+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4192 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Mycobacterium smegmatis RNA polymerase core | |||||||||
 Map data Map data | Mycobacterium smegmatis RNA polymerase core map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TRANSCRIPTION Sigma / TRANSCRIPTION | |||||||||
| Function / homology |  Function and homology information Function and homology informationDNA-directed RNA polymerase complex / ribonucleoside binding / DNA-directed RNA polymerase / DNA-directed RNA polymerase activity / protein dimerization activity / response to antibiotic / DNA-templated transcription / magnesium ion binding / DNA binding / zinc ion binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Mycobacterium smegmatis str. MC2 155 (bacteria) / Mycobacterium smegmatis str. MC2 155 (bacteria) /  Mycolicibacterium smegmatis MC2 155 (bacteria) Mycolicibacterium smegmatis MC2 155 (bacteria) | |||||||||
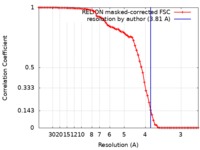
| Method | single particle reconstruction / cryo EM / Resolution: 3.81 Å | |||||||||
 Authors Authors | Kouba T / Barvik I | |||||||||
 Citation Citation |  Journal: J Bacteriol / Year: 2019 Journal: J Bacteriol / Year: 2019Title: The Core and Holoenzyme Forms of RNA Polymerase from . Authors: Tomáš Kouba / Jiří Pospíšil / Jarmila Hnilicová / Hana Šanderová / Ivan Barvík / Libor Krásný /   Abstract: Bacterial RNA polymerase (RNAP) is essential for gene expression and as such is a valid drug target. Hence, it is imperative to know its structure and dynamics. Here, we present two as-yet-unreported ...Bacterial RNA polymerase (RNAP) is essential for gene expression and as such is a valid drug target. Hence, it is imperative to know its structure and dynamics. Here, we present two as-yet-unreported forms of RNAP: core and holoenzyme containing σ but no other factors. Each form was detected by cryo-electron microscopy in two major conformations. Comparisons of these structures with known structures of other RNAPs reveal a high degree of conformational flexibility of the mycobacterial enzyme and confirm that region 1.1 of σ is directed into the primary channel of RNAP. Taken together, we describe the conformational changes of unrestrained mycobacterial RNAP. We describe here three-dimensional structures of core and holoenzyme forms of mycobacterial RNA polymerase (RNAP) solved by cryo-electron microscopy. These structures fill the thus-far-empty spots in the gallery of the pivotal forms of mycobacterial RNAP and illuminate the extent of conformational dynamics of this enzyme. The presented findings may facilitate future designs of antimycobacterial drugs targeting RNAP. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4192.map.gz emd_4192.map.gz | 3.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4192-v30.xml emd-4192-v30.xml emd-4192.xml emd-4192.xml | 21.4 KB 21.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4192_fsc.xml emd_4192_fsc.xml | 7.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_4192.png emd_4192.png | 81.4 KB | ||
| Filedesc metadata |  emd-4192.cif.gz emd-4192.cif.gz | 7.6 KB | ||
| Others |  emd_4192_half_map_1.map.gz emd_4192_half_map_1.map.gz emd_4192_half_map_2.map.gz emd_4192_half_map_2.map.gz | 26.2 MB 26.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4192 http://ftp.pdbj.org/pub/emdb/structures/EMD-4192 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4192 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4192 | HTTPS FTP |
-Related structure data
| Related structure data |  6f6wMC  3983C  6eydC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4192.map.gz / Format: CCP4 / Size: 34.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4192.map.gz / Format: CCP4 / Size: 34.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mycobacterium smegmatis RNA polymerase core map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.34 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: Mycobacterium smegmatis RNA polymerase core half1
