+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20966 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
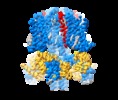
| Title | structure of human KCNQ1-KCNE3-CaM complex | |||||||||
 Map data Map data | human KCNQ1-KCNE3-CaM | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | potassium channel / KCNQ1 / CaM / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of membrane repolarization during ventricular cardiac muscle cell action potential / gastrin-induced gastric acid secretion / corticosterone secretion / voltage-gated potassium channel activity involved in atrial cardiac muscle cell action potential repolarization / negative regulation of voltage-gated potassium channel activity / basolateral part of cell / lumenal side of membrane / negative regulation of delayed rectifier potassium channel activity / rhythmic behavior / stomach development ...negative regulation of membrane repolarization during ventricular cardiac muscle cell action potential / gastrin-induced gastric acid secretion / corticosterone secretion / voltage-gated potassium channel activity involved in atrial cardiac muscle cell action potential repolarization / negative regulation of voltage-gated potassium channel activity / basolateral part of cell / lumenal side of membrane / negative regulation of delayed rectifier potassium channel activity / rhythmic behavior / stomach development / regulation of gastric acid secretion / voltage-gated potassium channel activity involved in cardiac muscle cell action potential repolarization / Phase 3 - rapid repolarisation / membrane repolarization during action potential / negative regulation of potassium ion export across plasma membrane / membrane repolarization during atrial cardiac muscle cell action potential / Phase 2 - plateau phase / iodide transport / regulation of atrial cardiac muscle cell membrane repolarization / membrane repolarization during ventricular cardiac muscle cell action potential / intracellular chloride ion homeostasis / membrane repolarization during cardiac muscle cell action potential / potassium ion export across plasma membrane / voltage-gated potassium channel activity involved in ventricular cardiac muscle cell action potential repolarization / atrial cardiac muscle cell action potential / renal sodium ion absorption / regulation of membrane repolarization / auditory receptor cell development / protein phosphatase 1 binding / detection of mechanical stimulus involved in sensory perception of sound / delayed rectifier potassium channel activity / ventricular cardiac muscle cell action potential / potassium ion homeostasis / Voltage gated Potassium channels / positive regulation of potassium ion transmembrane transport / regulation of ventricular cardiac muscle cell membrane repolarization / non-motile cilium assembly / outward rectifier potassium channel activity / intestinal absorption / CaM pathway / Cam-PDE 1 activation / Sodium/Calcium exchangers / Calmodulin induced events / inner ear morphogenesis / cardiac muscle cell contraction / Reduction of cytosolic Ca++ levels / Activation of Ca-permeable Kainate Receptor / CREB1 phosphorylation through the activation of CaMKII/CaMKK/CaMKIV cascasde / Loss of phosphorylation of MECP2 at T308 / CREB1 phosphorylation through the activation of Adenylate Cyclase / negative regulation of high voltage-gated calcium channel activity / PKA activation / CaMK IV-mediated phosphorylation of CREB / adrenergic receptor signaling pathway / Glycogen breakdown (glycogenolysis) / CLEC7A (Dectin-1) induces NFAT activation / Activation of RAC1 downstream of NMDARs / negative regulation of ryanodine-sensitive calcium-release channel activity / organelle localization by membrane tethering / renal absorption / mitochondrion-endoplasmic reticulum membrane tethering / autophagosome membrane docking / negative regulation of calcium ion export across plasma membrane / regulation of ryanodine-sensitive calcium-release channel activity / ciliary base / regulation of cardiac muscle cell action potential / protein kinase A regulatory subunit binding / protein kinase A catalytic subunit binding / presynaptic endocytosis / neuronal cell body membrane / regulation of heart contraction / Synthesis of IP3 and IP4 in the cytosol / inner ear development / sodium ion transport / potassium ion import across plasma membrane / Phase 0 - rapid depolarisation / Negative regulation of NMDA receptor-mediated neuronal transmission / regulation of heart rate by cardiac conduction / Unblocking of NMDA receptors, glutamate binding and activation / RHO GTPases activate PAKs / calcineurin-mediated signaling / regulation of cell communication by electrical coupling involved in cardiac conduction / Ion transport by P-type ATPases / monoatomic ion channel complex / Uptake and function of anthrax toxins / protein phosphatase activator activity / cochlea development / Long-term potentiation / Calcineurin activates NFAT / action potential / Regulation of MECP2 expression and activity / DARPP-32 events / Smooth Muscle Contraction / potassium channel regulator activity / voltage-gated potassium channel activity / detection of calcium ion / social behavior / regulation of cardiac muscle contraction / positive regulation of heart rate / catalytic complex Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Mackinnon R / Sun J | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2020 