[English] 日本語
 Yorodumi
Yorodumi- EMDB-22115: CryoEM Structure of E. coli Rho-dependent Transcription Pre-termi... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22115 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM Structure of E. coli Rho-dependent Transcription Pre-termination Complex bound with NusG | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Rho-dependent transcription termination / TRANSCRIPTION | |||||||||
| Function / homology |  Function and homology information Function and homology informationATP-dependent activity, acting on RNA / transcription elongation-coupled chromatin remodeling / RNA polymerase complex / submerged biofilm formation / cellular response to cell envelope stress / bacterial-type RNA polymerase core enzyme binding / cytosolic DNA-directed RNA polymerase complex / regulation of DNA-templated transcription initiation / bacterial-type flagellum assembly / bacterial-type flagellum-dependent cell motility ...ATP-dependent activity, acting on RNA / transcription elongation-coupled chromatin remodeling / RNA polymerase complex / submerged biofilm formation / cellular response to cell envelope stress / bacterial-type RNA polymerase core enzyme binding / cytosolic DNA-directed RNA polymerase complex / regulation of DNA-templated transcription initiation / bacterial-type flagellum assembly / bacterial-type flagellum-dependent cell motility / nitrate assimilation / transcription elongation factor complex / regulation of DNA-templated transcription elongation / helicase activity / transcription antitermination / DNA-templated transcription initiation / cell motility / DNA-templated transcription termination / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / protein complex oligomerization / ribonucleoside binding / DNA-directed 5'-3' RNA polymerase activity / DNA-directed RNA polymerase / ribosome biogenesis / response to heat / protein-containing complex assembly / intracellular iron ion homeostasis / protein dimerization activity / DNA-binding transcription factor activity / protein domain specific binding / response to antibiotic / nucleotide binding / magnesium ion binding / ATP hydrolysis activity / DNA binding / RNA binding / zinc ion binding / ATP binding / identical protein binding / membrane / cytosol / cytoplasm Similarity search - Function | |||||||||
| Biological species |   | |||||||||
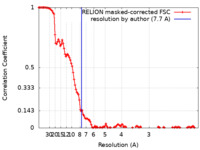
| Method | single particle reconstruction / cryo EM / Resolution: 7.7 Å | |||||||||
 Authors Authors | Hao ZT / Kim HK | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2021 Journal: Mol Cell / Year: 2021Title: Pre-termination Transcription Complex: Structure and Function. Authors: Zhitai Hao / Vitaly Epshtein / Kelly H Kim / Sergey Proshkin / Vladimir Svetlov / Venu Kamarthapu / Binod Bharati / Alexander Mironov / Thomas Walz / Evgeny Nudler /   Abstract: Rho is a general transcription termination factor playing essential roles in RNA polymerase (RNAP) recycling, gene regulation, and genomic stability in most bacteria. Traditional models of ...Rho is a general transcription termination factor playing essential roles in RNA polymerase (RNAP) recycling, gene regulation, and genomic stability in most bacteria. Traditional models of transcription termination postulate that hexameric Rho loads onto RNA prior to contacting RNAP and then translocates along the transcript in pursuit of the moving RNAP to pull RNA from it. Here, we report the cryoelectron microscopy (cryo-EM) structures of two termination process intermediates. Prior to interacting with RNA, Rho forms a specific "pre-termination complex" (PTC) with RNAP and elongation factors NusA and NusG, which stabilize the PTC. RNA exiting RNAP interacts with NusA before entering the central channel of Rho from the distal C-terminal side of the ring. We map the principal interactions in the PTC and demonstrate their critical role in termination. Our results support a mechanism in which the formation of a persistent PTC is a prerequisite for termination. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22115.map.gz emd_22115.map.gz | 11.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22115-v30.xml emd-22115-v30.xml emd-22115.xml emd-22115.xml | 26.1 KB 26.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_22115_fsc.xml emd_22115_fsc.xml | 12.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_22115.png emd_22115.png | 66.2 KB | ||
| Filedesc metadata |  emd-22115.cif.gz emd-22115.cif.gz | 9 KB | ||
| Others |  emd_22115_additional_1.map.gz emd_22115_additional_1.map.gz | 9.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22115 http://ftp.pdbj.org/pub/emdb/structures/EMD-22115 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22115 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22115 | HTTPS FTP |
-Validation report
| Summary document |  emd_22115_validation.pdf.gz emd_22115_validation.pdf.gz | 371.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_22115_full_validation.pdf.gz emd_22115_full_validation.pdf.gz | 370.8 KB | Display | |
| Data in XML |  emd_22115_validation.xml.gz emd_22115_validation.xml.gz | 12.8 KB | Display | |
| Data in CIF |  emd_22115_validation.cif.gz emd_22115_validation.cif.gz | 17.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22115 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22115 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22115 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22115 | HTTPS FTP |
-Related structure data
| Related structure data |  6xavMC  6xasC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22115.map.gz / Format: CCP4 / Size: 163.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22115.map.gz / Format: CCP4 / Size: 163.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.048 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: PTC18 composite map
| File | emd_22115_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | PTC18 composite map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : E. coli Rho-dependent Transcription Pre-termination Complex
+Supramolecule #1: E. coli Rho-dependent Transcription Pre-termination Complex
+Macromolecule #1: Transcription termination/antitermination protein NusG
+Macromolecule #5: DNA-directed RNA polymerase subunit omega
+Macromolecule #6: DNA-directed RNA polymerase subunit alpha
+Macromolecule #7: DNA-directed RNA polymerase subunit beta
+Macromolecule #8: DNA-directed RNA polymerase subunit beta'
+Macromolecule #9: Transcription termination/antitermination protein NusA
+Macromolecule #10: Transcription termination factor Rho
+Macromolecule #2: DNA (29-MER)
+Macromolecule #3: DNA (29-MER)
+Macromolecule #4: RNA (18-MER)
+Macromolecule #11: MAGNESIUM ION
+Macromolecule #12: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-6xav: |
 Movie
Movie Controller
Controller
























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)