+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-30343 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Structure of RyR1 (Ca2+/CHL) | |||||||||||||||
 マップデータ マップデータ | rRyR1 Ca2 /CHL | |||||||||||||||
 試料 試料 |
| |||||||||||||||
 キーワード キーワード | Rabbit / Ryanodine receptor1 / CHL. / MEMBRANE PROTEIN | |||||||||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報ATP-gated ion channel activity / : / negative regulation of calcium-mediated signaling / negative regulation of insulin secretion involved in cellular response to glucose stimulus / neuronal action potential propagation / negative regulation of release of sequestered calcium ion into cytosol / insulin secretion involved in cellular response to glucose stimulus / terminal cisterna / ryanodine-sensitive calcium-release channel activity / ryanodine receptor complex ...ATP-gated ion channel activity / : / negative regulation of calcium-mediated signaling / negative regulation of insulin secretion involved in cellular response to glucose stimulus / neuronal action potential propagation / negative regulation of release of sequestered calcium ion into cytosol / insulin secretion involved in cellular response to glucose stimulus / terminal cisterna / ryanodine-sensitive calcium-release channel activity / ryanodine receptor complex / release of sequestered calcium ion into cytosol by sarcoplasmic reticulum / response to redox state / ossification involved in bone maturation / negative regulation of heart rate / cellular response to caffeine / 'de novo' protein folding / skin development / FK506 binding / organelle membrane / intracellularly gated calcium channel activity / smooth endoplasmic reticulum / outflow tract morphogenesis / smooth muscle contraction / toxic substance binding / T cell proliferation / striated muscle contraction / regulation of cardiac muscle contraction by regulation of the release of sequestered calcium ion / voltage-gated calcium channel activity / calcium channel inhibitor activity / skeletal muscle fiber development / regulation of release of sequestered calcium ion into cytosol by sarcoplasmic reticulum / Ion homeostasis / release of sequestered calcium ion into cytosol / calcium channel complex / sarcoplasmic reticulum membrane / cellular response to calcium ion / muscle contraction / sarcoplasmic reticulum / protein maturation / peptidylprolyl isomerase / calcium channel regulator activity / peptidyl-prolyl cis-trans isomerase activity / calcium-mediated signaling / sarcolemma / calcium ion transmembrane transport / Stimuli-sensing channels / calcium channel activity / Z disc / intracellular calcium ion homeostasis / disordered domain specific binding / positive regulation of cytosolic calcium ion concentration / protein refolding / protein homotetramerization / transmembrane transporter binding / calmodulin binding / signaling receptor binding / calcium ion binding / ATP binding / identical protein binding / membrane / cytoplasm 類似検索 - 分子機能 | |||||||||||||||