Journal: Cell / Year: 2020Title: Structural Basis of Human KCNQ1 Modulation and Gating. Authors: Ji Sun / Roderick MacKinnon /  Abstract: KCNQ1, also known as Kv7.1, is a voltage-dependent K channel that regulates gastric acid secretion, salt and glucose homeostasis, and heart rhythm. Its functional properties are regulated in a tissue- ...KCNQ1, also known as Kv7.1, is a voltage-dependent K channel that regulates gastric acid secretion, salt and glucose homeostasis, and heart rhythm. Its functional properties are regulated in a tissue-specific manner through co-assembly with beta subunits KCNE1-5. In non-excitable cells, KCNQ1 forms a complex with KCNE3, which suppresses channel closure at negative membrane voltages that otherwise would close it. Pore opening is regulated by the signaling lipid PIP2. Using cryoelectron microscopy (cryo-EM), we show that KCNE3 tucks its single-membrane-spanning helix against KCNQ1, at a location that appears to lock the voltage sensor in its depolarized conformation. Without PIP2, the pore remains closed. Upon addition, PIP2 occupies a site on KCNQ1 within the inner membrane leaflet, which triggers a large conformational change that leads to dilation of the pore's gate. It is likely that this mechanism of PIP2 activation is conserved among Kv7 channels. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20966.map.gz emd_20966.map.gz | 93.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20966-v30.xml emd-20966-v30.xml emd-20966.xml emd-20966.xml | 17.2 KB 17.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_20966.png emd_20966.png | 176.9 KB | ||
| Filedesc metadata |  emd-20966.cif.gz emd-20966.cif.gz | 6.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20966 http://ftp.pdbj.org/pub/emdb/structures/EMD-20966 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20966 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20966 | HTTPS FTP |
-Related structure data
| Related structure data |  6v00MC  6uzzC  6v01C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_20966.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20966.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | human KCNQ1-KCNE3-CaM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.03 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : KCNQ1-CaM complex
| Entire | Name: KCNQ1-CaM complex |
|---|---|
| Components |
|
-Supramolecule #1: KCNQ1-CaM complex
| Supramolecule | Name: KCNQ1-CaM complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Potassium voltage-gated channel subfamily KQT member 1
| Macromolecule | Name: Potassium voltage-gated channel subfamily KQT member 1 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 63.258574 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MASDLGPRPP VSLDPRVSIY STRRPVLART HVQGRVYNFL ERPTGWKCFV YHFAVFLIVL VCLIFSVLST IEQYAALATG TLFWMEIVL VVFFGTEYVV RLWSAGCRSK YVGLWGRLRF ARKPISIIDL IVVVASMVVL CVGSKGQVFA TSAIRGIRFL Q ILRMLHVD ...String: MASDLGPRPP VSLDPRVSIY STRRPVLART HVQGRVYNFL ERPTGWKCFV YHFAVFLIVL VCLIFSVLST IEQYAALATG TLFWMEIVL VVFFGTEYVV RLWSAGCRSK YVGLWGRLRF ARKPISIIDL IVVVASMVVL CVGSKGQVFA TSAIRGIRFL Q ILRMLHVD RQGGTWRLLG SVVFIHRQEL ITTLYIGFLG LIFSSYFVYL AEKDAVNESG RVEFGSYADA LWWGVVTVTT IG YGDKVPQ TWVGKTIASC FSVFAISFFA LPAGILGSGF ALKVQQKQRQ KHFNRQIPAA ASLIQTAWRC YAAENPDSST WKI YIRKAP RSHTLLSPSP KPKKSVVVKK KKFKLDKDNG VTPGEKMLTV PHITCDPPEE RRLDHFSVDG YDSSVRKSPT LLEV SMPHF MRTNSFAEDL DLEGETLLTP ITHISQLREH HRATIKVIRR MQYFVAKKKF QQARKPYDVR DVIEQYSQGH LNLMV RIKE LQRRLDQSIG KPSLFISVSE KSKDRGSNTI GARLNRVEDK VTQLDQRLAL ITDMLHQLLS LHSNSLEVLF QGP UniProtKB: Potassium voltage-gated channel subfamily KQT member 1 |
-Macromolecule #2: Calmodulin-1
| Macromolecule | Name: Calmodulin-1 / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 16.852545 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MADQLTEEQI AEFKEAFSLF DKDGDGTITT KELGTVMRSL GQNPTEAELQ DMINEVDADG NGTIDFPEFL TMMARKMKDT DSEEEIREA FRVFDKDGNG YISAAELRHV MTNLGEKLTD EEVDEMIREA DIDGDGQVNY EEFVQMMTAK UniProtKB: Calmodulin-1 |
-Macromolecule #3: MCherry fluorescent protein,Potassium voltage-gated channel subfa...
| Macromolecule | Name: MCherry fluorescent protein,Potassium voltage-gated channel subfamily E member 3 type: protein_or_peptide / ID: 3 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.095168 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GGMVSKGEED NMAIIKEFMR FKVHMEGSVN GHEFEIEGEG EGRPYEGTQT AKLKVTKGGP LPFAWDILSP QFMYGSKAYV KHPADIPDY LKLSFPEGFN WERVMNFEDG GVVTVTQDSS LQDGEFIYKV KLRGTNFPSD GPVMQCRTMG WEASTERMYP E DGALKGEI ...String: GGMVSKGEED NMAIIKEFMR FKVHMEGSVN GHEFEIEGEG EGRPYEGTQT AKLKVTKGGP LPFAWDILSP QFMYGSKAYV KHPADIPDY LKLSFPEGFN WERVMNFEDG GVVTVTQDSS LQDGEFIYKV KLRGTNFPSD GPVMQCRTMG WEASTERMYP E DGALKGEI KQRLKLKDGG HYDAEVKTTY KAKKPVQLPG AYNVDIKLDI LSHNEDYTIV EQYERAEGRH STGGMDELYK GS GENLYFQ SSRATMETTN GTETWYESLH AVLKALNATL HSNLLCRPGP GLGPDNQTEE RRASLPGRDD NSYMYILFVM FLF AVTVGS LILGYTRSRK VDKRSDPYHV YIKNRVSMI UniProtKB: MCherry fluorescent protein, Potassium voltage-gated channel subfamily E member 3 |
-Macromolecule #4: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 4 / Number of copies: 8 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 94.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-6v00: |
 Movie
Movie Controller
Controller


































 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)





















