+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-24718 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Envelope-associated Adeno-associated virus serotype 2 | |||||||||
 Map data Map data | High resolution envelope-associated adeno-associated virus serotype 2. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | exo-AAV / exosome-associated adeno-associated virus / EA-AAV / Envelope-associated adeno-associated virus / VIRUS | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont entry into host cell via permeabilization of host membrane / host cell nucleolus / T=1 icosahedral viral capsid / clathrin-dependent endocytosis of virus by host cell / virion attachment to host cell / structural molecule activity Similarity search - Function | |||||||||
| Biological species |  Adeno-associated dependoparvovirus A / Adeno-associated dependoparvovirus A /  Adeno-associated virus - 2 Adeno-associated virus - 2 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.14 Å | |||||||||
 Authors Authors | Hull JA / Mietzsch M | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Virology / Year: 2022 Journal: Virology / Year: 2022Title: Structural characterization of an envelope-associated adeno-associated virus type 2 capsid. Authors: Joshua A Hull / Mario Mietzsch / Paul Chipman / David Strugatsky / Robert McKenna /  Abstract: Adeno-associated virus (AAV) are classified as non-enveloped ssDNA viruses. However, AAV capsids embedded within exosomes have been observed, and it has been suggested that the AAV membrane ...Adeno-associated virus (AAV) are classified as non-enveloped ssDNA viruses. However, AAV capsids embedded within exosomes have been observed, and it has been suggested that the AAV membrane associated accessory protein (MAAP) may play a role in envelope-associated AAV (EA-AAV) capsid formation. Here, we observed and selected sufficient homogeneous EA-AAV capsids of AAV2, produced using the Sf9 baculoviral expression system, to determine the cryo-electron microscopy (cryo-EM) structure at 3.14 Å resolution. The reconstructed map confirmed that the EA-AAV capsid, showed no significant structural variation compared to the non-envelope capsid. In addition, the Sf9 expression system used implies the notion that MAAP may enhance exosome AAV encapsulation. Furthermore, we speculate that these EA-AAV capsids may have therapeutic benefits over the currently used non-envelope AAV capsids, with advantages in immune evasion and/or improved infectivity. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24718.map.gz emd_24718.map.gz | 1.6 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24718-v30.xml emd-24718-v30.xml emd-24718.xml emd-24718.xml | 14.9 KB 14.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_24718.png emd_24718.png | 38.5 KB | ||
| Filedesc metadata |  emd-24718.cif.gz emd-24718.cif.gz | 5.9 KB | ||
| Others |  emd_24718_additional_1.map.gz emd_24718_additional_1.map.gz | 1.6 GB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24718 http://ftp.pdbj.org/pub/emdb/structures/EMD-24718 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24718 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24718 | HTTPS FTP |
-Related structure data
| Related structure data |  7rwlMC  7rwtC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_24718.map.gz / Format: CCP4 / Size: 1.7 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24718.map.gz / Format: CCP4 / Size: 1.7 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | High resolution envelope-associated adeno-associated virus serotype 2. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
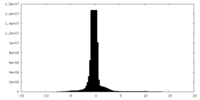
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Low resolution envelope-associated adeno-associated virus serotype 2.
| File | emd_24718_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Low resolution envelope-associated adeno-associated virus serotype 2. | ||||||||||||
| Projections & Slices |
| ||||||||||||
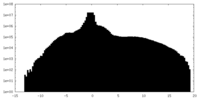
| Density Histograms |
- Sample components
Sample components
-Entire : Adeno-associated virus - 2
| Entire | Name:  Adeno-associated virus - 2 Adeno-associated virus - 2 |
|---|---|
| Components |
|
-Supramolecule #1: Adeno-associated virus - 2
| Supramolecule | Name: Adeno-associated virus - 2 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 10804 / Sci species name: Adeno-associated virus - 2 / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: SEROTYPE / Virus enveloped: Yes / Virus empty: Yes |
|---|---|
| Virus shell | Shell ID: 1 / T number (triangulation number): 1 |
-Macromolecule #1: Capsid protein VP1
| Macromolecule | Name: Capsid protein VP1 / type: protein_or_peptide / ID: 1 / Number of copies: 60 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Adeno-associated dependoparvovirus A Adeno-associated dependoparvovirus A |
| Molecular weight | Theoretical: 82.031352 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAADGYLPDW LEDTLSEGIR QWWKLKPGPP PPKPAERHKD DSRGLVLPGY KYLGPFNGLD KGEPVNEADA AALEHDKAYD RQLDSGDNP YLKYNHADAE FQERLKEDTS FGGNLGRAVF QAKKRVLEPL GLVEEPVKTA PGKKRPVEHS PVEPDSSSGT G KAGQQPAR ...String: MAADGYLPDW LEDTLSEGIR QWWKLKPGPP PPKPAERHKD DSRGLVLPGY KYLGPFNGLD KGEPVNEADA AALEHDKAYD RQLDSGDNP YLKYNHADAE FQERLKEDTS FGGNLGRAVF QAKKRVLEPL GLVEEPVKTA PGKKRPVEHS PVEPDSSSGT G KAGQQPAR KRLNFGQTGD ADSVPDPQPL GQPPAAPSGL GTNTMATGSG APMADNNEGA DGVGNSSGNW HCDSTWMGDR VI TTSTRTW ALPTYNNHLY KQISSQSGAS NDNHYFGYST PWGYFDFNRF HCHFSPRDWQ RLINNNWGFR PKRLNFKLFN IQV KEVTQN DGTTTIANNL TSTVQVFTDS EYQLPYVLGS AHQGCLPPFP ADVFMVPQYG YLTLNNGSQA VGRSSFYCLE YFPS QMLRT GNNFTFSYTF EDVPFHSSYA HSQSLDRLMN PLIDQYLYYL SRTNTPSGTT TQSRLQFSQA GASDIRDQSR NWLPG PCYR QQRVSKTSAD NNNSEYSWTG ATKYHLNGRD SLVNPGPAMA SHKDDEEKFF PQSGVLIFGK QGSEKTNVDI EKVMIT DEE EIRTTNPVAT EQYGSVSTNL QRGNRQAATA DVNTQGVLPG MVWQDRDVYL QGPIWAKIPH TDGHFHPSPL MGGFGLK HP PPQILIKNTP VPANPSTTFS AAKFASFITQ YSTGQVSVEI EWELQKENSK RWNPEIQYTS NYNKSVNVDF TVDTNGVY S EPRPIGTRYL TRNL UniProtKB: Capsid protein VP1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 34.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) X (Row.)
X (Row.) Y (Col.)
Y (Col.)