+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-3295 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Cryo-EM structure of human p97 bound UPCDC30245 inhibitor | |||||||||
 マップデータ マップデータ | Reconstruction of human p97 bound to UPCDC30245 inhibitor (bfactor corrected map). | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | cryo-electron microscopy / single-particle / p97 / AAA ATPase | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報protein binding / positive regulation of Lys63-specific deubiquitinase activity / flavin adenine dinucleotide catabolic process / positive regulation of oxidative phosphorylation / VCP-NSFL1C complex / endosome to lysosome transport via multivesicular body sorting pathway / protein-DNA covalent cross-linking repair / endoplasmic reticulum stress-induced pre-emptive quality control / cellular response to arsenite ion / Derlin-1 retrotranslocation complex ...protein binding / positive regulation of Lys63-specific deubiquitinase activity / flavin adenine dinucleotide catabolic process / positive regulation of oxidative phosphorylation / VCP-NSFL1C complex / endosome to lysosome transport via multivesicular body sorting pathway / protein-DNA covalent cross-linking repair / endoplasmic reticulum stress-induced pre-emptive quality control / cellular response to arsenite ion / Derlin-1 retrotranslocation complex / BAT3 complex binding / positive regulation of protein K63-linked deubiquitination / deubiquitinase activator activity / aggresome assembly / regulation of protein localization to chromatin / vesicle-fusing ATPase / mitotic spindle disassembly / VCP-NPL4-UFD1 AAA ATPase complex / NADH metabolic process / cellular response to misfolded protein / stress granule disassembly / K48-linked polyubiquitin modification-dependent protein binding / positive regulation of mitochondrial membrane potential / negative regulation of protein localization to chromatin / ubiquitin-modified protein reader activity / retrograde protein transport, ER to cytosol / regulation of aerobic respiration / positive regulation of ATP biosynthetic process / regulation of synapse organization / ATPase complex / ubiquitin-specific protease binding / ubiquitin-like protein ligase binding / MHC class I protein binding / RHOH GTPase cycle / autophagosome maturation / HSF1 activation / polyubiquitin modification-dependent protein binding / proteasomal protein catabolic process / Protein methylation / endoplasmic reticulum to Golgi vesicle-mediated transport / translesion synthesis / interstrand cross-link repair / negative regulation of smoothened signaling pathway / ERAD pathway / ATP metabolic process / endoplasmic reticulum unfolded protein response / Attachment and Entry / lipid droplet / viral genome replication / proteasome complex / Josephin domain DUBs / N-glycan trimming in the ER and Calnexin/Calreticulin cycle / Hh mutants are degraded by ERAD / macroautophagy / Defective CFTR causes cystic fibrosis / Hedgehog ligand biogenesis / Translesion Synthesis by POLH / positive regulation of protein-containing complex assembly / establishment of protein localization / ABC-family proteins mediated transport / ADP binding / autophagy / cytoplasmic stress granule / Aggrephagy / positive regulation of non-canonical NF-kappaB signal transduction / positive regulation of protein catabolic process / activation of cysteine-type endopeptidase activity involved in apoptotic process / KEAP1-NFE2L2 pathway / azurophil granule lumen / double-strand break repair / Ovarian tumor domain proteases / positive regulation of proteasomal ubiquitin-dependent protein catabolic process / positive regulation of canonical Wnt signaling pathway / E3 ubiquitin ligases ubiquitinate target proteins / Neddylation / site of double-strand break / cellular response to heat / ubiquitin-dependent protein catabolic process / regulation