca, ...Transient receptor potential melastatin 2 (TRPM2) is a Ca-permeable cation channel required for immune cell activation, insulin secretion, and body heat control. TRPM2 is activated by cytosolic Ca, phosphatidyl-inositol-4,5-bisphosphate and ADP ribose. Here, we present the ~3 Å resolution electron cryo-microscopic structure of TRPM2 from , 63% similar in sequence to human TRPM2, in the Ca-bound closed state. Compared to other TRPM channels, TRPM2 exhibits unique structural features that correlate with its function. The pore is larger and more negatively charged, consistent with its high Ca selectivity and larger conductance. The intracellular Ca binding sites are connected to the pore and cytosol, explaining the unusual dependence of TRPM2 activity on intra- and extracellular Ca. In addition, the absence of a post-filter motif is likely the cause of the rapid inactivation of human TRPM2. Together, our cryo-EM and electrophysiology studies provide a molecular understanding of the unique gating mechanism of TRPM2. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7542.map.gz emd_7542.map.gz | 202.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7542-v30.xml emd-7542-v30.xml emd-7542.xml emd-7542.xml | 14.5 KB 14.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7542.png emd_7542.png | 201.6 KB | ||
| Filedesc metadata |  emd-7542.cif.gz emd-7542.cif.gz | 6.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7542 http://ftp.pdbj.org/pub/emdb/structures/EMD-7542 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7542 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7542 | HTTPS FTP |
-Related structure data
| Related structure data |  6co7MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_7542.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7542.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Structure of the nvTRPM2 channel in complex with Ca2+ | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.03 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
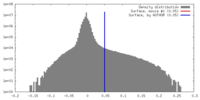
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : nvTRPM2 tetramer
| Entire | Name: nvTRPM2 tetramer |
|---|---|
| Components |
|
-Supramolecule #1: nvTRPM2 tetramer
| Supramolecule | Name: nvTRPM2 tetramer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Nematostella vectensis (starlet sea anemone) Nematostella vectensis (starlet sea anemone) |
-Macromolecule #1: Predicted protein
| Macromolecule | Name: Predicted protein / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Nematostella vectensis (starlet sea anemone) Nematostella vectensis (starlet sea anemone) |
| Molecular weight | Theoretical: 176.663438 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGKDSFTPLY DGGDSSHVHL NKFGSNQLSQ SKKSWIARNF SRRECIRFVP KSHDVSRCKC GRPRERHSQQ ALESGQGSEE WNVASCTTK HPTNAYGEID FEGYGGQKRA PYLRMSHDTD ANLVITLMLK RWNLEIPNLV ISVTGGAKSF VLKPRLREMF R RGLIKAAK ...String: MGKDSFTPLY DGGDSSHVHL NKFGSNQLSQ SKKSWIARNF SRRECIRFVP KSHDVSRCKC GRPRERHSQQ ALESGQGSEE WNVASCTTK HPTNAYGEID FEGYGGQKRA PYLRMSHDTD ANLVITLMLK RWNLEIPNLV ISVTGGAKSF VLKPRLREMF R RGLIKAAK TTGAWIITGG TNTGVMKHVG EAVKEQQLMF GSDTQVNVIG IATWGIVDKQ SDLISEKNGK YPALYSMEPT PG HQGAMLD PNHSHFFLVD DGTEGKYGVE IGMRSRIEEA IMKVKTDSRS EAGSIGVPVV LLVLEGGPNT VATMYELIKK KVP AVVIDG SGRAASVVGF AYNHTIKRNV DGQTINVIDP QYEDEVRAKV VEVFGAKGAD KTYSMIKDVL EDEKMISVYS LDGE ISQDI DLAILKALLK ANRSSPVAQL NLALAWNRID LAKSDIFTEE QQWTTETLSA AMLTALLDDK AEFAELFLQN GLSMR EFLS LDILCKLYAE VPGNTTIKPL LQKEMGKRQV KTIDMDVVGE VIEELMGDMF ESYYRKDGHY FGELASYAEG LVLKNR KSS KDLLANINRI DPLPTPYLDV FLWAVLCNRR ELARVLWEAG REPMAAALMA SRLLKRMASR AQEDNTITDI SSDLYDH AR LFEERAVGVL DECFNENETL SQTLLVRELD HYSRMTALEL AVSAESQDFI AHTSCQVLLT RLWMGTMAMN TRWWKVLV C LYLPVLIFPI IYFVPDEQHE RQAAEREHQK SLNQKSSKVK SHKEKNDAPV VPVYRSKEEK AVSNDEEARV GTENEEEDF QLEDYIPEIR EDDSMEVIMR NKKLGFCDRI MHFYSAPFSK FVGNVVGYLA FIFLYAYVVL FNFPRFDPAK TLGGIHPTEI VLYFWVFTI LIEEIRQLAA KPPKYIKDKV SVYFSDTWNF VDIFSLTVFI IAIILRFFTN SRIFTASRII LSLDIIFFIV R SLQIFSVN RLLGPKLVMI QKMMQDLAQF IIILAVFTIA YGIALHAVMF PSPGIYARNN TWVTITSVVQ YPYWQMYGEL FL DEIQGEK PKEFGEVDPD GRWLSPLLLA IYMVFTNILL LNLLIAIFNY TFERVQEDSD KVWKFQRYDL VQEYHSRPVF APP LVLLGH ILIFIRWVWR MCRCGHPPRG STMKIGLSPA EMEQMDNWEF QAAEMYIHQQ QQKNSGTLEE RVRALGDRVD CINS QLNRV LDSMSGTRAH ALTDGNGLEG GHDSEGRLAR MEVELSSNSE SLQKILALLQ QQPPVKGQAA VPIQLTLLHY KARSS PYPG STAKRFAVQD NMVDWQVPFP DYKPVNYTAP VVLANPVWAD KDLMAMSPRP ELPYNQMDHT CNVNRVSYNG TYVVKD GLP LNPMGRTGMQ GRGLLGRFGP NHAADPVVTR WKRTSAGVML QGGKKVLEFV AIQRKDNNQW AIPGGMVEPG QLVTQAL KA EFGEEAMAKL NVSQEEKERI AKQIERLFQQ GQEIYKGYVD DPRNTDNAWM ETVAVNFHDD KGDLFGDITL QAGDDAAA V RWQRVSGNIP LYASHVSILE KVAKMRDAAF SNSLEVLFQ UniProtKB: Transient receptor potential cation channel subfamily M member-like 2 |
-Macromolecule #3: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 3 / Number of copies: 4 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Macromolecule #4: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(tri...
| Macromolecule | Name: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate type: ligand / ID: 4 / Number of copies: 56 / Formula: POV |
|---|---|
| Molecular weight | Theoretical: 760.076 Da |
| Chemical component information |  ChemComp-POV: |
-Macromolecule #5: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 5 / Number of copies: 4 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #6: SODIUM ION
| Macromolecule | Name: SODIUM ION / type: ligand / ID: 6 / Number of copies: 3 |
|---|---|
| Molecular weight | Theoretical: 22.99 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 400 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 20 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 298 K / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Dimensions - Width: 3710 pixel / Digitization - Dimensions - Height: 3838 pixel / Digitization - Frames/image: 1-50 / Number grids imaged: 1 / Number real images: 1550 / Average exposure time: 0.2 sec. / Average electron dose: 1.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated defocus max: 2.5 µm / Calibrated defocus min: 0.7000000000000001 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-6co7: |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)