+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-31500 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | ghrelin-bound ghrelin receptor in complex with Gq | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ghrelin / GPCR / Gq / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationghrelin receptor binding / positive regulation of bone development / positive regulation of gastric mucosal blood circulation / negative regulation of locomotion / cortisol secretion / growth hormone-releasing hormone activity / negative regulation of circadian sleep/wake cycle, REM sleep / regulation of response to food / positive regulation of small intestinal transit / regulation of gastric motility ...ghrelin receptor binding / positive regulation of bone development / positive regulation of gastric mucosal blood circulation / negative regulation of locomotion / cortisol secretion / growth hormone-releasing hormone activity / negative regulation of circadian sleep/wake cycle, REM sleep / regulation of response to food / positive regulation of small intestinal transit / regulation of gastric motility / regulation of transmission of nerve impulse / positive regulation of circadian sleep/wake cycle, non-REM sleep / positive regulation of cortisol secretion / positive regulation of corticotropin secretion / positive regulation of growth rate / gastric acid secretion / growth hormone secretion / neuronal dense core vesicle lumen / positive regulation of small intestine smooth muscle contraction / positive regulation of eating behavior / positive regulation of appetite / adult feeding behavior / positive regulation of growth hormone secretion / positive regulation of growth hormone receptor signaling pathway / actin polymerization or depolymerization / positive regulation of multicellular organism growth / cartilage development / positive regulation of synapse assembly / regulation of postsynapse organization / positive regulation of vascular endothelial cell proliferation / negative regulation of interleukin-1 beta production / positive regulation of sprouting angiogenesis / negative regulation of endothelial cell proliferation / decidualization / dendrite development / negative regulation of interleukin-6 production / protein tyrosine kinase activator activity / Synthesis, secretion, and deacylation of Ghrelin / negative regulation of tumor necrosis factor production / response to electrical stimulus / postsynaptic modulation of chemical synaptic transmission / hormone-mediated signaling pathway / response to hormone / positive regulation of adipose tissue development / synapse assembly / negative regulation of angiogenesis / Peptide ligand-binding receptors / positive regulation of insulin secretion involved in cellular response to glucose stimulus / excitatory postsynaptic potential / negative regulation of insulin secretion / G protein-coupled receptor binding / response to estrogen / negative regulation of inflammatory response / Schaffer collateral - CA1 synapse / glucose metabolic process / Olfactory Signaling Pathway / Activation of the phototransduction cascade / positive regulation of insulin secretion / G protein-coupled acetylcholine receptor signaling pathway / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / G beta:gamma signalling through BTK / photoreceptor disc membrane / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / Glucagon-type ligand receptors / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / cellular response to catecholamine stimulus / ADP signalling through P2Y purinoceptor 1 / ADORA2B mediated anti-inflammatory cytokines production / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / regulation of cell population proliferation / GPER1 signaling / cellular response to prostaglandin E stimulus / heterotrimeric G-protein complex / positive regulation of cold-induced thermogenesis / G alpha (12/13) signalling events / Inactivation, recovery and regulation of the phototransduction cascade / G-protein beta-subunit binding / extracellular vesicle / sensory perception of taste / Thrombin signalling through proteinase activated receptors (PARs) / signaling receptor complex adaptor activity / positive regulation of cytosolic calcium ion concentration / retina development in camera-type eye / GTPase binding / fibroblast proliferation Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||
 Authors Authors | Wang Y / Zhuang Y | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Molecular recognition of an acyl-peptide hormone and activation of ghrelin receptor. Authors: Yue Wang / Shimeng Guo / Youwen Zhuang / Ying Yun / Peiyu Xu / Xinheng He / Jia Guo / Wanchao Yin / H Eric Xu / Xin Xie / Yi Jiang /  Abstract: Ghrelin, also called "the hunger hormone", is a gastric peptide hormone that regulates food intake, body weight, as well as taste sensation, reward, cognition, learning and memory. One unique feature ...Ghrelin, also called "the hunger hormone", is a gastric peptide hormone that regulates food intake, body weight, as well as taste sensation, reward, cognition, learning and memory. One unique feature of ghrelin is its acylation, primarily with an octanoic acid, which is essential for its binding and activation of the ghrelin receptor, a G protein-coupled receptor. The multifaceted roles of ghrelin make ghrelin receptor a highly attractive drug target for growth retardation, obesity, and metabolic disorders. Here we present two cryo-electron microscopy structures of G-coupled ghrelin receptor bound to ghrelin and a synthetic agonist, GHRP-6. Analysis of these two structures reveals a unique binding pocket for the octanoyl group, which guides the correct positioning of the peptide to initiate the receptor activation. Together with mutational and functional data, our structures define the rules for recognition of the acylated peptide hormone and activation of ghrelin receptor, and provide structural templates to facilitate drug design targeting ghrelin receptor. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31500.map.gz emd_31500.map.gz | 27.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31500-v30.xml emd-31500-v30.xml emd-31500.xml emd-31500.xml | 16.2 KB 16.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_31500.png emd_31500.png | 54.9 KB | ||
| Filedesc metadata |  emd-31500.cif.gz emd-31500.cif.gz | 6.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31500 http://ftp.pdbj.org/pub/emdb/structures/EMD-31500 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31500 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31500 | HTTPS FTP |
-Related structure data
| Related structure data |  7f9yMC  7f9zC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_31500.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31500.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.071 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : ghrelin-bound ghrelin receptor in complex with Gq
| Entire | Name: ghrelin-bound ghrelin receptor in complex with Gq |
|---|---|
| Components |
|
-Supramolecule #1: ghrelin-bound ghrelin receptor in complex with Gq
| Supramolecule | Name: ghrelin-bound ghrelin receptor in complex with Gq / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Engineered G-alpha-q subunit
