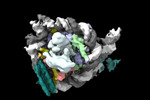
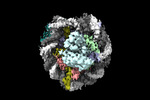
登録情報 データベース : EMDB / ID : EMD-26258タイトル structure of LIN28b nucleosome bound 3 OCT4 複合体 : Complex of nucleosome bound to three OCT-4sタンパク質・ペプチド : Histone H3.1タンパク質・ペプチド : Histone H4タンパク質・ペプチド : Histone H2A type 2-Cタンパク質・ペプチド : Histone H2B type 2-EDNA : DNA (162-MER)DNA : DNA (162-MER)タンパク質・ペプチド : Maltodextrin-binding protein,POU domain, class 5, transcription factor 1タンパク質・ペプチド : Single-chain variable fragment / / / / 機能・相同性 分子機能 ドメイン・相同性 構成要素
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / 生物種 Homo sapiens (ヒト) / Mus musculus (ハツカネズミ)手法 / / 解像度 : 2.6 Å Lian T / Guan R / Bai Y 資金援助 Organization Grant number 国 National Institutes of Health/National Cancer Institute (NIH/NCI)
ジャーナル : Mol Cell / 年 : 2023タイトル : Structural mechanism of LIN28B nucleosome targeting by OCT4.著者 : Ruifang Guan / Tengfei Lian / Bing-Rui Zhou / David Wheeler / Yawen Bai / 要旨 : Pioneer transcription factors are essential for cell fate changes by targeting closed chromatin. OCT4 is a crucial pioneer factor that can induce cell reprogramming. However, the structural basis of ... Pioneer transcription factors are essential for cell fate changes by targeting closed chromatin. OCT4 is a crucial pioneer factor that can induce cell reprogramming. However, the structural basis of how pioneer factors recognize the in vivo nucleosomal DNA targets is unknown. Here, we determine the high-resolution structures of the nucleosome containing human LIN28B DNA and its complexes with the OCT4 DNA binding region. Three OCT4s bind the pre-positioned nucleosome by recognizing non-canonical DNA sequences. Two use their POUS domains while the other uses the POUS-loop-POUHD region; POUHD serves as a wedge to unwrap ∼25 base pair DNA. Our analysis of previous genomic data and determination of the ESRRB-nucleosome-OCT4 structure confirmed the generality of these structural features. Moreover, biochemical studies suggest that multiple OCT4s cooperatively open the H1-condensed nucleosome array containing the LIN28B nucleosome. Thus, our study suggests a mechanism of how OCT4 can target the nucleosome and open closed chromatin. 履歴 登録 2022年2月18日 - ヘッダ(付随情報) 公開 2023年6月28日 - マップ公開 2023年6月28日 - 更新 2023年6月28日 - 現状 2023年6月28日 処理サイト : RCSB / 状態 : 公開
すべて表示 表示を減らす
 データを開く
データを開く 基本情報
基本情報
 マップデータ
マップデータ 試料
試料 キーワード
キーワード 機能・相同性情報
機能・相同性情報 Homo sapiens (ヒト) /
Homo sapiens (ヒト) / 
 データ登録者
データ登録者 米国, 1件
米国, 1件  引用
引用 ジャーナル: Mol Cell / 年: 2023
ジャーナル: Mol Cell / 年: 2023
 構造の表示
構造の表示 ダウンロードとリンク
ダウンロードとリンク emd_26258.map.gz
emd_26258.map.gz EMDBマップデータ形式
EMDBマップデータ形式 emd-26258-v30.xml
emd-26258-v30.xml emd-26258.xml
emd-26258.xml EMDBヘッダ
EMDBヘッダ emd_26258.png
emd_26258.png http://ftp.pdbj.org/pub/emdb/structures/EMD-26258
http://ftp.pdbj.org/pub/emdb/structures/EMD-26258 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26258
ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26258 emd_26258_validation.pdf.gz
emd_26258_validation.pdf.gz EMDB検証レポート
EMDB検証レポート emd_26258_full_validation.pdf.gz
emd_26258_full_validation.pdf.gz emd_26258_validation.xml.gz
emd_26258_validation.xml.gz emd_26258_validation.cif.gz
emd_26258_validation.cif.gz https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26258
https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26258 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26258
ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26258 リンク
リンク EMDB (EBI/PDBe) /
EMDB (EBI/PDBe) /  EMDataResource
EMDataResource マップ
マップ ダウンロード / ファイル: emd_26258.map.gz / 形式: CCP4 / 大きさ: 52.7 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
ダウンロード / ファイル: emd_26258.map.gz / 形式: CCP4 / 大きさ: 52.7 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) 試料の構成要素
試料の構成要素 Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト)
 Homo sapiens (ヒト)
Homo sapiens (ヒト)
 Homo sapiens (ヒト)
Homo sapiens (ヒト)
 Homo sapiens (ヒト)
Homo sapiens (ヒト)
 Homo sapiens (ヒト)
Homo sapiens (ヒト)


 Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト) 解析
解析 試料調製
試料調製 電子顕微鏡法
電子顕微鏡法 FIELD EMISSION GUN
FIELD EMISSION GUN
 画像解析
画像解析 ムービー
ムービー コントローラー
コントローラー