| 生物種 |   Homo sapiens (ヒト) Homo sapiens (ヒト) | |||||||||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 4.7 Å | |||||||||||||||
 データ登録者 データ登録者 | Ma R / Haji-Ghassemi O | |||||||||||||||
| 資金援助 |  中国, 中国,  カナダ, 4件 カナダ, 4件
| |||||||||||||||
 引用 引用 |  ジャーナル: Nat Chem Biol / 年: 2020 ジャーナル: Nat Chem Biol / 年: 2020タイトル: Structural basis for diamide modulation of ryanodine receptor. 著者: Ruifang Ma / Omid Haji-Ghassemi / Dan Ma / Heng Jiang / Lianyun Lin / Li Yao / Arthur Samurkas / Yuxin Li / Yiwen Wang / Peng Cao / Shian Wu / Yan Zhang / Takashi Murayama / Bernard Moussian ...著者: Ruifang Ma / Omid Haji-Ghassemi / Dan Ma / Heng Jiang / Lianyun Lin / Li Yao / Arthur Samurkas / Yuxin Li / Yiwen Wang / Peng Cao / Shian Wu / Yan Zhang / Takashi Murayama / Bernard Moussian / Filip Van Petegem / Zhiguang Yuchi /      要旨: The diamide insecticide class is one of the top-selling insecticides globally. They are used to control a wide range of pests by targeting their ryanodine receptors (RyRs). Here, we report the ...The diamide insecticide class is one of the top-selling insecticides globally. They are used to control a wide range of pests by targeting their ryanodine receptors (RyRs). Here, we report the highest-resolution cryo-electron microscopy (cryo-EM) structure of RyR1 in the open state, in complex with the anthranilic diamide chlorantraniliprole (CHL). The 3.2-Å local resolution map facilitates unambiguous assignment of the CHL binding site. The molecule induces a conformational change by affecting the S4-S5 linker, triggering channel opening. The binding site is further corroborated by mutagenesis data, which reveal how diamide insecticides are selective to the Lepidoptera group of insects over honeybee or mammalian RyRs. Our data reveal that several pests have developed resistance via two mechanisms, steric hindrance and loss of contact. Our results provide a foundation for the development of highly selective pesticides aimed at overcoming resistance and therapeutic molecules to treat human myopathies. | |||||||||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_30343.map.gz emd_30343.map.gz | 278.6 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-30343-v30.xml emd-30343-v30.xml emd-30343.xml emd-30343.xml | 28.9 KB 28.9 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_30343_fsc.xml emd_30343_fsc.xml | 16 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_30343.png emd_30343.png | 142.2 KB | ||
| Filedesc metadata |  emd-30343.cif.gz emd-30343.cif.gz | 8.7 KB | ||