| Macromolecule | Name: Engineered G-alpha-q subunit / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 41.855578 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MMGCTLSAED KAAVERSKMI EKQLQKDKQV YRRTLRLLLL GADNSGKSTI VKQMRIYHVN GYSEEECKQY KAVVYSNTIQ SIIAIIRAM GRLKIDFGDS ARADDARQLF VLAGAAEEGF MTAELAGVIK RLWKDSGVQA CFNRSREYQL NDSAAYYLND L DRIAQPNY ...String: MMGCTLSAED KAAVERSKMI EKQLQKDKQV YRRTLRLLLL GADNSGKSTI VKQMRIYHVN GYSEEECKQY KAVVYSNTIQ SIIAIIRAM GRLKIDFGDS ARADDARQLF VLAGAAEEGF MTAELAGVIK RLWKDSGVQA CFNRSREYQL NDSAAYYLND L DRIAQPNY IPTQQDVLRT RVKTSGIFET KFQVDKVNFH MFDVGAQRDE RRKWIQCFND VTAIIFVVDS SDYNRLQEAL ND FKSIWNN RWLRTISVIL FLNKQDLLAE KVLAGKSKIE DYFPEFARYT TPEDATPEPG EDPRVTRAKY FIRKEFVDIS TAS GDGRHI CYPHFTCSVD TENARRIFND CKDIILQMNL REYNLV |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.226992 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSLLQSELD QLRQEAEQLK NQIRDARKAC ADATLSQITN NIDPVGRIQM RTRRTLRGHL AKIYAMHWGT DSRLLVSASQ DGKLIIWDS YTTNKVHAIP LRSSWVMTCA YAPSGNYVAC GGLDNICSIY NLKTREGNVR VSRELAGHTG YLSCCRFLDD N QIVTSSGD ...String: MGSLLQSELD QLRQEAEQLK NQIRDARKAC ADATLSQITN NIDPVGRIQM RTRRTLRGHL AKIYAMHWGT DSRLLVSASQ DGKLIIWDS YTTNKVHAIP LRSSWVMTCA YAPSGNYVAC GGLDNICSIY NLKTREGNVR VSRELAGHTG YLSCCRFLDD N QIVTSSGD TTCALWDIET GQQTTTFTGH TGDVMSLSLA PDTRLFVSGA CDASAKLWDV REGMCRQTFT GHESDINAIC FF PNGNAFA TGSDDATCRL FDLRADQELM TYSHDNIICG ITSVSFSKSG RLLLAGYDDF NCNVWDALKA DRAGVLAGHD NRV SCLGVT DDGMAVATGS WDSFLKIWNG SSGGGGSGGG GSSGVSGWRL FKKIS UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: Ghrelin-28
| Macromolecule | Name: Ghrelin-28 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 3.252726 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GSSFLSPEHQ RVQQRKESKK PPAKLQPR UniProtKB: Appetite-regulating hormone |
-Macromolecule #4: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #5: NB35
| Macromolecule | Name: NB35 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 15.343019 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAQVQLQESG GGLVQPGGSL RLSCAASGFT FSNYKMNWVR QAPGKGLEWV SDISQSGASI SYTGSVKGRF TISRDNAKNT LYLQMNSLK PEDTAVYYCA RCPAPFTRDC FDVTSTTYAY RGQGTQVTVS SHHHHHHEPE A |
-Macromolecule #6: Soluble cytochrome b562
| Macromolecule | Name: Soluble cytochrome b562 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 74.282609 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKTIIALSYI FCLVFAHHHH HHHHHHADLE DNWETLNDNL KVIEKADNAA QVKDALTKMR AAALDAQKAT PPKLEDKSPD SPEMKDFRH GFDILVGQID DALKLANEGK VKEAQAAAEQ LKTTRNAYIQ KYLMWNATPS EEPGFNLTLA DLDWDASPGN D SLGDELLQ ...String: MKTIIALSYI FCLVFAHHHH HHHHHHADLE DNWETLNDNL KVIEKADNAA QVKDALTKMR AAALDAQKAT PPKLEDKSPD SPEMKDFRH GFDILVGQID DALKLANEGK VKEAQAAAEQ LKTTRNAYIQ KYLMWNATPS EEPGFNLTLA DLDWDASPGN D SLGDELLQ LFPAPLLAGV TATCVALFVV GIAGNLLTML VVSRFRELRT TTNLYLSSMA FSDLLIFLCM PLDLVRLWQY RP WNFGDLL CKLFQFVSES CTYATVLTIT ALSVERYFAI CFPLRAKVVV TKGRVKLVIF VIWAVAFCSA GPIFVLVGVE HEN GTDPWD TNECRPTEFA VRSGLLTVMV WVSSIFFFLP VFCLTVLYSL IGRKLWRRRR GDAVVGASLR DQNHKQTVKM LAVV VFAFI LCWLPFHVGR YLFSKSFEPG SLEIAQISQY CNLVSFVLFY LSAAINPILY NIMSKKYRVA VFRLLGFEPF SQRKL STLK DESSRAWTES SINTGSAGSV FTLEDFVGDW EQTAAYNLDQ VLEQGGVSSL LQNLAVSVTP IQRIVRSGEN ALKIDI HVI IPYEGLSADQ MAQIEEVFKV VYPVDDHHFK VILPYGTLVI DGVTPNMLNY FGRPYEGIAV FDGKKITVTG TLWNGNK II DERLITPDGS MLFRVTINS |
-Macromolecule #7: OCTANOIC ACID (CAPRYLIC ACID)
| Macromolecule | Name: OCTANOIC ACID (CAPRYLIC ACID) / type: ligand / ID: 7 / Number of copies: 1 / Formula: OCA |
|---|---|
| Molecular weight | Theoretical: 144.211 Da |
| Chemical component information | 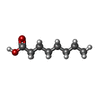 ChemComp-OCA: |
-Macromolecule #8: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 8 / Number of copies: 2 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 80.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.9 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 522055 |
| Initial angle assignment | Type: RANDOM ASSIGNMENT |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller






































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