| File | emd_4192_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mycobacterium smegmatis RNA polymerase core half1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Mycobacterium smegmatis RNA polymerase core half2
| File | emd_4192_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mycobacterium smegmatis RNA polymerase core half2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Mycobacterium smegmatis RNA polymerase Sigma-A holoenzyme
| Entire | Name: Mycobacterium smegmatis RNA polymerase Sigma-A holoenzyme |
|---|---|
| Components |
|
-Supramolecule #1: Mycobacterium smegmatis RNA polymerase Sigma-A holoenzyme
| Supramolecule | Name: Mycobacterium smegmatis RNA polymerase Sigma-A holoenzyme type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  Mycobacterium smegmatis str. MC2 155 (bacteria) Mycobacterium smegmatis str. MC2 155 (bacteria) |
| Molecular weight | Theoretical: 364 KDa |
-Macromolecule #1: DNA-directed RNA polymerase subunit alpha
| Macromolecule | Name: DNA-directed RNA polymerase subunit alpha / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: DNA-directed RNA polymerase |
|---|---|
| Source (natural) | Organism:  Mycolicibacterium smegmatis MC2 155 (bacteria) Mycolicibacterium smegmatis MC2 155 (bacteria) |
| Molecular weight | Theoretical: 37.959441 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MLISQRPTLS EETVAENRSR FVIEPLEPGF GYTLGNSLRR TLLSSIPGAA VTSIRIDGVL HEFTTVPGVK EDVTDIILNL KGLVVSSDD DEPVTMYLRK QGPGVVTAGD IVPPAGVTVH NPDMHIATLN DKGKLEVELV VERGRGYVPA VQNKASGAEI G RIPVDSIY ...String: MLISQRPTLS EETVAENRSR FVIEPLEPGF GYTLGNSLRR TLLSSIPGAA VTSIRIDGVL HEFTTVPGVK EDVTDIILNL KGLVVSSDD DEPVTMYLRK QGPGVVTAGD IVPPAGVTVH NPDMHIATLN DKGKLEVELV VERGRGYVPA VQNKASGAEI G RIPVDSIY SPVLKVTYKV EATRVEQRTD FDKLIIDVET KNSISPRDAL ASAGGTLVEL FGLARELNAD SEHIEIGPSP AE ADHIASF ALPIDDLDLT VRSYNCLKRE GVHTVGELVA RTESDLLDIR NFGQKSIDEV KIKLHQLGLS LKDSPATFDP SEV AGYDAA TGTWTSDAGY DLDDNQDYAE TEQL UniProtKB: DNA-directed RNA polymerase subunit alpha |
-Macromolecule #2: DNA-directed RNA polymerase subunit beta
| Macromolecule | Name: DNA-directed RNA polymerase subunit beta / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: DNA-directed RNA polymerase |
|---|---|
| Source (natural) | Organism:  Mycolicibacterium smegmatis MC2 155 (bacteria) Mycolicibacterium smegmatis MC2 155 (bacteria) |
| Molecular weight | Theoretical: 129.456938 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: VLEGCILAVS SQSKSNAITN NSVPGAPNRV SFAKLREPLE VPGLLDVQTD SFEWLVGSDR WRQAAIDRGE ENPVGGLEEV LAELSPIED FSGSMSLSFS DPRFDEVKAS VDECKDKDMT YAAPLFVTAE FINNNTGEIK SQTVFMGDFP MMTEKGTFII N GTERVVVS ...String: VLEGCILAVS SQSKSNAITN NSVPGAPNRV SFAKLREPLE VPGLLDVQTD SFEWLVGSDR WRQAAIDRGE ENPVGGLEEV LAELSPIED FSGSMSLSFS DPRFDEVKAS VDECKDKDMT YAAPLFVTAE FINNNTGEIK SQTVFMGDFP MMTEKGTFII N GTERVVVS QLVRSPGVYF DETIDKSTEK TLHSVKVIPG RGAWLEFDVD KRDTVGVRID RKRRQPVTVL LKALGWTNEQ IV ERFGFSE IMMGTLEKDT TSGTDEALLD IYRKLRPGEP PTKESAQTLL ENLFFKEKRY DLARVGRYKV NKKLGLNAGK PIT SSTLTE EDVVATIEYL VRLHEGQTSM TVPGGVEVPV EVDDIDHFGN RRLRTVGELI QNQIRVGLSR MERVVRERMT TQDV EAITP QTLINIRPVV AAIKEFFGTS QLSQFMDQNN PLSGLTHKRR LSALGPGGLS RERAGLEVRD VHPSHYGRMC PIETP EGPN IGLIGSLSVY ARVNPFGFIE TPYRKVENGV VTDQIDYLTA DEEDRHVVAQ ANSPTDENGR FTEDRVMVRK KGGEVE FVS ADQVDYMDVS PRQMVSVATA MIPFLEHDDA NRALMGANMQ RQAVPLVRSE APLVGTGMEL RAAIDAGDVV VADKTGV IE EVSADYITVM ADDGTRQSYR LRKFARSNHG TCANQRPIVD AGQRVEAGQV IADGPCTQNG EMALGKNLLV AIMPWEGH N YEDAIILSNR LVEEDVLTSI HIEEHEIDAR DTKLGAEEIT RDIPNVSDEV LADLDERGIV RIGAEVRDGD ILVGKVTPK GETELTPEER LLRAIFGEKA REVRDTSLKV PHGESGKVIG IRVFSREDDD ELPAGVNELV RVYVAQKRKI SDGDKLAGRH GNKGVIGKI LPVEDMPFLP DGTPVDIILN THGVPRRMNI GQILETHLGW VAKAGWNIDV AAGVPDWASK LPEELYSAPA D STVATPVF DGAQEGELAG LLGSTLPNRD GEVMVDADGK STLFDGRSGE PFPYPVTVGY MYILKLHHLV DDKIHARSTG PY SMITQQP LGGKAQFGGQ RFGEMECWAM QAYGAAYTLQ ELLTIKSDDT VGRVKVYEAI VKGENIPEPG IPESFKVLLK ELQ SLCLNV EVLSSDGAAI EMRDGDDEDL ERAAANLGIN LSRNESASVE DLALARHGGS GA UniProtKB: DNA-directed RNA polymerase subunit beta |