| その他 |  emd_30343_additional.map.gz emd_30343_additional.map.gz emd_30343_additional_1.map.gz emd_30343_additional_1.map.gz emd_30343_half_map_1.map.gz emd_30343_half_map_1.map.gz emd_30343_half_map_2.map.gz emd_30343_half_map_2.map.gz | 85.9 MB 85.9 MB 254.5 MB 257 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30343 http://ftp.pdbj.org/pub/emdb/structures/EMD-30343 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30343 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30343 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_30343.map.gz / 形式: CCP4 / 大きさ: 343 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_30343.map.gz / 形式: CCP4 / 大きさ: 343 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | rRyR1 Ca2 /CHL | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
-追加マップ: rRyR1 Ca2 /CHL
| ファイル | emd_30343_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | rRyR1 Ca2 /CHL | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-追加マップ: rRyR1 Ca2 /CHL
| ファイル | emd_30343_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | rRyR1 Ca2 /CHL | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: rRyR1 Ca2 /CHL, half map
| ファイル | emd_30343_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | rRyR1 Ca2 /CHL, half map | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: rRyR1 Ca2 /CHL, half map
| ファイル | emd_30343_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | rRyR1 Ca2 /CHL, half map | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : RyR1 Ca2+/CHL
| 全体 | 名称: RyR1 Ca2+/CHL |
|---|---|
| 要素 |
|
-超分子 #1: RyR1 Ca2+/CHL
| 超分子 | 名称: RyR1 Ca2+/CHL / タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: #1-#2 |
|---|---|
| 分子量 | 理論値: 2.54 MDa |
-超分子 #2: RyR1
| 超分子 | 名称: RyR1 / タイプ: complex / ID: 2 / 親要素: 1 / 含まれる分子: #1 |
|---|---|
| 由来(天然) | 生物種:  |
-超分子 #3: FKBP1B
| 超分子 | 名称: FKBP1B / タイプ: complex / ID: 3 / 親要素: 1 / 含まれる分子: #2 |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
-分子 #1: Ryanodine receptor 1,RyR1
| 分子 | 名称: Ryanodine receptor 1,RyR1 / タイプ: protein_or_peptide / ID: 1 / コピー数: 4 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  |
| 分子量 | 理論値: 539.836938 KDa |
| 配列 | 文字列: MGDGGEGEDE VQFLRTDDEV VLQCSATVLK EQLKLCLAAE GFGNRLCFLE PTSNAQNVPP DLAICCFTLE QSLSVRALQE MLANTVEAG VESSQGGGHR TLLYGHAILL RHAHSRMYLS CLTTSRSMTD KLAFDVGLQE DATGEACWWT MHPASKQRSE G EKVRVGDD ...文字列: MGDGGEGEDE VQFLRTDDEV VLQCSATVLK EQLKLCLAAE GFGNRLCFLE PTSNAQNVPP DLAICCFTLE QSLSVRALQE MLANTVEAG VESSQGGGHR TLLYGHAILL RHAHSRMYLS CLTTSRSMTD KLAFDVGLQE DATGEACWWT MHPASKQRSE G EKVRVGDD LILVSVSSER YLHLSTASGE LQVDASFMQT LWNMNPICSC CEEGYVTGGH VLRLFHGHMD ECLTISAADS DD QRRLVYY EGGAVCTHAR SLWRLEPLRI SWSGSHLRWG QPLRIRHVTT GRYLALTEDQ GLVVVDACKA HTKATSFCFR VSK EKLDTA PKRDVEGMGP PEIKYGESLC FVQHVASGLW LTYAAPDPKA LRLGVLKKKA ILHQEGHMDD ALFLTRCQQE ESQA ARMIH STAGLYNQFI KGLDSFSGKP RGSGPPAGPA LPIEAVILSL QDLIGYFEPP SEELQHEEKQ SKLRSLRNRQ SLFQE EGML SLVLNCIDRL NVYTTAAHFA EYAGEEAAES WKEIVNLLYE LLASLIRGNR ANCALFSTNL DWVVSKLDRL EASSGI LEV LYCVLIESPE VLNIIQENHI KSIISLLDKH GRNHKVLDVL CSLCVCNGVA VRSNQDLITE NLLPGRELLL QTNLINY VT SIRPNIFVGR AEGSTQYGKW YFEVMVDEVV PFLTAQATHL RVGWALTEGY SPYPGGGEGW GGNGVGDDLY SYGFDGLH L WTGHVARPVT SPGQHLLAPE DVVSCCLDLS VPSISFRING CPVQGVFEAF NLDGLFFPVV SFSAGVKVRF LLGGRHGEF KFLPPPGYAP CHEAVLPRER LRLEPIKEYR REGPRGPHLV GPSRCLSHTD FVPCPVDTVQ IVLPPHLERI REKLAENIHE LWALTRIEQ GWTYGPVRDD NKRLHPCLVN FHSLPEPERN YNLQMSGETL KTLLALGCHV GMADEKAEDN LKKTKLPKTY M MSNGYKPA PLDLSHVRLT PAQTTLVDRL AENGHNVWAR DRVAQGWSYS AVQDIPARRN PRLVPYRLLD EATKRSNRDS LC QAVRTLL GYGYNIEPPD QEPSQVENQS RWDRVRIFRA EKSYTVQSGR WYFEFEAVTT GEMRVGWARP ELRPDVELGA DEL AYVFNG HRGQRWHLGS EPFGRPWQSG DVVGCMIDLT ENTIIFTLNG EVLMSDSGSE TAFREIEIGD GFLPVCSLGP GQVG HLNLG QDVSSLRFFA ICGLQEGFEP FAINMQRPVT TWFSKSLPQF EPVPPEHPHY EVARMDGTVD TPPCLRLAHR TWGSQ NSLV EMLFLRLSLP VQFHQHFRCT AGATPLAPPG LQPPAEDEAR AAEPDPDYEN LRRSAGGWGE AEGGKEGTAK EGTPGG TPQ PGVEAQPVRA ENEKDATTEK NKKRGFLFKA KKAAMMTQPP ATPALPRLPH DVVPADNRDD PEIILNTTTY YYSVRVF AG QEPSCVWVGW VTPDYHQHDM NFDLSKVRAV TVTMGDEQGN VHSSLKCSNC YMVWGGDFVS PGQQGRISHT DLVIGCLV D LATGLMTFTA NGKESNTFFQ VEPNTKLFPA VFVLPTHQNV IQFELGKQKN IMPLSAAMFL SERKNPAPQC PPRLEVQML MPVSWSRMPN HFLQVETRRA GERLGWAVQC QDPLTMMALH IPEENRCMDI LELSERLDLQ RFHSHTLRLY RAVCALGNNR VAHALCSHV DQAQLLHALE DAHLPGPLRA GYYDLLISIH LESACRSRRS MLSEYIVPLT PETRAITLFP PGRKGGNARR H GLPGVGVT TSLRPPHHFS PPCFVAALPA AGVAEAPARL SPAIPLEALR DKALRMLGEA VRDGGQHARD PVGGSVEFQF VP VLKLVST LLVMGIFGDE DVKQILKMIE PEVFTEEEEE EEEEEEEEEE EEEDEEEKEE DEEEEEKEDA EKEEEEAPEG EKE DLEEGL LQMKLPESVK LQMCNLLEYF CDQELQHRVE SLAAFAERYV DKLQANQRSR YALLMRAFTM SAAETARRTR EFRS PPQEQ INMLLHFKDE ADEEDCPLPE DIRQDLQDFH QDLLAHCGIQ LEGEEEEPEE ETSLSSRLRS LLETVRLVKK KEEKP EEEL PAEEKKPQSL QELVSHMVVR WAQEDYVQSP ELVRAMFSLL HRQYDGLGEL LRALPRAYTI SPSSVEDTMS LLECLG QIR SLLIVQMGPQ EENLMIQSIG NIMNNKVFYQ HPNLMRALGM HETVMEVMVN VLGGGETKEI RFPKMVTSCC RFLCYFC RI SRQNQRSMFD HLSYLLENSG IGLGMQGSTP LDVAAASVID NNELALALQE QDLEKVVSYL AGCGLQSCPM LLAKGYPD I GWNPCGGERY LDFLRFAVFV NGESVEENAN VVVRLLIRKP ECFGPALRGE GGSGLLAAIE EAIRISEDPA RDGPGVRRD RRREHFGEEP PEENRVHLGH AIMSFYAALI DLLGRCAPEM HLIQAGKGEA LRIRAILRSL VPLDDLVGII SLPLQIPTLG KDGALVQPK MSASFVPDHK ASMVLFLDRV YGIENQDFLL HVLDVGFLPD MRAAASLDTA TFSTTEMALA LNRYLCLAVL P LITKCAPL FAGTEHRAIM VDSMLHTVYR LSRGRSLTKA QRDVIEDCLM ALCRYIRPSM LQHLLRRLVF DVPILN(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)E TLNVIIPEKL DSFINKFAEY THEKWAFDKI QNNW SYGEN VDEELKTHPM LRPYKTFSEK DKEIYRWPIK ESLKAMIAWE WTIEKAREGE EERTEKKKTR KISQTAQTYD PREGY NPQP PDLSGVTLSR ELQAMAEQLA ENYHNTWGRK KKQELEAKGG GTHPLLVPYD TLTAKEKARD REKAQELLKF LQMNGY AVT R(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)A(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)LLSK QRRRAVVACF