of apoptotic process / protein phosphatase binding / proteasome-mediated ubiquitin-dependent protein catabolic process / secretory granule lumen / ficolin-1-rich granule lumen / Attachment and Entry / protein ubiquitination / protein domain specific binding / intracellular membrane-bounded organelle / DNA repair / signaling receptor binding / glutamatergic synapse / lipid binding / ubiquitin protein ligase binding / DNA damage response / Neutrophil degranulation / endoplasmic reticulum membrane / perinuclear region of cytoplasm / endoplasmic reticulum / ATP hydrolysis activity / protein-containing complex / RNA binding 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) / unidentified (未定義) Homo sapiens (ヒト) / unidentified (未定義) | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 2.3 Å | |||||||||
 データ登録者 データ登録者 | Banerjee S / Bartesaghi A / Merk A / Rao P / Bulfer SL / Yan Y / Green N / Mroczkowski B / Neitz RJ / Wipf P ...Banerjee S / Bartesaghi A / Merk A / Rao P / Bulfer SL / Yan Y / Green N / Mroczkowski B / Neitz RJ / Wipf P / Falconieri V / Deshaies RJ / Milne JLS / Huryn D / Arkin M / Subramaniam S | |||||||||
 引用 引用 |  ジャーナル: Science / 年: 2016 ジャーナル: Science / 年: 2016タイトル: 2.3 Å resolution cryo-EM structure of human p97 and mechanism of allosteric inhibition. 著者: Soojay Banerjee / Alberto Bartesaghi / Alan Merk / Prashant Rao / Stacie L Bulfer / Yongzhao Yan / Neal Green / Barbara Mroczkowski / R Jeffrey Neitz / Peter Wipf / Veronica Falconieri / ...著者: Soojay Banerjee / Alberto Bartesaghi / Alan Merk / Prashant Rao / Stacie L Bulfer / Yongzhao Yan / Neal Green / Barbara Mroczkowski / R Jeffrey Neitz / Peter Wipf / Veronica Falconieri / Raymond J Deshaies / Jacqueline L S Milne / Donna Huryn / Michelle Arkin / Sriram Subramaniam /  要旨: p97 is a hexameric AAA+ adenosine triphosphatase (ATPase) that is an attractive target for cancer drug development. We report cryo-electron microscopy (cryo-EM) structures for adenosine diphosphate ...p97 is a hexameric AAA+ adenosine triphosphatase (ATPase) that is an attractive target for cancer drug development. We report cryo-electron microscopy (cryo-EM) structures for adenosine diphosphate (ADP)-bound, full-length, hexameric wild-type p97 in the presence and absence of an allosteric inhibitor at resolutions of 2.3 and 2.4 angstroms, respectively. We also report cryo-EM structures (at resolutions of ~3.3, 3.2, and 3.3 angstroms, respectively) for three distinct, coexisting functional states of p97 with occupancies of zero, one, or two molecules of adenosine 5'-O-(3-thiotriphosphate) (ATPγS) per protomer. A large corkscrew-like change in molecular architecture, coupled with upward displacement of the N-terminal domain, is observed only when ATPγS is bound to both the D1 and D2 domains of the protomer. These cryo-EM structures establish the sequence of nucleotide-driven structural changes in p97 at atomic resolution. They also enable elucidation of the binding mode of an allosteric small-molecule inhibitor to p97 and illustrate how inhibitor binding at the interface between the D1 and D2 domains prevents propagation of the conformational changes necessary for p97 function. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_3295.map.gz emd_3295.map.gz | 106.3 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-3295-v30.xml emd-3295-v30.xml emd-3295.xml emd-3295.xml | 12 KB 12 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_3295.png emd_3295.png | 253.4 KB | ||
| その他 |  emd_3295_additional_1.map.gz emd_3295_additional_1.map.gz | 106.4 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3295 http://ftp.pdbj.org/pub/emdb/structures/EMD-3295 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3295 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3295 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_3295_validation.pdf.gz emd_3295_validation.pdf.gz | 381.9 KB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_3295_full_validation.pdf.gz emd_3295_full_validation.pdf.gz | 381 KB | 表示 | |
