[English] 日本語
 Yorodumi
Yorodumi- EMDB-20206: Symmetric reconstruction of human norovirus GII.4 Minerva strain ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20206 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Symmetric reconstruction of human norovirus GII.4 Minerva strain VLP in T=4 symmetry | |||||||||
 Map data Map data | Symmetric reconstruction of human norovirus GII.4 Minerva strain VLP in T=4 symmetry | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Caliciviridae / Calicivirus / Norovirus / GII.4 / VIRUS LIKE PARTICLE | |||||||||
| Function / homology | Calicivirus coat protein C-terminal / Calicivirus coat protein C-terminal / Calicivirus coat protein / Calicivirus coat protein / virion component / Picornavirus/Calicivirus coat protein / Viral coat protein subunit / host cell cytoplasm / Major capsid protein Function and homology information Function and homology information | |||||||||
| Biological species |  Norovirus Hu/GII.4/Minerva/2006/USA Norovirus Hu/GII.4/Minerva/2006/USA | |||||||||
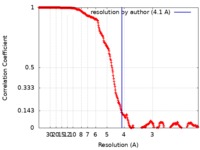
| Method | single particle reconstruction / cryo EM / Resolution: 4.1 Å | |||||||||
 Authors Authors | Jung J / Grant T | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2019 Journal: Proc Natl Acad Sci U S A / Year: 2019Title: High-resolution cryo-EM structures of outbreak strain human norovirus shells reveal size variations. Authors: James Jung / Timothy Grant / Dennis R Thomas / Chris W Diehnelt / Nikolaus Grigorieff / Leemor Joshua-Tor /  Abstract: Noroviruses are a leading cause of foodborne illnesses worldwide. Although GII.4 strains have been responsible for most norovirus outbreaks, the assembled virus shell structures have been available ...Noroviruses are a leading cause of foodborne illnesses worldwide. Although GII.4 strains have been responsible for most norovirus outbreaks, the assembled virus shell structures have been available in detail for only a single strain (GI.1). We present high-resolution (2.6- to 4.1-Å) cryoelectron microscopy (cryo-EM) structures of GII.4, GII.2, GI.7, and GI.1 human norovirus outbreak strain virus-like particles (VLPs). Although norovirus VLPs have been thought to exist in a single-sized assembly, our structures reveal polymorphism between and within genogroups, with small, medium, and large particle sizes observed. Using asymmetric reconstruction, we were able to resolve a Zn metal ion adjacent to the coreceptor binding site, which affected the structural stability of the shell. Our structures serve as valuable templates for facilitating vaccine formulations. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20206.map.gz emd_20206.map.gz | 1.8 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20206-v30.xml emd-20206-v30.xml emd-20206.xml emd-20206.xml | 15.3 KB 15.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_20206_fsc.xml emd_20206_fsc.xml | 32.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_20206.png emd_20206.png | 200.3 KB | ||
| Filedesc metadata |  emd-20206.cif.gz emd-20206.cif.gz | 6.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20206 http://ftp.pdbj.org/pub/emdb/structures/EMD-20206 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20206 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20206 | HTTPS FTP |
-Related structure data
| Related structure data |  6ouuMC  6otfC  6ou9C  6oucC  6outC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_20206.map.gz / Format: CCP4 / Size: 1.9 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20206.map.gz / Format: CCP4 / Size: 1.9 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Symmetric reconstruction of human norovirus GII.4 Minerva strain VLP in T=4 symmetry | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Norovirus Hu/GII.4/Minerva/2006/USA
| Entire | Name:  Norovirus Hu/GII.4/Minerva/2006/USA Norovirus Hu/GII.4/Minerva/2006/USA |
|---|---|
| Components |
|
-Supramolecule #1: Norovirus Hu/GII.4/Minerva/2006/USA
| Supramolecule | Name: Norovirus Hu/GII.4/Minerva/2006/USA / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 1182144 / Sci species name: Norovirus Hu/GII.4/Minerva/2006/USA / Sci species strain: GII.4 / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: Yes |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 14.16 MDa |
| Virus shell | Shell ID: 1 / Name: VP1 / Diameter: 490.0 Å / T number (triangulation number): 4 |
-Macromolecule #1: Major capsid protein
| Macromolecule | Name: Major capsid protein / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Norovirus Hu/GII.4/Minerva/2006/USA Norovirus Hu/GII.4/Minerva/2006/USA |
| Molecular weight | Theoretical: 58.938098 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKMASNDANP SDGSAANLVP EVNNEVMALE PVVGAAIAAP VAGQQNVIDP WIRNNFVQAP GGEFTVSPRN APGEILWSAP LGPDLNPYL SHLARMYNGY AGGFEVQVIL AGNAFTAGKI IFAAVPPNFP TEGLSPSQVT MFPHIIVDVR QLEPVLIPLP D VRNNFYHY ...String: MKMASNDANP SDGSAANLVP EVNNEVMALE PVVGAAIAAP VAGQQNVIDP WIRNNFVQAP GGEFTVSPRN APGEILWSAP LGPDLNPYL SHLARMYNGY AGGFEVQVIL AGNAFTAGKI IFAAVPPNFP TEGLSPSQVT MFPHIIVDVR QLEPVLIPLP D VRNNFYHY NQSNDSTIKL IAMLYTPLRA NNAGEDVFTV SCRVLTRPSP DFDFIFLVPP TVESRTKPFT VPILTVEEMT NS RFPIPLE KLFTGPSGAF VVQPQNGRCT TDGVLLGTTQ LSPVNICTFR GDVTHIAGSR NYTMNLASLN WNNYDPTEEI PAP LGTPDF VGKIQGVLTQ TTKGDGSTRG HKATVYTGSA PFTPKLGSVQ FSTDTENDFE THQNTKFTPV GVIQDGSTTH RNEP QQWVL PSYSGRNVHN VHLAPAVAPT FPGEQLLFFR STMPGCSGYP NMDLDCLLPQ EWVQHFYQEA APAQSDVALL RFVNP DTGR VLFECKLHKS GYVTVAHTGQ HDLVIPPNGY FRFDSWVNQF YTLAPMGNGT GGRRAL UniProtKB: Major capsid protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4 mg/mL |
|---|---|
| Buffer | pH: 5.75 |
| Grid | Material: COPPER / Support film - Material: CARBON / Support film - topology: LACEY / Pretreatment - Type: GLOW DISCHARGE / Details: unspecified |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 295 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Number real images: 2951 / Average exposure time: 7.0 sec. / Average electron dose: 69.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.6 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A / Chain - Residue range: 225-529 / Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
| Output model |  PDB-6ouu: |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)