RMTPLY NLP THRACNMFLE SYKAAWILTE DHSFEDRMID DLSKAGEQEE EEEEVEEKKP DPLHQLVLHF SRTALTEKSK LDEDYLY MA YADIMAKSCH LEEGGENGEA EEEEVEVSFE EKEMEKQRLL YQQSRLHTRG AAEMVLQMIS ACKGETGAMV SSTLKLGI S ILNGGNAEVQ QKMLDYLKDK KEVGFFQSIQ ALMQTCSVLD LNAFERQNKA EGLGMVNEDG TVINRQNGEK VMADDEFTQ DLFRFLQLLC EGHNNDFQNY LRTQTGNTTT INIIICTVDY LLRLQESISD FYWYYSGKDV IEEQGKRNFS KAMSVAKQVF NSLTEYIQG PCTGNQQSLA HSRLWDAVVG FLHVFAHMMM KLAQDSSQIE LLKELLDLQK DMVVMLLSLL EGNVVNGMIA R QMVDMLVE SSSNVEMILK FFDMFLKLKD IVGSEAFQDY VTDPRGLISK KDFQKAMDSQ KQFTGPEIQF LLSCSEADEN EM INFEEFA NRFQEPARDI GFNVAVLLTN LSEHVPHDPR LRNFLELAES ILEYFRPYLG RIEIMGASRR IERIYFEISE TNR AQWEMP QVKESKRQFI FDVVNEGGEA EKMELFVSFC EDTIFEMQIA AQISE(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) WGELEVQRVK FLNYLSRNFY TLRFLALFLA FAINFILLF YKVSDSPPGE DDMEGSAAGD LAGAGSGGGS GWGSGAGEEA EGDEDENMVY YFLEESTGYM EPALWCLSLL H TLVAFLCI IGYNCLKVPL VIFKREKELA RKLEFDGLYI TEQPGDDDVK GQWDRLVLNT PSFPSNYWDK FVKRKVLDKH GD IFGRERI AELLGMDLAS LEITAHNERK PDPPPGLLTW LMSIDVKYQI WKFGVIFTDN SFLYLGWYMV MSLLGHYNNF FFA AHLLDI AMGVKTLRTI LSSVTHNGKQ LVMTVGLLAV VVYLYTVVAF NFFRKFYNKS EDEDEPDMKC DDMMTCYLFH MYVG VRAGG GIGDEIEDPA GDEYELYRVV FDITFFFFVI VILLAIIQGL IIDAFGELRD QQEQVKEDME TKCFICGIGS DYFDT TPHG FETHTLEEHN LANYMFFLMY LINKDETEHT GQESYVWKMY QERCWDFFPA GDCFRKQYED QLS UniProtKB: Ryanodine receptor 1, Ryanodine receptor 1, Ryanodine receptor 1, Ryanodine receptor 1 |
-分子 #2: Peptidyl-prolyl cis-trans isomerase FKBP1B
| 分子 | 名称: Peptidyl-prolyl cis-trans isomerase FKBP1B / タイプ: protein_or_peptide / ID: 2 / コピー数: 4 / 光学異性体: LEVO / EC番号: peptidylprolyl isomerase |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 11.667305 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: GVEIETISPG DGRTFPKKGQ TCVVHYTGML QNGKKFDSSR DRNKPFKFRI GKQEVIKGFE EGAAQMSLGQ RAKLTCTPDV AYGATGHPG VIPPNATLIF DVELLNLE UniProtKB: Peptidyl-prolyl cis-trans isomerase FKBP1B |
-分子 #3: CALCIUM ION
| 分子 | 名称: CALCIUM ION / タイプ: ligand / ID: 3 / コピー数: 4 / 式: CA |
|---|---|
| 分子量 | 理論値: 40.078 Da |
-分子 #4: ZINC ION
| 分子 | 名称: ZINC ION / タイプ: ligand / ID: 4 / コピー数: 4 / 式: ZN |
|---|---|
| 分子量 | 理論値: 65.409 Da |
-分子 #5: 5-bromanyl-N-[4-chloranyl-2-methyl-6-(methylcarbamoyl)phenyl]-2-(...
| 分子 | 名称: 5-bromanyl-N-[4-chloranyl-2-methyl-6-(methylcarbamoyl)phenyl]-2-(3-chloranylpyridin-2-yl)pyrazole-3-carboxamide タイプ: ligand / ID: 5 / コピー数: 4 / 式: F0U |
|---|---|
| 分子量 | 理論値: 483.146 Da |
| Chemical component information | 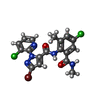 ChemComp-F0U: |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.5 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | TFS KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: FEI FALCON III (4k x 4k) 平均電子線量: 50.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)






















