| XML形式データ |  emd_3295_validation.xml.gz emd_3295_validation.xml.gz | 6.4 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3295 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3295 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3295 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3295 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_3295.map.gz / 形式: CCP4 / 大きさ: 113.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_3295.map.gz / 形式: CCP4 / 大きさ: 113.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Reconstruction of human p97 bound to UPCDC30245 inhibitor (bfactor corrected map). | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 0.637 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
-添付マップデータ: emd 3295 additional 1.map
| ファイル | emd_3295_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
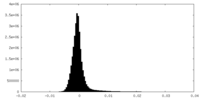
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : Full-length human p97 bound to UPCDC30245 inhibitor
| 全体 | 名称: Full-length human p97 bound to UPCDC30245 inhibitor |
|---|---|
| 要素 |
|
-超分子 #1000: Full-length human p97 bound to UPCDC30245 inhibitor
| 超分子 | 名称: Full-length human p97 bound to UPCDC30245 inhibitor / タイプ: sample / ID: 1000 / 詳細: The sample was monodisperse. / 集合状態: hexamer / Number unique components: 2 |
|---|---|
| 分子量 | 理論値: 540 KDa / 手法: sequence |
-分子 #1: p97/VCP Transitional endoplasmic reticulum ATPase
| 分子 | 名称: p97/VCP Transitional endoplasmic reticulum ATPase / タイプ: protein_or_peptide / ID: 1 / Name.synonym: p97 / コピー数: 1 / 集合状態: hexamer / 組換発現: Yes |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) / 別称: Human / 細胞中の位置: cytoplasm Homo sapiens (ヒト) / 別称: Human / 細胞中の位置: cytoplasm |
| 分子量 | 理論値: 540 KDa |
| 組換発現 | 生物種: Bacteria (unknown) / 組換細胞: E. coli / 組換プラスミド: Rosetta2 (DE3) |
| 配列 | UniProtKB: Transitional endoplasmic reticulum ATPase GO: ATP binding, signaling receptor binding, protein binding, lipid binding |
-分子 #2: 3-(5-fluoro-1H-indol-2-yl)phenyl)-N-(2-(4-isopropylpiperazin-1-yl...
| 分子 | 名称: 3-(5-fluoro-1H-indol-2-yl)phenyl)-N-(2-(4-isopropylpiperazin-1-yl)ethyl)piperidin-4-amine タイプ: ligand / ID: 2 / Name.synonym: UPCDC30245 / コピー数: 6 / 集合状態: monomer / 組換発現: No |
|---|---|
| 由来(天然) | 生物種: unidentified (未定義) |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 0.9 mg/mL |
|---|---|
| 緩衝液 | pH: 8 / 詳細: 25 mM Tris, 150 mM NaCl, 1 mM MgCl2, 1.0 mM TCEP |
| グリッド | 詳細: 200 mesh Quantifoil R1.2/1/3 grids |
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 90.15 K / 装置: FEI VITROBOT MARK IV / 手法: Blot for 2.5 seconds before plunging. |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 温度 | 最低: 79.6 K / 最高: 79.8 K / 平均: 79.7 K |
| 特殊光学系 | エネルギーフィルター - 名称: Gatan, Inc. エネルギーフィルター - エネルギー下限: 0.0 eV エネルギーフィルター - エネルギー上限: 20.0 eV |
| 詳細 | Parallel beam illumination |
| 日付 | 2015年2月16日 |
| 撮影 | カテゴリ: CCD フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 実像数: 2068 / 平均電子線量: 40 e/Å2 詳細: Every image is the average of 38 frames recorded by the direct electron detector. |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 倍率(補正後): 78490 / 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / Cs: 2.7 mm / 最大 デフォーカス(公称値): 2.5 µm / 最小 デフォーカス(公称値): 0.7 µm / 倍率(公称値): 215000 |
| 試料ステージ | 試料ホルダー: Liquid nitrogen cooled 試料ホルダーモデル: FEI TITAN KRIOS AUTOGRID HOLDER |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
- 画像解析
画像解析
| CTF補正 | 詳細: each particle |
|---|---|
| 最終 再構成 | 想定した対称性 - 点群: C6 (6回回転対称) / アルゴリズム: OTHER / 解像度のタイプ: BY AUTHOR / 解像度: 2.3 Å / 解像度の算出法: OTHER / ソフトウェア - 名称: FREALIGN 詳細: Map was corrected using a B-factor of -75 A^2. Unsharpened map is also part of this submission. 使用した粒子像数: 40913 |
 ムービー
ムービー コントローラー
コントローラー


































 Z
Z Y
Y X
X