+ Open data
Open data
- Basic information
Basic information
| Entry | 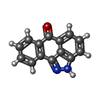 Database: PDB chemical components / ID: 537 Database: PDB chemical components / ID: 537 | ||
|---|---|---|---|
| Name | Name: | Comment | inhibitor*YM | |
-Chemical information
| Composition |  | ||||||
|---|---|---|---|---|---|---|---|
| Others | Type: NON-POLYMER / PDB classification: HETAIN / Three letter code: 537 / Model coordinates PDB-ID: 1PMV | ||||||
| History |
| ||||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  BindingDB BindingDB  / /  Brenda Brenda      / /  ChEBI / ChEBI /  ChEMBL / ChEMBL /  ChemicalBook / ChemicalBook /  CompTox / CompTox /  DrugBank / DrugBank /  GtoPharmacology / GtoPharmacology /  HMDB / HMDB /  Nikkaji / Nikkaji /  PubChem / PubChem /  PubChem_TPharma / PubChem_TPharma /  SureChEMBL / SureChEMBL /  ZINC / ZINC /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 10.04 | | CACTVS 3.341 | OpenEye OEToolkits 1.5.0 | |
|---|
-SMILES CANONICAL
| CACTVS 3.341 | | OpenEye OEToolkits 1.5.0 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| ACDLabs 10.04 |
|---|
-PDB entries
Showing all 5 items

PDB-1pmv: 
The structure of JNK3 in complex with a dihydroanthrapyrazole inhibitor

PDB-1uki: 
Structural basis for the selective inhibition of JNK1 by the scaffolding protein JIP1 and SP600125

PDB-2zmd: 
Crystal structure of human Mps1 catalytic domain T686A mutant in complex with SP600125 inhibitor

PDB-4feu: 
Crystal structure of the aminoglycoside phosphotransferase APH(3')-Ia, with substrate kanamycin and small molecule inhibitor anthrapyrazolone SP600125

PDB-5lvl: 
Human PDK1 Kinase Domain in Complex with Compound PS653 Bound to the ATP-Binding Site
 Movie
Movie Controller
Controller



