+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21258 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | AMC009 SOSIP.v4.2 in complex with PGV04 Fab | |||||||||||||||
 Map data Map data | AMC009 SOSIP.v4.1 in complex with PGV04 Fab | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | HIV / Env / antibody / VIRAL PROTEIN / IMMUNE SYSTEM | |||||||||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /  Homo sapiens (human) Homo sapiens (human) | |||||||||||||||
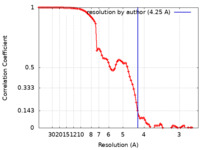
| Method | single particle reconstruction / cryo EM / Resolution: 4.25 Å | |||||||||||||||
 Authors Authors | Cottrell CA / de Val N | |||||||||||||||
| Funding support |  United States, 4 items United States, 4 items
| |||||||||||||||
 Citation Citation |  Journal: J Virol / Year: 2020 Journal: J Virol / Year: 2020Title: Neutralizing Antibody Responses Induced by HIV-1 Envelope Glycoprotein SOSIP Trimers Derived from Elite Neutralizers. Authors: Anna Schorcht / Tom L G M van den Kerkhof / Christopher A Cottrell / Joel D Allen / Jonathan L Torres / Anna-Janina Behrens / Edith E Schermer / Judith A Burger / Steven W de Taeye / Alba ...Authors: Anna Schorcht / Tom L G M van den Kerkhof / Christopher A Cottrell / Joel D Allen / Jonathan L Torres / Anna-Janina Behrens / Edith E Schermer / Judith A Burger / Steven W de Taeye / Alba Torrents de la Peña / Ilja Bontjer / Stephanie Gumbs / Gabriel Ozorowski / Celia C LaBranche / Natalia de Val / Anila Yasmeen / Per Johan Klasse / David C Montefiori / John P Moore / Hanneke Schuitemaker / Max Crispin / Marit J van Gils / Andrew B Ward / Rogier W Sanders /    Abstract: The induction of broadly neutralizing antibodies (bNAbs) is a major goal in vaccine research. HIV-1-infected individuals that develop exceptionally strong bNAb responses, termed elite neutralizers, ...The induction of broadly neutralizing antibodies (bNAbs) is a major goal in vaccine research. HIV-1-infected individuals that develop exceptionally strong bNAb responses, termed elite neutralizers, can inform vaccine design by providing blueprints for the induction of similar bNAb responses. We describe a new recombinant native-like envelope glycoprotein (Env) SOSIP trimer, termed AMC009, based on the viral founder sequences of an elite neutralizer. The subtype B AMC009 SOSIP protein formed stable native-like trimers that displayed multiple bNAb epitopes. Overall, its structure at 4.3-Å resolution was similar to that of BG505 SOSIP.664. The AMC009 trimer resembled one from a second elite neutralizer, AMC011, in having a dense and complete glycan shield. When tested as immunogens in rabbits, the AMC009 trimers did not induce autologous neutralizing antibody (NAb) responses efficiently while the AMC011 trimers did so very weakly, outcomes that may reflect the completeness of their glycan shields. The AMC011 trimer induced antibodies that occasionally cross-neutralized heterologous tier 2 viruses, sometimes at high titer. Cross-neutralizing antibodies were more frequently elicited by a trivalent combination of AMC008, AMC009, and AMC011 trimers, all derived from subtype B viruses. Each of these three individual trimers could deplete the NAb activity from the rabbit sera. Mapping the polyclonal sera by electron microscopy revealed that antibodies of multiple specificities could bind to sites on both autologous and heterologous trimers. These results advance our understanding of how to use Env trimers in multivalent vaccination regimens and the immunogenicity of trimers derived from elite neutralizers. Elite neutralizers, i.e., individuals who developed unusually broad and potent neutralizing antibody responses, might serve as blueprints for HIV-1 vaccine design. Here, we studied the immunogenicity of native-like recombinant envelope glycoprotein (Env) trimers based on viral sequences from elite neutralizers. While immunization with single trimers from elite neutralization did not recapitulate the breadth and potency of neutralization observed in these infected individuals, a combination of three subtype B Env trimers from elite neutralizers resulted in some neutralization breadth within subtype B viruses. These results should guide future efforts to design vaccines to induce broadly neutralizing antibodies. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21258.map.gz emd_21258.map.gz | 59.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21258-v30.xml emd-21258-v30.xml emd-21258.xml emd-21258.xml | 17.2 KB 17.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_21258_fsc.xml emd_21258_fsc.xml | 8.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_21258.png emd_21258.png | 98.9 KB | ||
