+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-31002 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human ABCA4 in ATP-bound state | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | lipid transport / MEMBRANE PROTEIN / TRANSLOCASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationrod photoreceptor disc membrane / flippase activity / N-retinylidene-phosphatidylethanolamine flippase activity / Defective visual phototransduction due to ABCA4 loss of function / all-trans retinal binding / phospholipid transfer to membrane / retinol transmembrane transporter activity / phospholipid transporter activity / 11-cis retinal binding / retinal metabolic process ...rod photoreceptor disc membrane / flippase activity / N-retinylidene-phosphatidylethanolamine flippase activity / Defective visual phototransduction due to ABCA4 loss of function / all-trans retinal binding / phospholipid transfer to membrane / retinol transmembrane transporter activity / phospholipid transporter activity / 11-cis retinal binding / retinal metabolic process / ATPase-coupled intramembrane lipid transporter activity / phosphatidylethanolamine flippase activity / retinoid binding / photoreceptor cell maintenance / P-type phospholipid transporter / phospholipid translocation / lipid transport / The canonical retinoid cycle in rods (twilight vision) / phototransduction, visible light / photoreceptor outer segment / ATPase-coupled transmembrane transporter activity / ABC-type transporter activity / retinoid metabolic process / visual perception / ABC-family proteins mediated transport / transmembrane transport / photoreceptor disc membrane / cytoplasmic vesicle / GTPase activity / intracellular membrane-bounded organelle / endoplasmic reticulum / ATP hydrolysis activity / ATP binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Xie T / Zhang ZK / Gong X | |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Structural basis of substrate recognition and translocation by human ABCA4. Authors: Tian Xie / Zike Zhang / Qi Fang / Bowen Du / Xin Gong /  Abstract: Human ATP-binding cassette (ABC) subfamily A (ABCA) transporters mediate the transport of various lipid compounds across the membrane. Mutations in human ABCA transporters have been described to ...Human ATP-binding cassette (ABC) subfamily A (ABCA) transporters mediate the transport of various lipid compounds across the membrane. Mutations in human ABCA transporters have been described to cause severe hereditary disorders associated with impaired lipid transport. However, little is known about the mechanistic details of substrate recognition and translocation by ABCA transporters. Here, we present three cryo-EM structures of human ABCA4, a retina-specific ABCA transporter, in distinct functional states at resolutions of 3.3-3.4 Å. In the nucleotide-free state, the two transmembrane domains (TMDs) exhibit a lateral-opening conformation, allowing the lateral entry of substrate from the lipid bilayer. The N-retinylidene-phosphatidylethanolamine (NRPE), the physiological lipid substrate of ABCA4, is sandwiched between the two TMDs in the luminal leaflet and is further stabilized by an extended loop from extracellular domain 1. In the ATP-bound state, the two TMDs display a closed conformation, which precludes the substrate binding. Our study provides a molecular basis to understand the mechanism of ABCA4-mediated NRPE recognition and translocation, and suggests a common 'lateral access and extrusion' mechanism for ABCA-mediated lipid transport. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31002.map.gz emd_31002.map.gz | 78.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31002-v30.xml emd-31002-v30.xml emd-31002.xml emd-31002.xml | 11.7 KB 11.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_31002.png emd_31002.png | 34 KB | ||
| Filedesc metadata |  emd-31002.cif.gz emd-31002.cif.gz | 6.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31002 http://ftp.pdbj.org/pub/emdb/structures/EMD-31002 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31002 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31002 | HTTPS FTP |
-Related structure data
| Related structure data |  7e7qMC  7e7iC  7e7oC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_31002.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31002.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
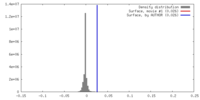
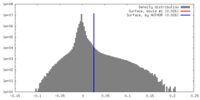
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : ATP-bound ABCA4
| Entire | Name: ATP-bound ABCA4 |
|---|---|
| Components |
|
-Supramolecule #1: ATP-bound ABCA4
| Supramolecule | Name: ATP-bound ABCA4 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Retinal-specific phospholipid-transporting ATPase ABCA4
| Macromolecule | Name: Retinal-specific phospholipid-transporting ATPase ABCA4 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: P-type phospholipid transporter |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 261.141578 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MADYKDDDDK SGPDEVDASG RMGFVRQIQL LLWKNWTLRK RQKIRFVVEL VWPLSLFLVL IWLRNANPLY SHHECHFPNK AMPSAGMLP WLQGIFCNVN NPCFQSPTPG ESPGIVSNYN NSILARVYRD FQELLMNAPE SQHLGRIWTE LHILSQFMDT L RTHPERIA ...String: MADYKDDDDK SGPDEVDASG RMGFVRQIQL LLWKNWTLRK RQKIRFVVEL VWPLSLFLVL IWLRNANPLY SHHECHFPNK AMPSAGMLP WLQGIFCNVN NPCFQSPTPG ESPGIVSNYN NSILARVYRD FQELLMNAPE SQHLGRIWTE LHILSQFMDT L RTHPERIA GRGIRIRDIL KDEETLTLFL IKNIGLSDSV VYLLINSQVR PEQFAHGVPD LALKDIACSE ALLERFIIFS QR RGAKTVR YALCSLSQGT LQWIEDTLYA NVDFFKLFRV LPTLLDSRSQ GINLRSWGGI LSDMSPRIQE FIHRPSMQDL LWV TRPLMQ NGGPETFTKL MGILSDLLCG YPEGGGSRVL SFNWYEDNNY KAFLGIDSTR KDPIYSYDRR TTSFCNALIQ SLES NPLTK IAWRAAKPLL MGKILYTPDS PAARRILKNA NSTFEELEHV RKLVKAWEEV GPQIWYFFDN STQMNMIRDT LGNPT VKDF LNRQLGEEGI TAEAILNFLY KGPRESQADD MANFDWRDIF NITDRTLRLV NQYLECLVLD KFESYNDETQ LTQRAL SLL EENMFWAGVV FPDMYPWTSS LPPHVKYKIR MDIDVVEKTN KIKDRYWDSG PRADPVEDFR YIWGGFAYLQ DMVEQGI TR SQVQAEAPVG IYLQQMPYPC FVDDSFMIIL NRCFPIFMVL AWIYSVSMTV KSIVLEKELR LKETLKNQGV SNAVIWCT W FLDSFSIMSM SIFLLTIFIM HGRILHYSDP FILFLFLLAF STATIMLCFL LSTFFSKASL AAACSGVIYF TLYLPHILC FAWQDRMTAE LKKAVSLLSP VAFGFGTEYL VRFEEQGLGL QWSNIGNSPT EGDEFSFLLS MQMMLLDAAV YGLLAWYLDQ VFPGDYGTP LPWYFLLQES YWLGGEGCST REERALEKTE PLTEETEDPE HPEGIHDSFF EREHPGWVPG VCVKNLVKIF E PCGRPAVD RLNITFYENQ ITAFLGHNGA GKTTTLSILT GLLPPTSGTV LVGGRDIETS LDAVRQSLGM CPQHNILFHH LT VAEHMLF YAQLKGKSQE EAQLEMEAML EDTGLHHKRN EEAQDLSGGM QRKLSVAIAF VGDAKVVILD QPTSGVDPYS RRS IWDLLL KYRSGRTIIM STHHMDEADL LGDRIAIIAQ GRLYCSGTPL FLKNCFGTGL YLTLVRKMKN IQSQRKGSEG TCSC SSKGF STTCPAHVDD LTPEQVLDGD VNELMDVVLH HVPEAKLVEC IGQELIFLLP NKNFKHRAYA SLFRELEETL ADLGL SSFG ISDTPLEEIF LKVTEDSDSG PLFAGGAQQK RENVNPRHPC LGPREKAGQT PQDSNVCSPG APAAHPEGQP PPEPEC PGP QLNTGTQLVL QHVQALLVKR FQHTIRSHKD FLAQIVLPAT FVFLALMLSI VIPPFGEYPA LTLHPWIYGQ QYTFFSM DE PGSEQFTVLA DVLLNKPGFG NRCLKEGWLP EYPCGNSTPW KTPSVSPNIT QLFQKQKWTQ VNPSPSCRCS TREKLTML P ECPEGAGGLP PPQRTQRSTE ILQDLTDRNI SDFLVKTYPA LIRSSLKSKF WVNEQRYGGI SIGGKLPVVP ITGEALVGF LSDLGRIMNV SGGPITREAS KEIPDFLKHL ETEDNIKVWF NNKGWHALVS FLNVAHNAIL RASLPKDRSP EEYGITVISQ PLNLTKEQL SEITVLTTSV DAVVAICVIF SMSFVPASFV LYLIQERVNK SKHLQFISGV SPTTYWVTNF LWDIMNYSVS A GLVVGIFI GFQKKAYTSP ENLPALVALL LLYGWAVIPM MYPASFLFDV PSTAYVALSC ANLFIGINSS AITFILELFE NN RTLLRFN AVLRKLLIVF PHFCLGRGLI DLALSQAVTD VYARFGEEHS ANPFHWDLIG KNLFAMVVEG VVYFLLTLLV QRH FFLSQW IAEPTKEPIV DEDDDVAEER QRIITGGNKT DILRLHELTK IYPGTSSPAV DRLCVGVRPG ECFGLLGVNG AGKT TTFKM LTGDTTVTSG DATVAGKSIL TNISEVHQNM GYCPQFDAID ELLTGREHLY LYARLRGVPA EEIEKVANWS IKSLG LTVY ADCLAGTYSG GNKRKLSTAI ALIGCPPLVL LDQPTTGMDP QARRMLWNVI VSIIREGRAV VLTSHSMEEC EALCTR LAI MVKGAFRCMG TIQHLKSKFG DGYIVTMKIK SPKDDLLPDL NPVEQFFQGN FPGSVQRERH YNMLQFQVSS SSLARIF QL LLSHKDSLLI EEYSVTQTTL DQVFVNFAKQ QTESHDLPLH PRAAGASRQA QDLEGSDEVD AVEGSHHHHH HHHHH UniProtKB: Retinal-specific phospholipid-transporting ATPase ABCA4 |
-Macromolecule #3: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 3 / Number of copies: 7 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #4: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 4 / Number of copies: 2 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Macromolecule #5: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 5 / Number of copies: 2 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.3 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 173278 |
| Initial angle assignment | Type: NOT APPLICABLE |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)