+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3j06 | ||||||
|---|---|---|---|---|---|---|---|
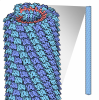
| Title | CryoEM Helical Reconstruction of TMV | ||||||
 Components Components |
| ||||||
 Keywords Keywords | VIRUS / RNA | ||||||
| Function / homology |  Function and homology information Function and homology informationhelical viral capsid / structural molecule activity / identical protein binding Similarity search - Function | ||||||
| Biological species |   Tobacco mosaic virus Tobacco mosaic virus | ||||||
| Method | ELECTRON MICROSCOPY / helical reconstruction / cryo EM / Resolution: 3.3 Å | ||||||
 Authors Authors | Ge, P. / Zhou, Z.H. | ||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2011 Journal: Proc Natl Acad Sci U S A / Year: 2011Title: Hydrogen-bonding networks and RNA bases revealed by cryo electron microscopy suggest a triggering mechanism for calcium switches. Authors: Peng Ge / Z Hong Zhou /  Abstract: Helical assemblies such as filamentous viruses, flagella, and F-actin represent an important category of structures in biology. As the first discovered virus, tobacco mosaic virus (TMV) was at the ...Helical assemblies such as filamentous viruses, flagella, and F-actin represent an important category of structures in biology. As the first discovered virus, tobacco mosaic virus (TMV) was at the center of virus research. Previously, the structure of TMV was solved at atomic detail by X-ray fiber diffraction but only for its dormant or high-calcium-concentration state, not its low-calcium-concentration state, which is relevant to viral assembly and disassembly inside host cells. Here we report a helical reconstruction of TMV in its calcium-free, metastable assembling state at 3.3 Å resolution by cryo electron microscopy, revealing both protein side chains and RNA bases. An atomic model was built de novo showing marked differences from the high-calcium, dormant-state structure. Although it could be argued that there might be inaccuracies in the latter structure derived from X-ray fiber diffraction, these differences can be interpreted as conformational changes effected by calcium-driven switches, a common regulatory mechanism in plant viruses. Our comparisons of the structures of the low- and high-calcium states indicate that hydrogen bonds formed by Asp116 and Arg92 in the place of the calcium ion of the dormant (high-calcium) state might trigger allosteric changes in the RNA base-binding pockets of the coat protein. In turn, the coat protein-RNA interactions in our structure favor an adenine-X-guanine (A*G) motif over the G*A motif of the dormant state, thus offering an explanation underlying viral assembly initiation by an AAG motif. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3j06.cif.gz 3j06.cif.gz | 43 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3j06.ent.gz pdb3j06.ent.gz | 29.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3j06.json.gz 3j06.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  3j06_validation.pdf.gz 3j06_validation.pdf.gz | 781.1 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  3j06_full_validation.pdf.gz 3j06_full_validation.pdf.gz | 798.7 KB | Display | |
| Data in XML |  3j06_validation.xml.gz 3j06_validation.xml.gz | 17.6 KB | Display | |
| Data in CIF |  3j06_validation.cif.gz 3j06_validation.cif.gz | 23 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/j0/3j06 https://data.pdbj.org/pub/pdb/validation_reports/j0/3j06 ftp://data.pdbj.org/pub/pdb/validation_reports/j0/3j06 ftp://data.pdbj.org/pub/pdb/validation_reports/j0/3j06 | HTTPS FTP |
-Related structure data
| Related structure data |  5185MC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 | x 49
|
| 2 |
|
| 3 | 
|
| Symmetry | Helical symmetry: (Circular symmetry: 1 / Dyad axis: no / N subunits divisor: 1 / Num. of operations: 49 / Rise per n subunits: 1.408 Å / Rotation per n subunits: 22.035 °) |
- Components
Components
| #1: Protein | Mass: 17636.621 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Tobacco mosaic virus / Strain: vulgare / References: UniProt: P69687 Tobacco mosaic virus / Strain: vulgare / References: UniProt: P69687 |
|---|---|
| #2: RNA chain | Mass: 935.620 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Details: viral RNA / Source: (natural)   Tobacco mosaic virus / Strain: vulgare Tobacco mosaic virus / Strain: vulgare |
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: FILAMENT / 3D reconstruction method: helical reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Tobacco Mosaic Virus (Vulgare) / Type: VIRUS / Details: 49 coat proteins in 3 turn right hand helix |
|---|---|
| Molecular weight | Value: 37.8 MDa / Experimental value: NO |
| Details of virus | Host category: PLANTAE (HIGHER PLANTS) / Isolate: STRAIN / Type: VIRION |
| Natural host | Organism: Nicotiana tabacum |
| Buffer solution | Name: Tris-buffered saline / pH: 7.4 / Details: 10 mM Tris, 130 mM sodium chloride |
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES / Details: 10 mM Tris, 130 mM sodium chloride |
| Specimen support | Details: 400 mesh Quantifoil 1.3/1.2 micro-meter |
| Vitrification | Instrument: FEI VITROBOT MARK IV / Cryogen name: ETHANE / Temp: 93 K / Humidity: 100 % / Details: 2 uL sample Method: 10 second wait time, 1-2 second blot time, 1-2 second drain time, blot force 1 |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS / Date: Jun 19, 2009 |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM / Electron beam tilt params: 0 FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM / Electron beam tilt params: 0 |
| Electron lens | Mode: BRIGHT FIELD / Nominal magnification: 75000 X / Calibrated magnification: 73000 X / Nominal defocus max: 2500 nm / Nominal defocus min: 1200 nm / Cs: 2.7 mm Astigmatism: objective lens astigmatism was corrected at 250000x magnification Camera length: 0 mm |
| Specimen holder | Specimen holder model: SIDE ENTRY, EUCENTRIC / Specimen holder type: Eucentric / Temperature: 90 K / Tilt angle max: 0 ° / Tilt angle min: 0 ° |
| Image recording | Electron dose: 25 e/Å2 / Film or detector model: KODAK SO-163 FILM |
| Image scans | Num. digital images: 386 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Relative weight: 1 |
- Processing
Processing
| Software | Name: EMAN / Classification: refinement | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| EM software |
| ||||||||||||
| Helical symmerty | Angular rotation/subunit: 22.035 ° / Axial rise/subunit: 1.407 Å / Axial symmetry: C1 | ||||||||||||
| 3D reconstruction | Method: IHRSR / Resolution: 3.3 Å / Resolution method: OTHER / Nominal pixel size: 0.847 Å / Actual pixel size: 0.869 Å / Symmetry type: HELICAL | ||||||||||||
| Refinement step | Cycle: LAST
|
 Movie
Movie Controller
Controller









 PDBj
PDBj