| Filedesc metadata |  emd-21258.cif.gz emd-21258.cif.gz | 6.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21258 http://ftp.pdbj.org/pub/emdb/structures/EMD-21258 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21258 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21258 | HTTPS FTP |
-Related structure data
| Related structure data |  6vo3MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_21258.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21258.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
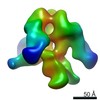
| Annotation | AMC009 SOSIP.v4.1 in complex with PGV04 Fab | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.31 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : AMC009 SOSIP.v4.2 in complex with PGV04 Fab
| Entire | Name: AMC009 SOSIP.v4.2 in complex with PGV04 Fab |
|---|---|
| Components |
|
-Supramolecule #1: AMC009 SOSIP.v4.2 in complex with PGV04 Fab
| Supramolecule | Name: AMC009 SOSIP.v4.2 in complex with PGV04 Fab / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|
-Supramolecule #2: AMC009 SOSIP.v4.2
| Supramolecule | Name: AMC009 SOSIP.v4.2 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
-Supramolecule #3: PGV04 Fab
| Supramolecule | Name: PGV04 Fab / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #3-#4 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: PGV04 heavy chain
| Macromolecule | Name: PGV04 heavy chain / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 24.644771 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLVQSGSG VKKPGASVRV SCWTSEDIFE RTELIHWVRQ APGQGLEWIG WVKTVTGAVN FGSPDFRQRV SLTRDRDLFT AHMDIRGLT QGDTATYFCA RQKFYTGGQG WYFDLWGRGT LIVVSSASTK GPSVFPLAPS SKSTSGGTAA LGCLVKDYFP E PVTVSWNS ...String: QVQLVQSGSG VKKPGASVRV SCWTSEDIFE RTELIHWVRQ APGQGLEWIG WVKTVTGAVN FGSPDFRQRV SLTRDRDLFT AHMDIRGLT QGDTATYFCA RQKFYTGGQG WYFDLWGRGT LIVVSSASTK GPSVFPLAPS SKSTSGGTAA LGCLVKDYFP E PVTVSWNS GALTSGVHTF PAVLQSSGLY SLSSVVTVPS SSLGTQTYIC NVNHKPSNTK VDKKVEPKSC |
-Macromolecule #2: PGV04 light chain
| Macromolecule | Name: PGV04 light chain / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.073822 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EIVLTQSPGT LSLSPGETAS LSCTAASYGH MTWYQKKPGQ PPKLLIFATS KRASGIPDRF SGSQFGKQYT LTITRMEPED FARYYCQQL EFFGQGTRLE IRRTVAAPSV FIFPPSDEQL KSGTASVVCL LNNFYPREAK VQWKVDNALQ SGNSQESVTE Q DSKDSTYS ...String: EIVLTQSPGT LSLSPGETAS LSCTAASYGH MTWYQKKPGQ PPKLLIFATS KRASGIPDRF SGSQFGKQYT LTITRMEPED FARYYCQQL EFFGQGTRLE IRRTVAAPSV FIFPPSDEQL KSGTASVVCL LNNFYPREAK VQWKVDNALQ SGNSQESVTE Q DSKDSTYS LSSTLTLSKA DYEKHKVYAC EVTHQGLSSP VTKSFNRGEC |
-Macromolecule #3: AMC009 SOSIP.v4.2 envelope glycoprotein gp120
| Macromolecule | Name: AMC009 SOSIP.v4.2 envelope glycoprotein gp120 / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 54.113422 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: ARADKLWVTV YYGVPVWKEA TTTLFCASDA KAYDTEVRNV WATHACVPTD PNPQEVVLEN VTENFNMWKN DMVEQMHEDI ISLWDQSLK PCVKLTPLCV TLNCTDYVGN ATNASTTNAT GGIGGTVERG EIKNCSFNIT TSIRDKVQKE YALFYKLDIV P IDNDNTNN ...String: ARADKLWVTV YYGVPVWKEA TTTLFCASDA KAYDTEVRNV WATHACVPTD PNPQEVVLEN VTENFNMWKN DMVEQMHEDI ISLWDQSLK PCVKLTPLCV TLNCTDYVGN ATNASTTNAT GGIGGTVERG EIKNCSFNIT TSIRDKVQKE YALFYKLDIV P IDNDNTNN SYRLINCNTS VIKQACPKVS FEPIPIHYCA PAGFAILKCN DKKFNGTGPC TNVSTVQCTH GIRPVVSTQL LL NGSLAEK EVVIRSQNFT NNAKVIIVQL NESVVINCTR PNNNTRKSIH IAPGRWFYTT GAIIGDIRQA HCNISRVKWN NTL KQIATK LREQFKNKTI AFNQSSGGDP EIVMHSFNCG GEFFYCNTTQ LFNSTWNDTE VSNYNDITHI TLPCRIKQII NMWQ KVGKA MYAPPIRGQI RCSSNITGLL LTRDGGSNEN KTSETETFRP AGGDMRDNWR SELYKYKVVK IEPLGVAPTK CKRRV VQ |
-Macromolecule #4: AMC009 SOSIP.v4.2 envelope glycoprotein gp41
| Macromolecule | Name: AMC009 SOSIP.v4.2 envelope glycoprotein gp41 / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 17.391799 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AVGAIGAVFL GFLGAAGSTM GAASMTLTVQ ARQLLSGIVQ QQNNLLRAPE AQQHMLKLTV WGIKQLQARV LAVERYLRDQ QLLGIWGCS GKLICCTAVP WNNSWSNRSL DMIWNNMTWI EWEREIDNYT GLIYNLLEES QNQQEKNEQE LLELD |
-Macromolecule #8: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 8 / Number of copies: 18 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE / Instrument: HOMEMADE PLUNGER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number real images: 3021 / Average exposure time: 7.0 sec. / Average electron dose: 32.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-6vo3: |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)