-Macromolecule #3: DNA-directed RNA polymerase subunit beta'
| Macromolecule | Name: DNA-directed RNA polymerase subunit beta' / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO / EC number: DNA-directed RNA polymerase |
|---|---|
| Source (natural) | Organism:  Mycolicibacterium smegmatis MC2 155 (bacteria) Mycolicibacterium smegmatis MC2 155 (bacteria) |
| Molecular weight | Theoretical: 147.785953 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: VLDVNFFDEL RIGLATADDI RNWSYGEVKK PETINYRTLK PEKDGLFCEK IFGPTRDWEC YCGKYKRVRF KGIICERCGV EVTRAKVRR ERMGHIELAA PVTHIWYFKG VPSRLGYLLD LAPKDLEKII YFAAYVITSV DDEMRHNELS TLEAEMAVEK K AVEDQRDA ...String: VLDVNFFDEL RIGLATADDI RNWSYGEVKK PETINYRTLK PEKDGLFCEK IFGPTRDWEC YCGKYKRVRF KGIICERCGV EVTRAKVRR ERMGHIELAA PVTHIWYFKG VPSRLGYLLD LAPKDLEKII YFAAYVITSV DDEMRHNELS TLEAEMAVEK K AVEDQRDA DLEARAQKLE ADLAELEAEG AKSDVRRKVR DSGEREMRQL RDRAQRELDR LDEIWNTFTK LAPKQLIVDE VL YRELQDR YGEYFTGAMG AESIKKLIEN FDIDAEAESL REVIRSGKGQ KKLRALKRLK VVAAFQQSGN SPMGMVLDAV PVI PPELRP MVQLDGGRFA TSDLNDLYRR VINRNNRLKR LIDLGAPEII VNNEKRMLQE SVDALFDNGR RGRPVTGPGN RPLK SLSDL LKGKQGRFRQ NLLGKRVDYS GRSVIVVGPQ LKLHQCGLPK LMALELFKPF VMKRLVDLNH AQNIKSAKRM VERQR PQVW DVLEEVIAEH PVLLNRAPTL HRLGIQAFEP QLVEGKAIQL HPLVCEAFNA DFDGDQMAVH LPLSAEAQAE ARILML SSN NILSPASGKP LAMPRLDMVT GLYYLTTLVE GATGEYQAAT KDAPEQGVYS SPAEAIMAMD RGALSVRAKI KVRLTEL RP PTDLEAQLFE NGWKPGDAWT AETTLGRVMF NELLPKSYPF VNEQMHKKVQ ARIINDLAER FPMIVVAQTV DKLKDAGF Y WATRSGVTVS MADVLVPPQK QEILERHEAE ADAIERKYQR GALNHTERNE SLVKIWQDAT EEVGKALEEF YPADNPIIT IVKSGATGNL TQTRTLAGMK GLVTNPKGEF IPRPIKSSFR EGLTVLEYFI NTHGARKGLA DTALRTADSG YLTRRLVDVS QDVIVREHD CETERGINVT LAERGPDGTL IRDAHVETSA FARTLATDAV DANGNVIIER GHDLGDPAID ALLAAGITTV K VRSVLTCT SATGVCAMCY GRSMATGKLV DIGEAVGIVA AQSIGEPGTQ LTMRTFHQGG VTGGADIVGG LPRVQELFEA RV PRNKAPI ADVAGRVRLE ESDKFFKITI VPDDGGEEVV YDKLSKRQRL RVITHEDGTE GVLSDGDHVE VGDQLMEGAA DPH EVLRVQ GPREVQIHLV KEVQEVYRAQ GVSIHDKHIE VIVRQMLRRV TIIDSGSTEF LPGSLTERAE FEAENRRVVA EGGE PAAGR PVLMGITKAS LATDSWLSAA SFQETTRVLT DAAINCRSDK LNGLKENVII GKLIPAGTGI SRYRNIQVQP TEEAR AAAY TIPSYEDQYY SPDFGQATGA AVPLDDYGYS DYRHHHHHHH H UniProtKB: DNA-directed RNA polymerase subunit beta' |
-Macromolecule #4: DNA-directed RNA polymerase subunit omega
| Macromolecule | Name: DNA-directed RNA polymerase subunit omega / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO / EC number: DNA-directed RNA polymerase |
|---|---|
| Source (natural) | Organism:  Mycolicibacterium smegmatis MC2 155 (bacteria) Mycolicibacterium smegmatis MC2 155 (bacteria) |
| Molecular weight | Theoretical: 11.512698 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: VSTPHADAQL NAADDLGIDS SAASAYDTPL GITNPPIDEL LSRASSKYAL VIYAAKRARQ INDYYNQLGD GILEYVGPLV EPGLQEKPL SIALREIHGD LLEHTEGE UniProtKB: DNA-directed RNA polymerase subunit omega |
-Macromolecule #5: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 5 / Number of copies: 2 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Macromolecule #6: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 6 / Number of copies: 1 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.8 |
|---|---|
| Grid | Material: GRAPHENE OXIDE |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: INTEGRATING / Average exposure time: 2.0 sec. / Average electron dose: 52.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 3.8000000000000003 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 78000 |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: RECIPROCAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-6f6w: |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)