+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2b57 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Guanine Riboswitch C74U mutant bound to 2,6-diaminopurine | ||||||
 Components Components | 65-MER | ||||||
 Keywords Keywords | RNA / RNA-ligand complex / Double helix / base triples / base quadruples / mRNA / purine | ||||||
| Function / homology | 9H-PURINE-2,6-DIAMINE / ACETATE ION / COBALT HEXAMMINE(III) / : / RNA / RNA (> 10) Function and homology information Function and homology information | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.15 Å MOLECULAR REPLACEMENT / Resolution: 2.15 Å | ||||||
 Authors Authors | Gilbert, S.D. / Stoddard, C.D. / Wise, S.J. / Batey, R.T. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2006 Journal: J.Mol.Biol. / Year: 2006Title: Thermodynamic and kinetic characterization of ligand binding to the purine riboswitch aptamer domain. Authors: Gilbert, S.D. / Stoddard, C.D. / Wise, S.J. / Batey, R.T. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2b57.cif.gz 2b57.cif.gz | 52.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2b57.ent.gz pdb2b57.ent.gz | 37.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2b57.json.gz 2b57.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/b5/2b57 https://data.pdbj.org/pub/pdb/validation_reports/b5/2b57 ftp://data.pdbj.org/pub/pdb/validation_reports/b5/2b57 ftp://data.pdbj.org/pub/pdb/validation_reports/b5/2b57 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 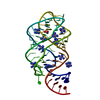
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: RNA chain | Mass: 20857.381 Da / Num. of mol.: 1 / Fragment: G-box RNA / Mutation: C74U / Source method: obtained synthetically Details: RNA was prepared by in vitro transcription. The sequence of this RNA can be found naturally in Bacillus Subtilis References: GenBank: 1256615 | ||||
|---|---|---|---|---|---|
| #2: Chemical | ChemComp-ACT / | ||||
| #3: Chemical | ChemComp-NCO / #4: Chemical | ChemComp-6AP / | #5: Water | ChemComp-HOH / | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.29 Å3/Da / Density % sol: 46 % | ||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 7.5 Details: 20% PEG 2K, 480 mM Ammonium Acetate, 12 mM Cobalt Hexammine, pH 7.5, VAPOR DIFFUSION, HANGING DROP, temperature 298K | ||||||||||||||||||||||||||||||||
| Components of the solutions |
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU / Wavelength: 1.5418 Å ROTATING ANODE / Type: RIGAKU / Wavelength: 1.5418 Å |
| Detector | Type: RIGAKU RAXIS IV / Detector: IMAGE PLATE / Date: Mar 18, 2005 |
| Radiation | Monochromator: NI Filter / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.15→20 Å / Num. all: 10478 / Num. obs: 10387 / % possible obs: 99.1 % / Observed criterion σ(F): 2 / Observed criterion σ(I): 2 / Biso Wilson estimate: 17.7 Å2 |
| Reflection shell | Resolution: 2.15→2.23 Å / % possible all: 90.71 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 2.15→19.86 Å / Rfactor Rfree error: 0.01 / Data cutoff high absF: 423665.58 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 MOLECULAR REPLACEMENT / Resolution: 2.15→19.86 Å / Rfactor Rfree error: 0.01 / Data cutoff high absF: 423665.58 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 52.7638 Å2 / ksol: 0.352404 e/Å3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 40.5 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.15→19.86 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.15→2.28 Å / Rfactor Rfree error: 0.032 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller








 PDBj
PDBj