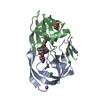
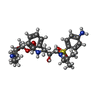
Entry Database : PDB / ID : 5bs4Title HIV-1 wild Type protease with GRL-047-11A (a methylamine bis-Tetrahydrofuran P2-Ligand, 4-amino sulfonamide derivative) Protease Keywords / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Method / / / Resolution : 1.29 Å Authors Wang, Y.-F. / Agniswamy, J. / Weber, I.T. Funding support Organization Grant number Country National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) GM53386 National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) GM62920 Department of Energy (DOE, United States) W-31-109-Eng-38 National Institutes of Health/National Cancer Institute (NIH/NCI) by the Intramural Research Program of the Center for Cancer Research the Ministry of Education, Culture, Sports, Science, and Technology by a Grant-in-Aid for Scientific Research (Priority Areas) the Ministry of Education, Culture, Sports, Science, and Technology the Grant to the Cooperative Research Project on Clinical and Epidemiological Studies of Emerging and Reemerging Infectious Diseases (Renkei Jigyo) Ministry of Health, Welfare, and Labor Grant for Promotion of AIDS Research
Journal : J.Med.Chem. / Year : 2015Title : Design of HIV-1 Protease Inhibitors with Amino-bis-tetrahydrofuran Derivatives as P2-Ligands to Enhance Backbone-Binding Interactions: Synthesis, Biological Evaluation, and Protein-Ligand X-ray Studies.Authors : Ghosh, A.K. / Martyr, C.D. / Osswald, H.L. / Sheri, V.R. / Kassekert, L.A. / Chen, S. / Agniswamy, J. / Wang, Y.F. / Hayashi, H. / Aoki, M. / Weber, I.T. / Mitsuya, H. History Deposition Jun 1, 2015 Deposition site / Processing site Revision 1.0 Sep 9, 2015 Provider / Type Revision 1.1 Sep 23, 2015 Group Revision 1.2 Sep 20, 2017 Group / Database references / Derived calculationsCategory / pdbx_audit_support / pdbx_struct_oper_listItem / _pdbx_audit_support.funding_organization / _pdbx_struct_oper_list.symmetry_operationRevision 1.3 Dec 4, 2019 Group / Category / Item Revision 1.4 Sep 27, 2023 Group Data collection / Database references ... Data collection / Database references / Derived calculations / Refinement description Category chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model / pdbx_struct_conn_angle / struct_conn Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.value / _struct_conn.pdbx_dist_value / _struct_conn.ptnr2_auth_seq_id
Show all Show less
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Human immunodeficiency virus type 1 BH10
Human immunodeficiency virus type 1 BH10 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.29 Å
MOLECULAR REPLACEMENT / Resolution: 1.29 Å  Authors
Authors United States,
United States,  Japan, 7items
Japan, 7items  Citation
Citation Journal: J.Med.Chem. / Year: 2015
Journal: J.Med.Chem. / Year: 2015 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 5bs4.cif.gz
5bs4.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb5bs4.ent.gz
pdb5bs4.ent.gz PDB format
PDB format 5bs4.json.gz
5bs4.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/bs/5bs4
https://data.pdbj.org/pub/pdb/validation_reports/bs/5bs4 ftp://data.pdbj.org/pub/pdb/validation_reports/bs/5bs4
ftp://data.pdbj.org/pub/pdb/validation_reports/bs/5bs4
 Links
Links Assembly
Assembly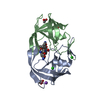
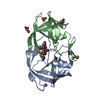
 Components
Components Human immunodeficiency virus type 1 BH10
Human immunodeficiency virus type 1 BH10








 X-RAY DIFFRACTION
X-RAY DIFFRACTION Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  APS
APS  / Beamline: 22-BM / Wavelength: 1 Å
/ Beamline: 22-BM / Wavelength: 1 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller






















 PDBj
PDBj






